A Swiss Army Knife of Methods
Deciphering the structure, dynamics and interactions of non-canonical DNA
ARNA, INSERM U1212, CNRS UMR 5320, Université de Bordeaux
UFR des Sciences Pharmaceutiques, Université de Bordeaux
May 12, 2026
Hello from Bordeaux



Conformational polymorphism
of DNA
DNA can adopt many secondary structures
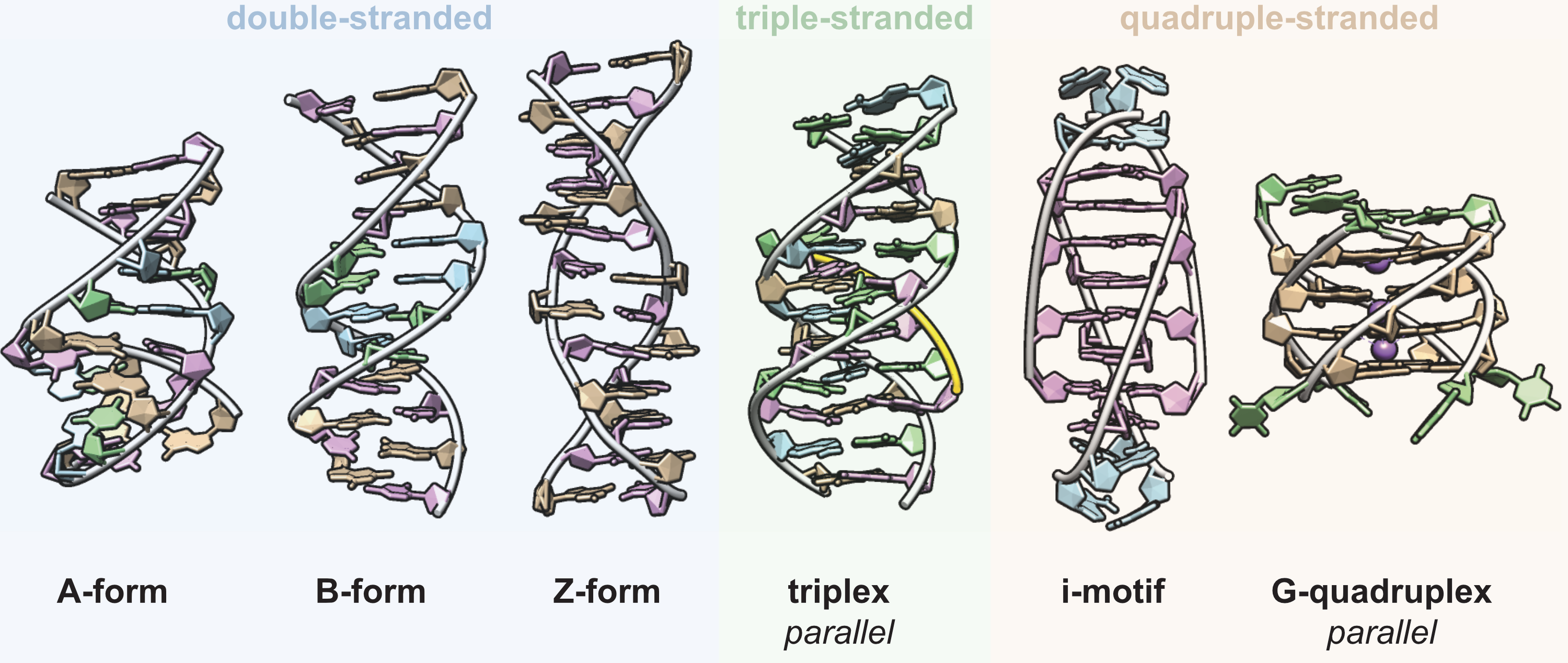
Largy et al., Chem. Rev. 2022, 122, 8, 7720
G-quadruplexes are particularly peculiar
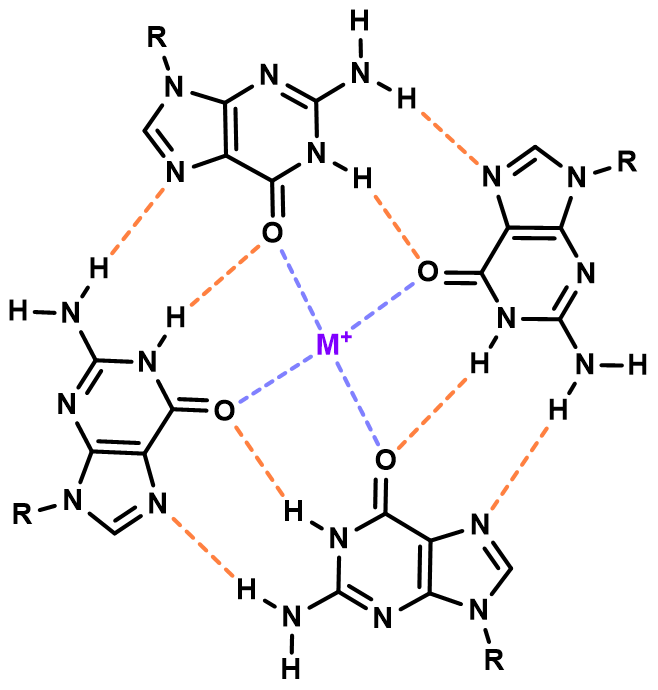
G-quadruplexes are particularly peculiar

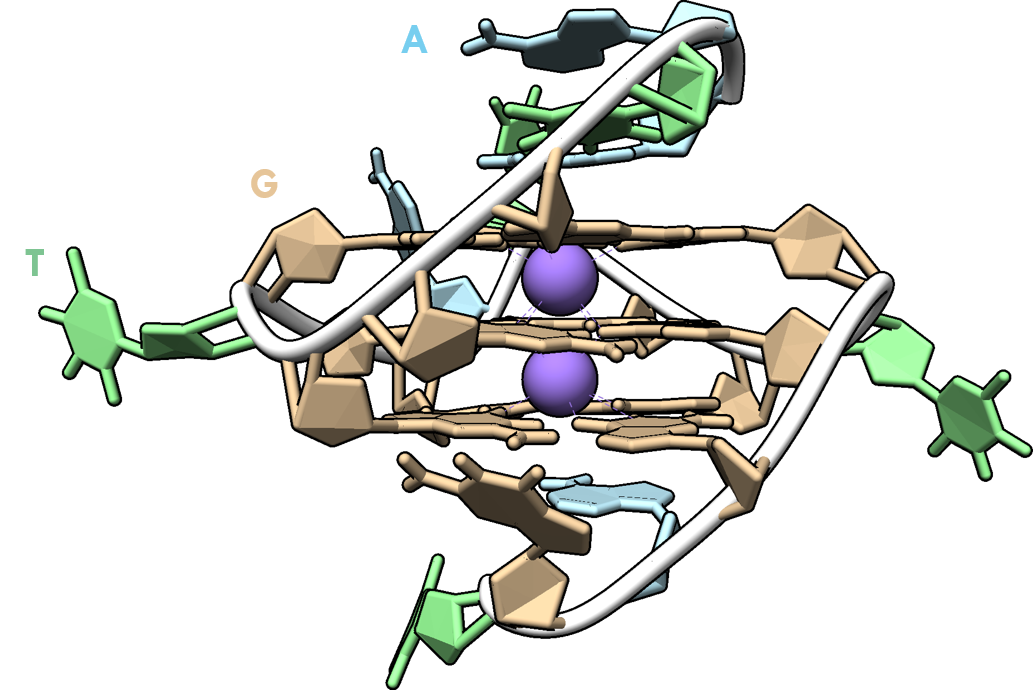
G4s are DNA/RNA drug targets distinct from dsDNA
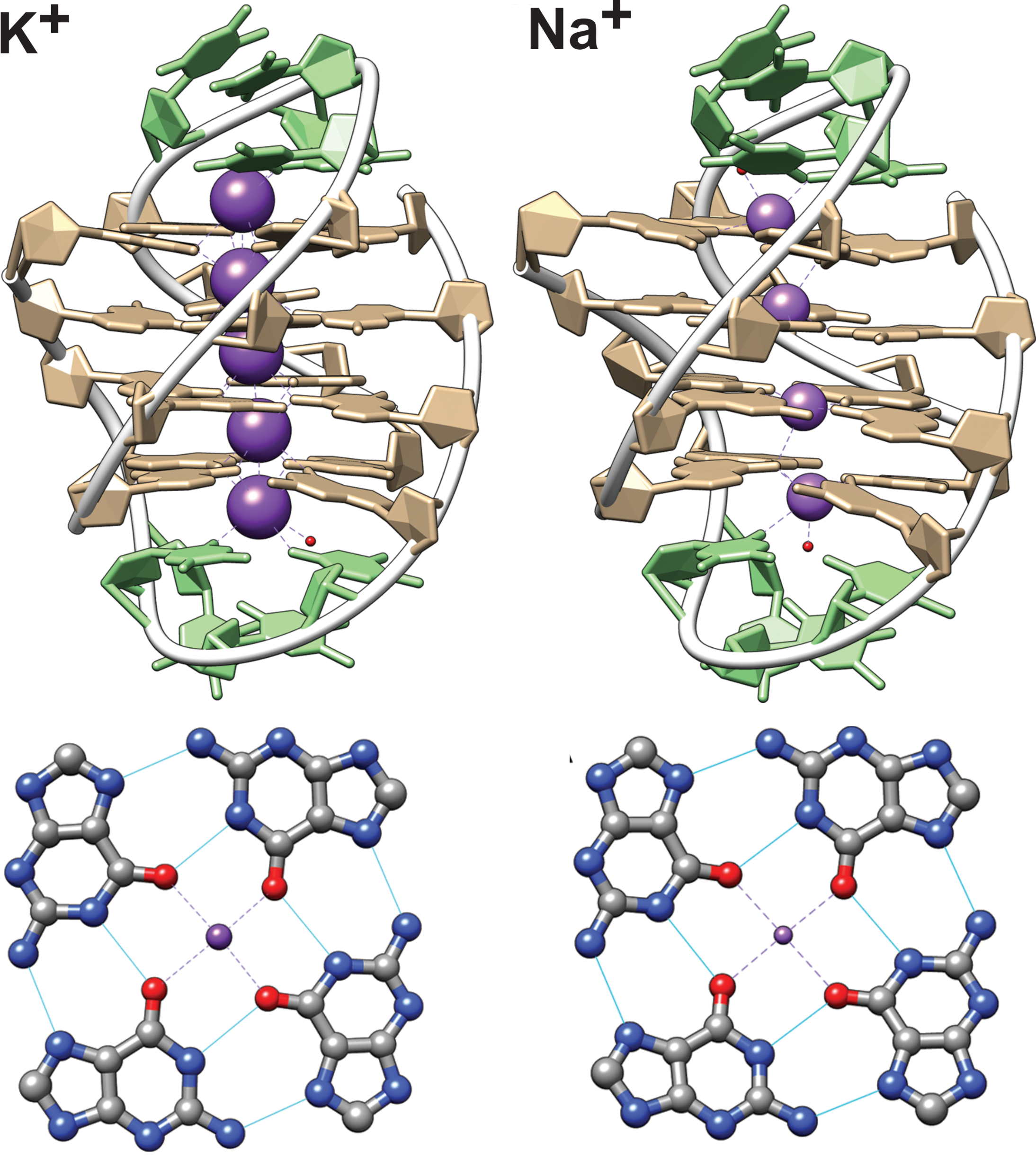
Largy, E., Mergny, J.-L. et Gabelica, V. Role of Alkali Metal Ions in G-Quadruplex Nucleic Acid Structure and Stability. 2016, p.203‑258.
G4s may also be drugs themselves

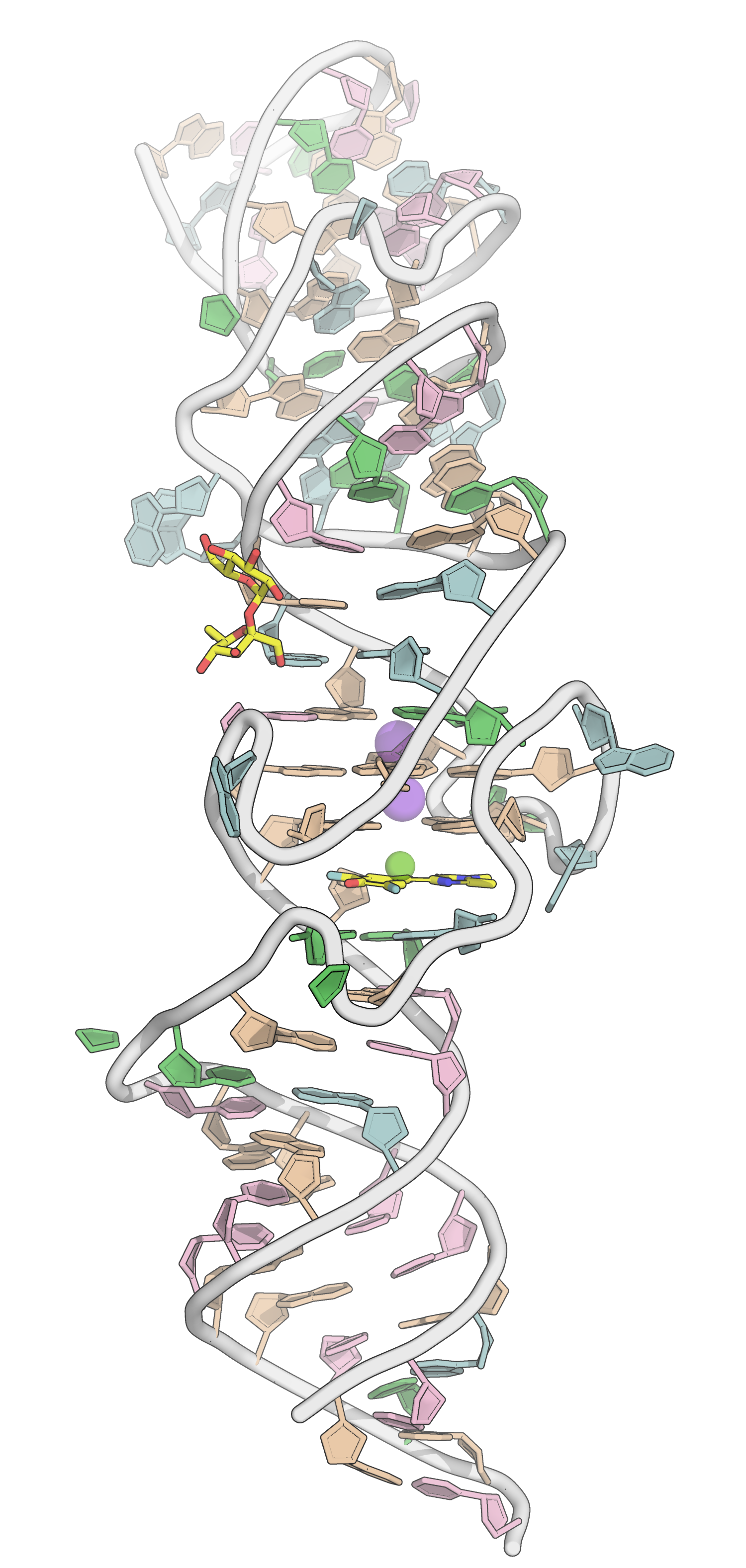
PDB: 4TS2: Warner et al., Nat. Struc. Mol. Biol., 2014, 21, 658
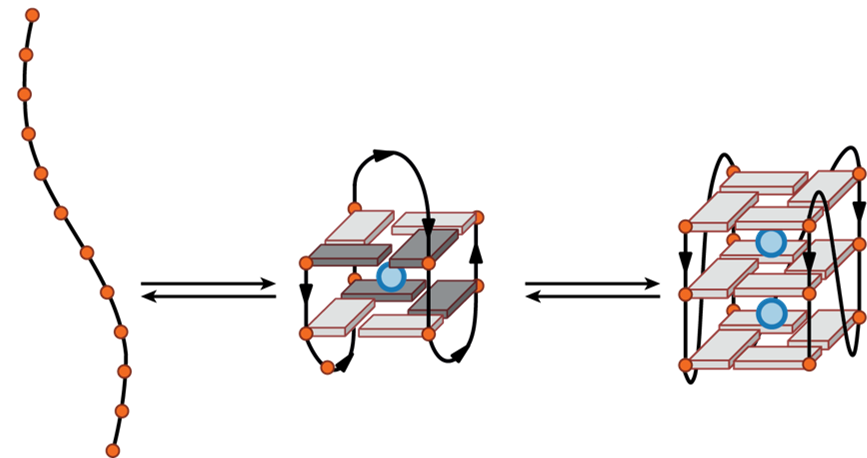
G4s are very polymorphic…
…and in equilibrium

Topology, stability and stoichiometry determination
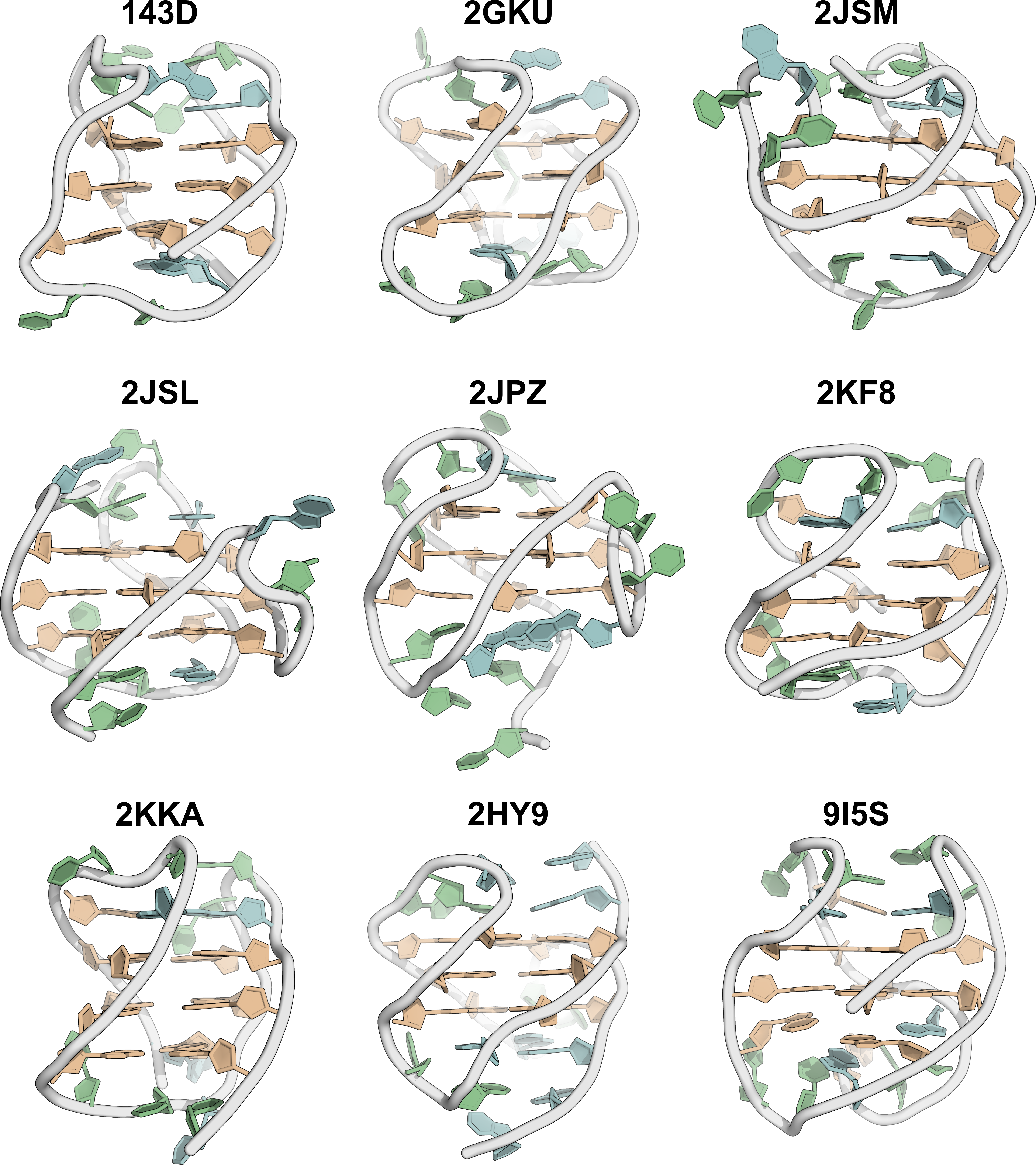
Topologies are poor structural descriptors
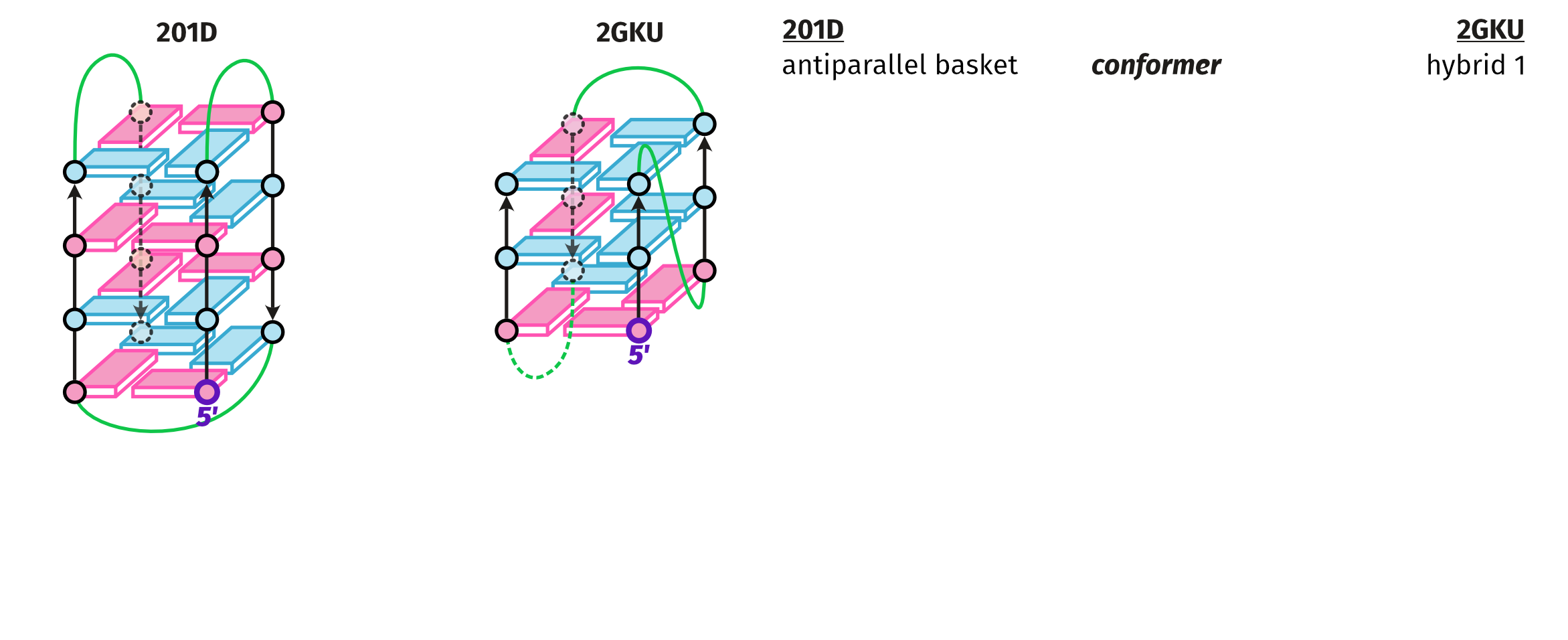
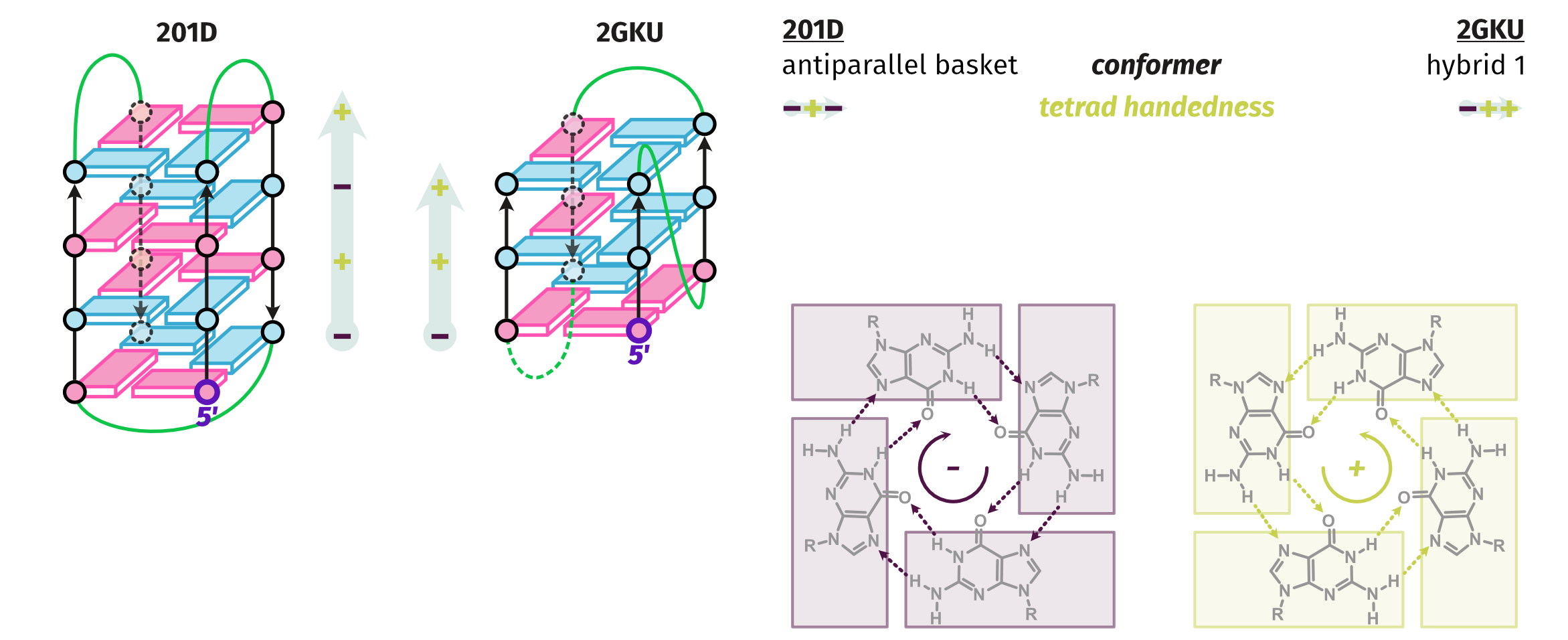
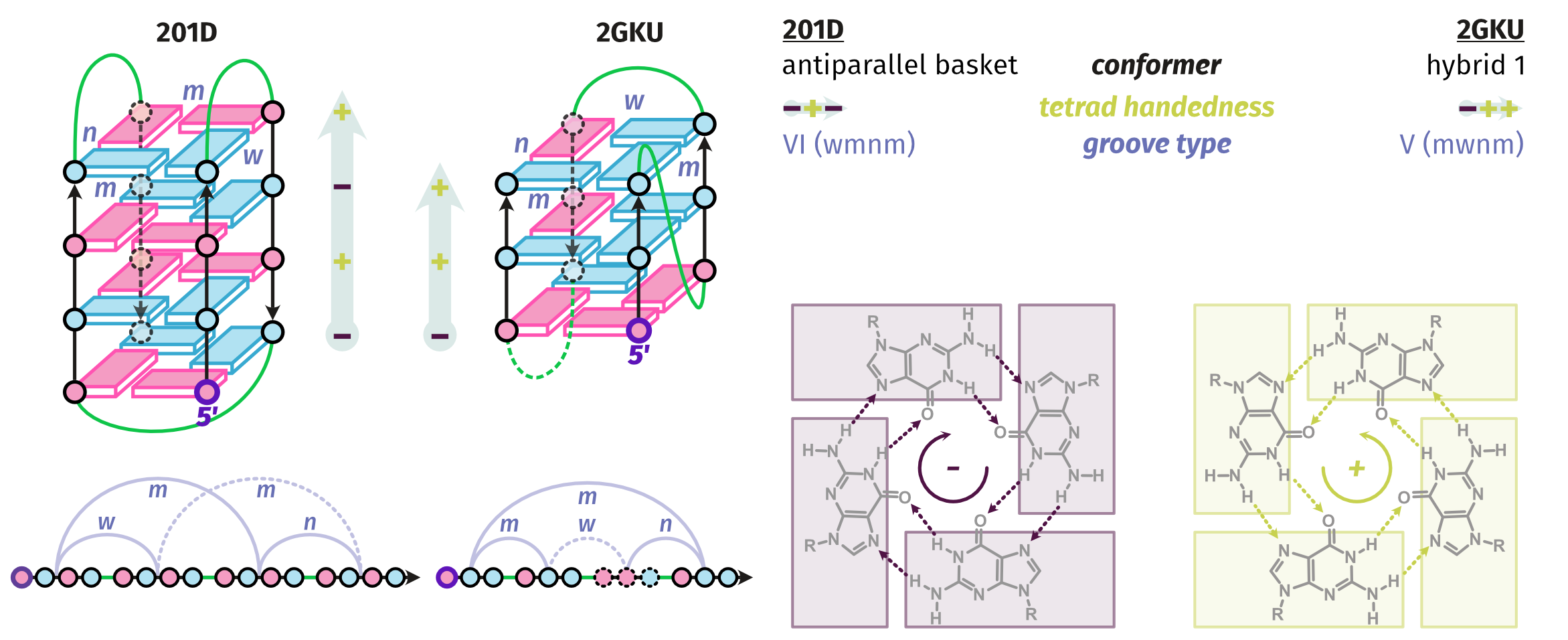
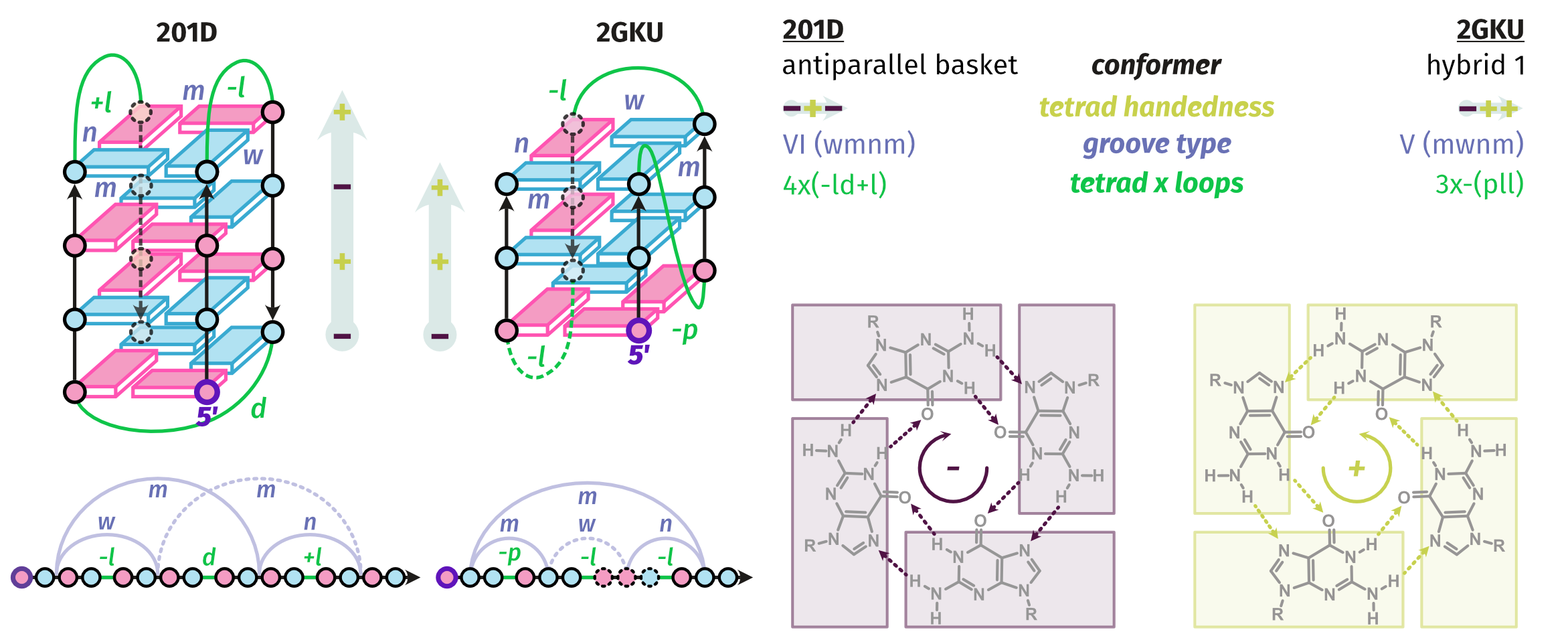
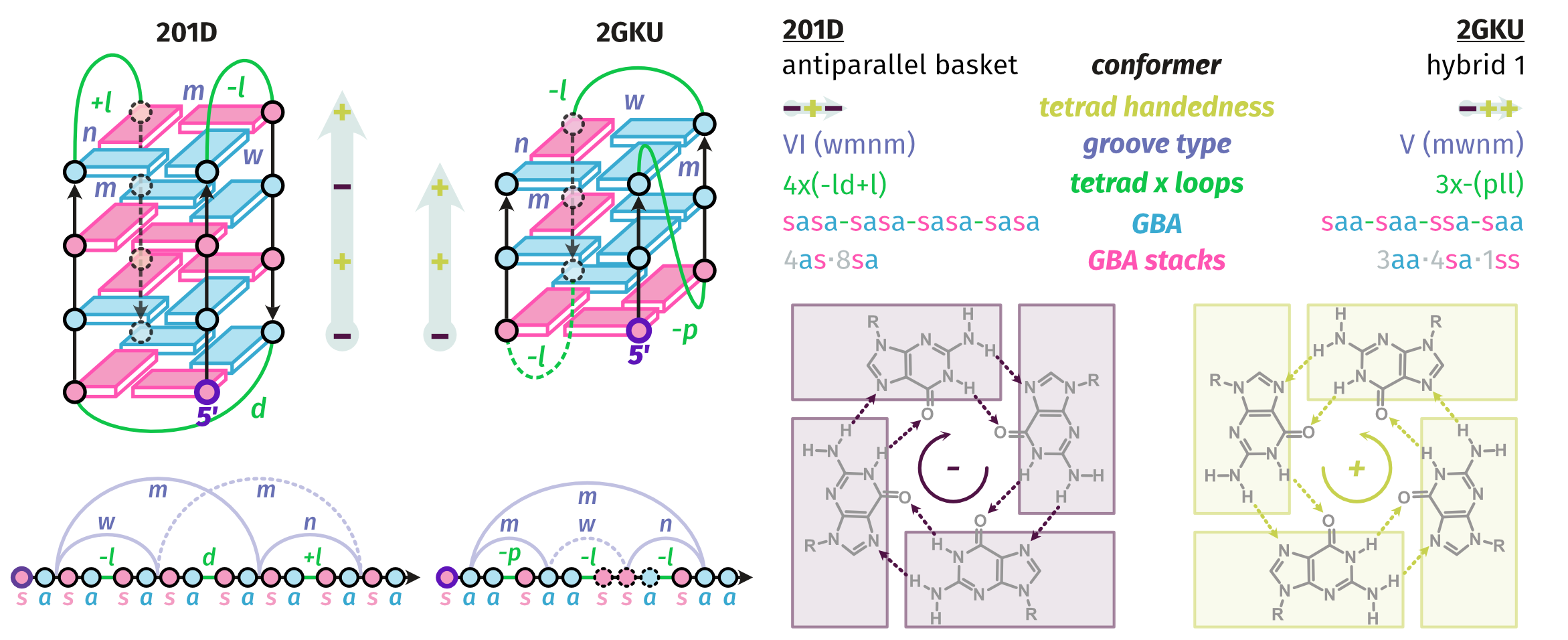


Topologies are poor structural descriptors
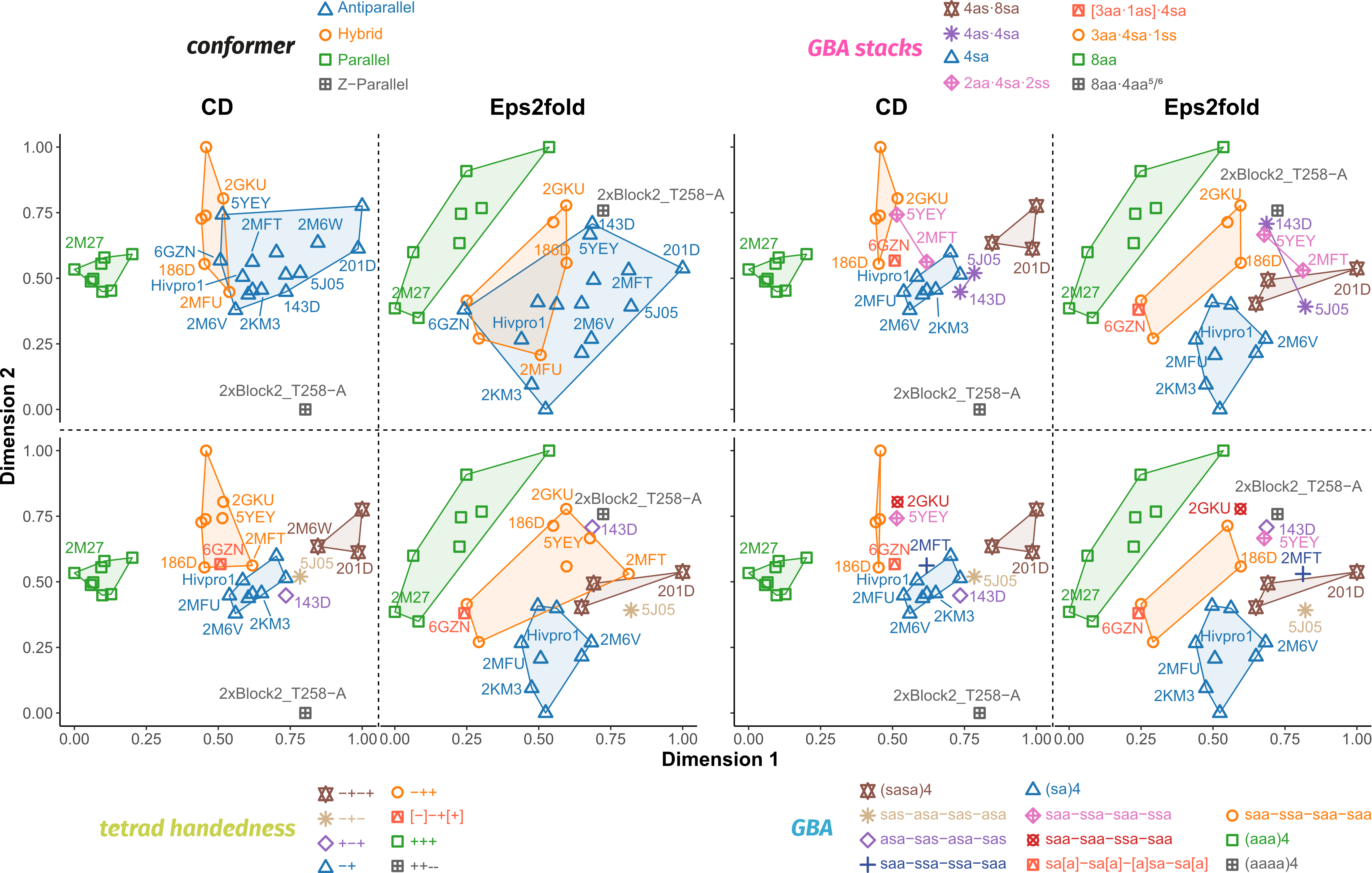
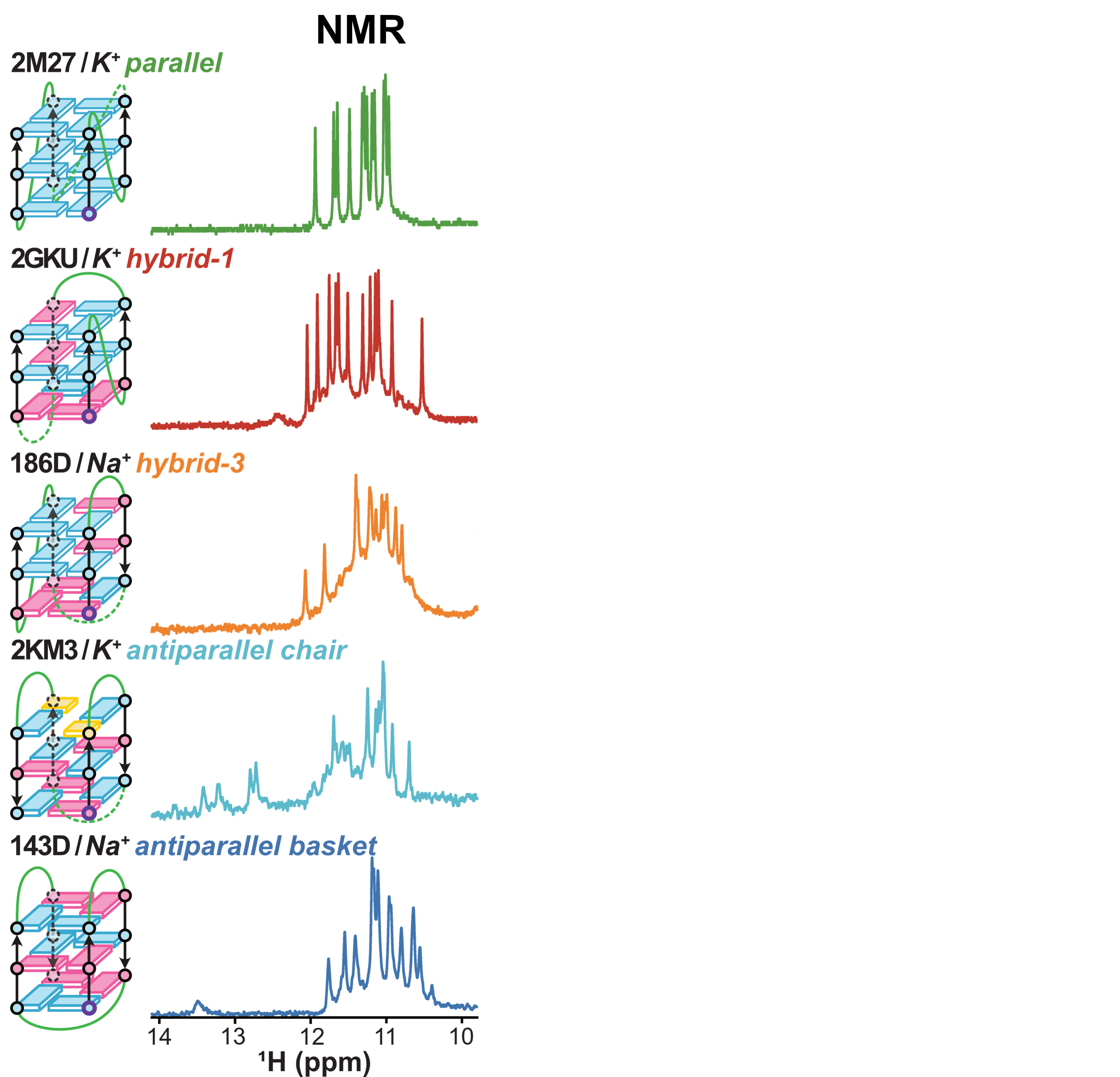
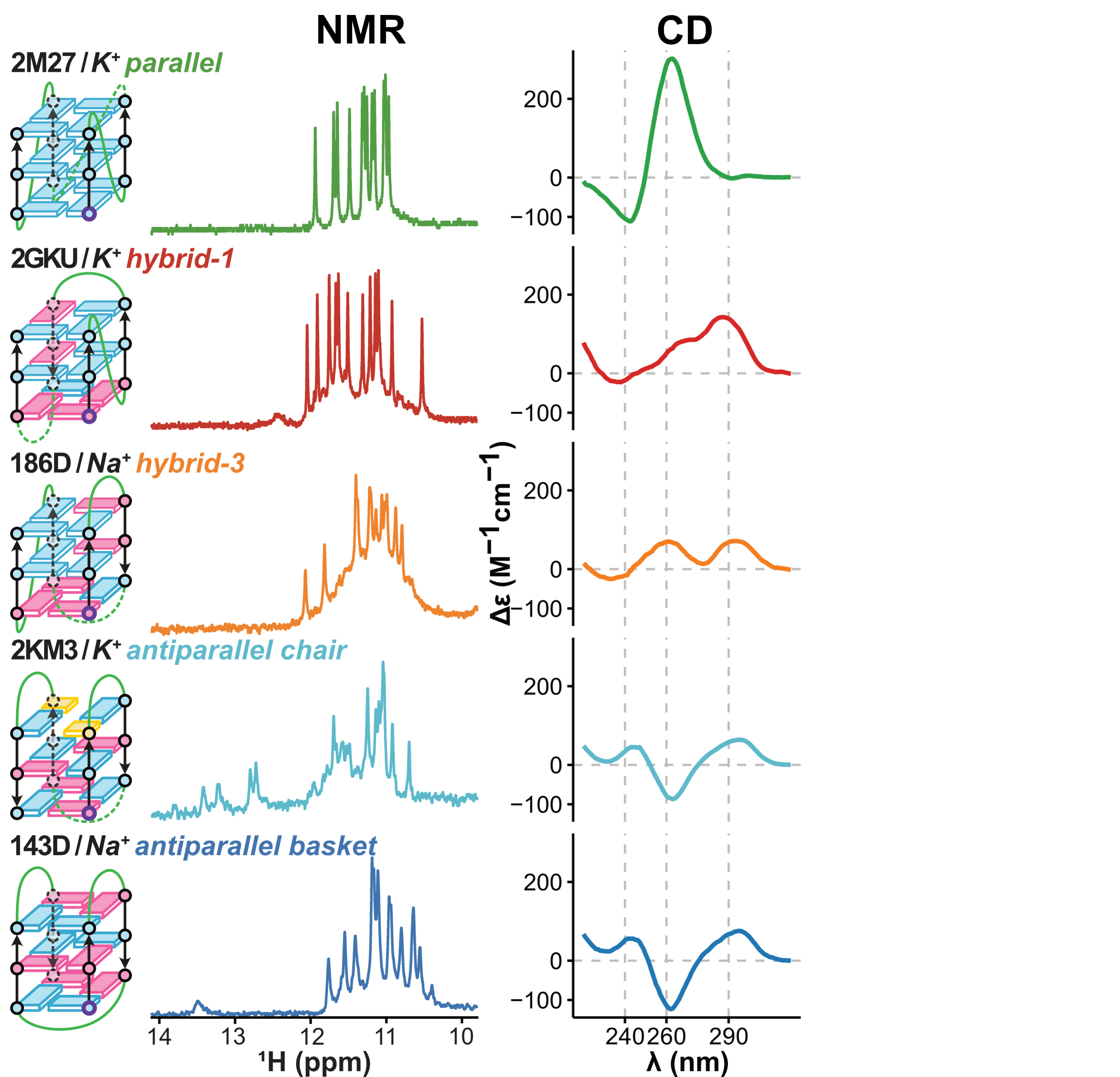
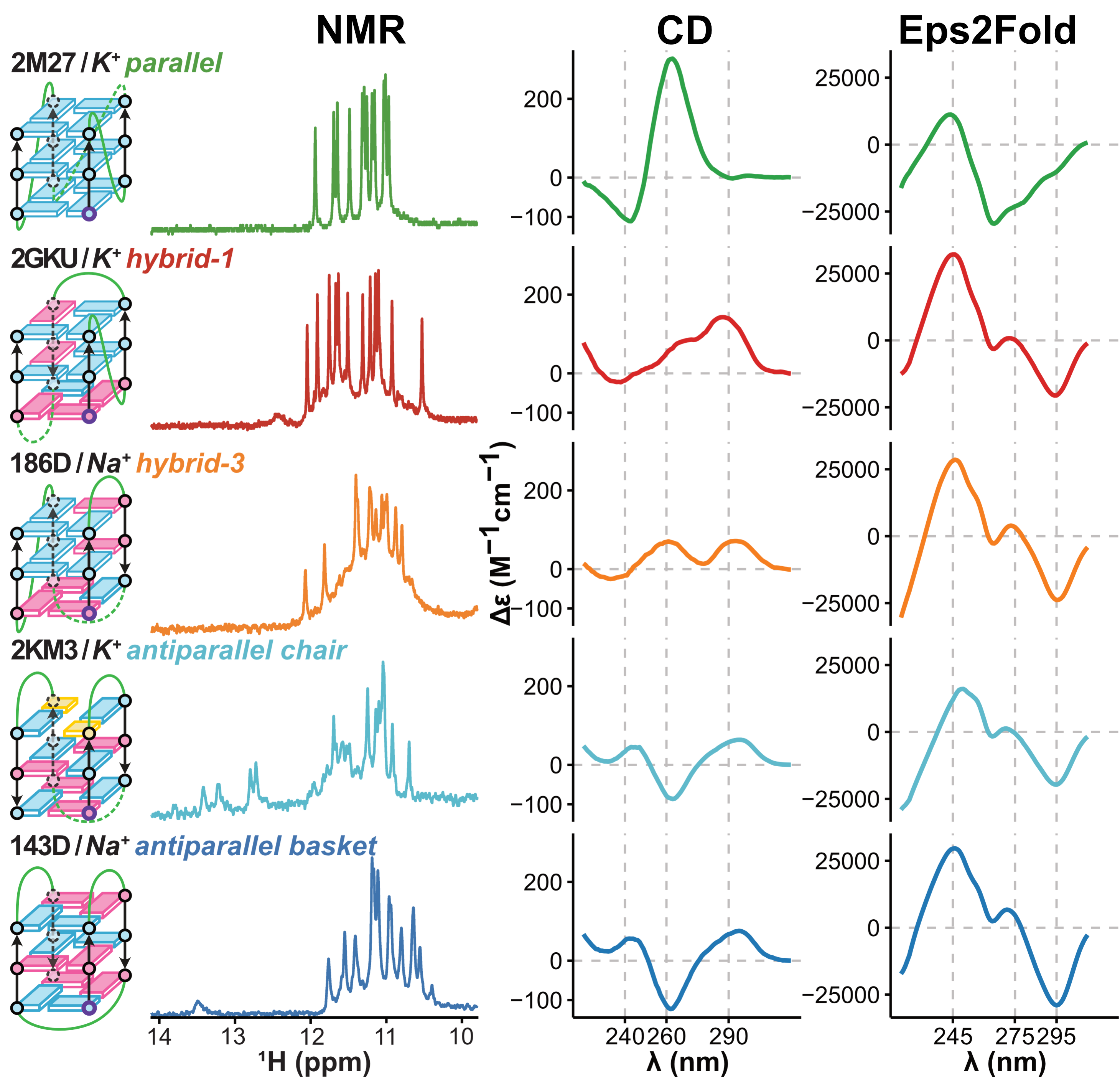


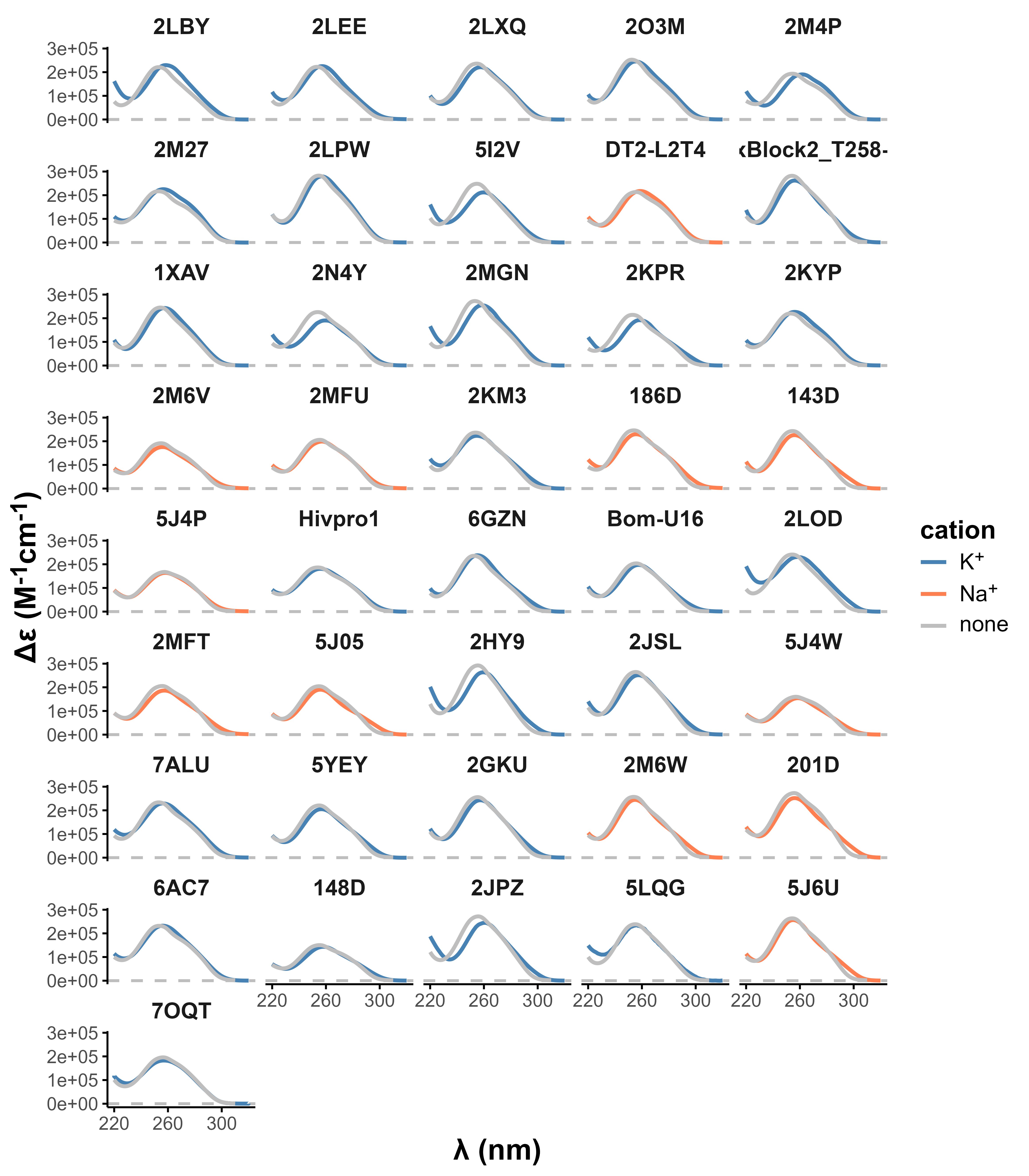
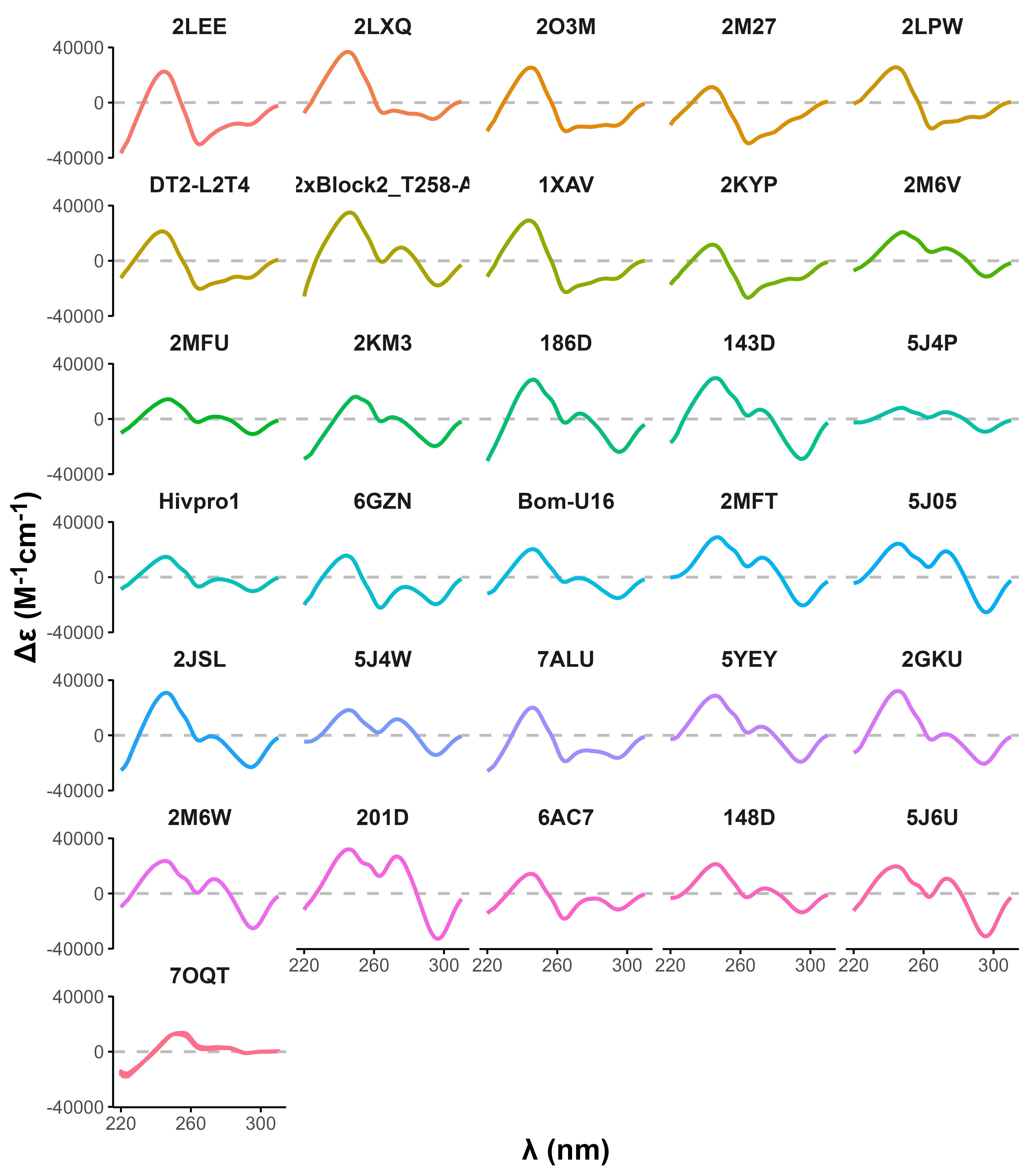


Largy, E., Guédin, A., Kabbara, A., Mergny, J.-L. et Amrane, S. Nucleic Acids Res., 2025, 53, gkaf953
Stability measurements are often innacurate
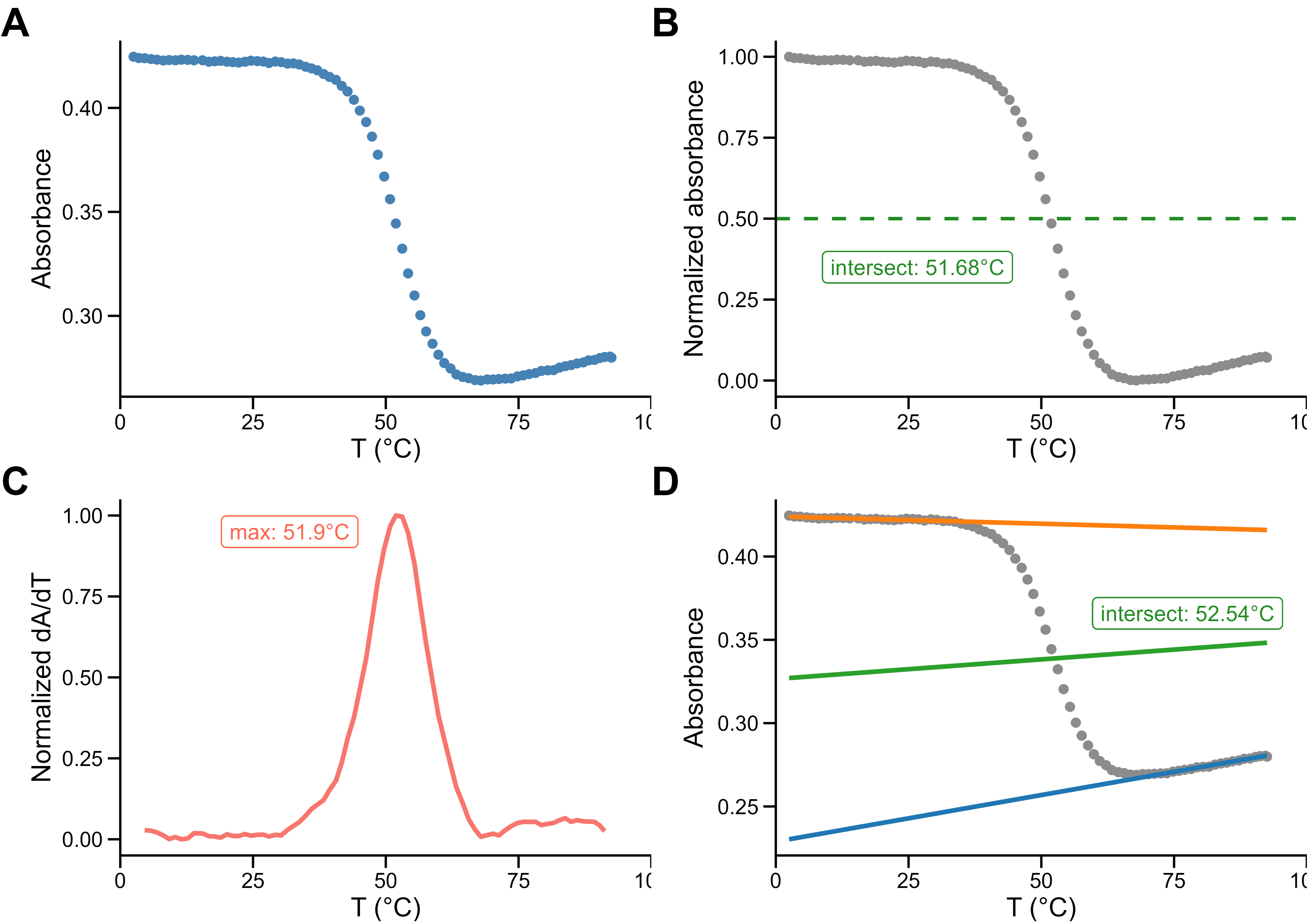
Stability measurements are often innacurate
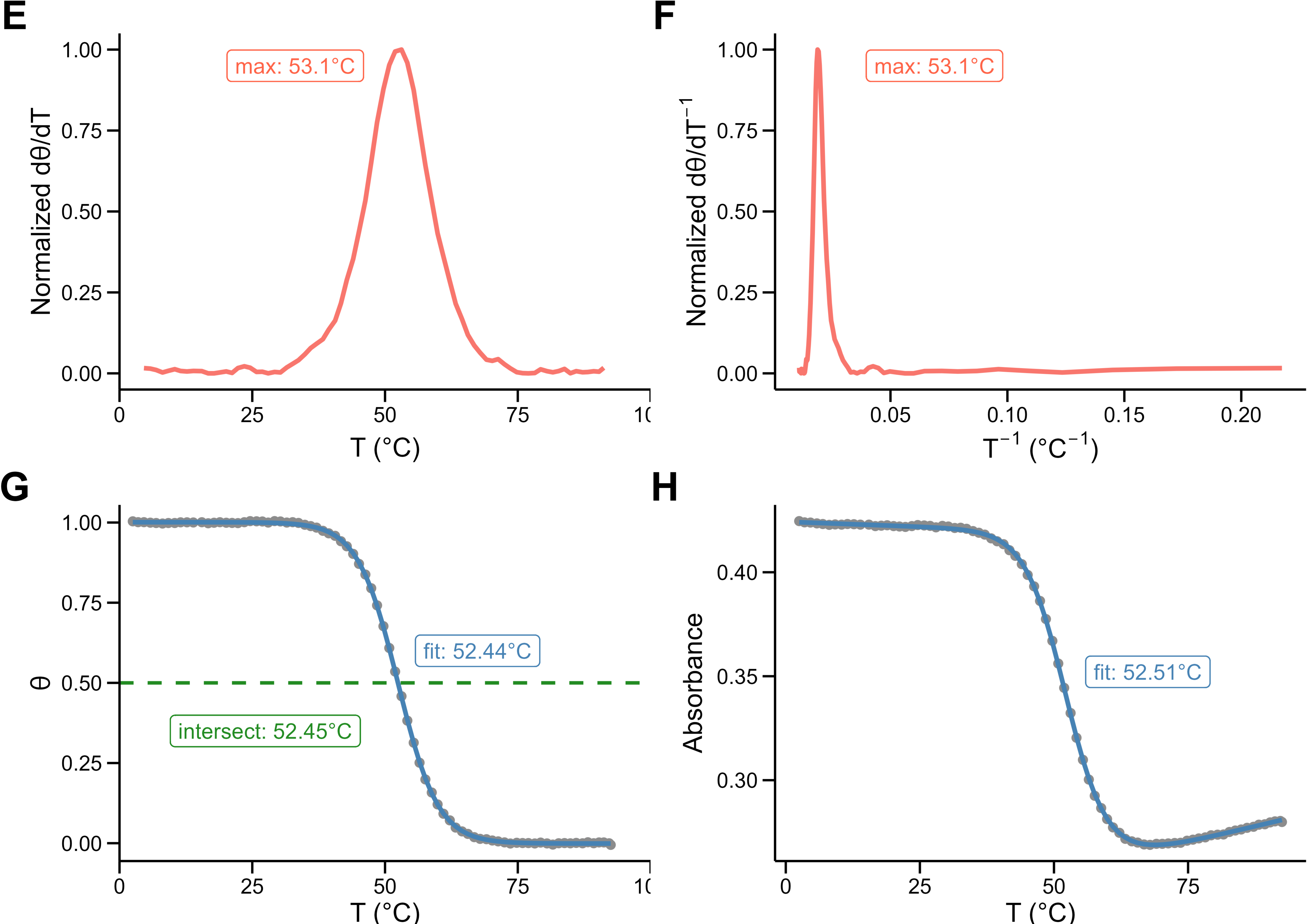
\[ A_T=(a^FT+b^F) \times \frac{1}{1+exp(-\frac{\Delta H^0 (1- \frac{T}{T_m})}{RT})} + (a^UT+b^U) \times \frac{exp(-\frac{\Delta H^0 (1- \frac{T}{T_m})}{RT})}{1+exp(-\frac{\Delta H^0 (1- \frac{T}{T_m})}{RT})} \]
\[ A_T=\textcolor{forestgreen}{(a^FT+b^F)} \times \frac{1}{1+exp(-\frac{\Delta H^0 (1- \frac{T}{T_m})}{RT})} + \textcolor{forestgreen}{(a^UT+b^U)} \times \frac{exp(-\frac{\Delta H^0 (1- \frac{T}{T_m})}{RT})}{1+exp(-\frac{\Delta H^0 (1- \frac{T}{T_m})}{RT})} \]
\[ A_T=(a^FT+b^F) \times \frac{1}{1+exp(-\frac{\Delta H^0 (1- \frac{T}{\textcolor{coral}{T_m}})}{RT})} + (a^UT+b^U) \times \frac{exp(-\frac{\Delta H^0 (1- \frac{T}{\textcolor{coral}{T_m}})}{RT})}{1+exp(-\frac{\Delta H^0 (1- \frac{T}{\textcolor{coral}{T_m}})}{RT})} \]
\[ A_T=(a^FT+b^F) \times \frac{1}{1+exp(-\frac{\textcolor{steelblue}{\Delta H^0} (1- \frac{T}{T_m})}{RT})} + (a^UT+b^U) \times \frac{exp(-\frac{\textcolor{steelblue}{\Delta H^0} (1- \frac{T}{T_m})}{RT})}{1+exp(-\frac{\textcolor{steelblue}{\Delta H^0} (1- \frac{T}{T_m})}{RT})} \]
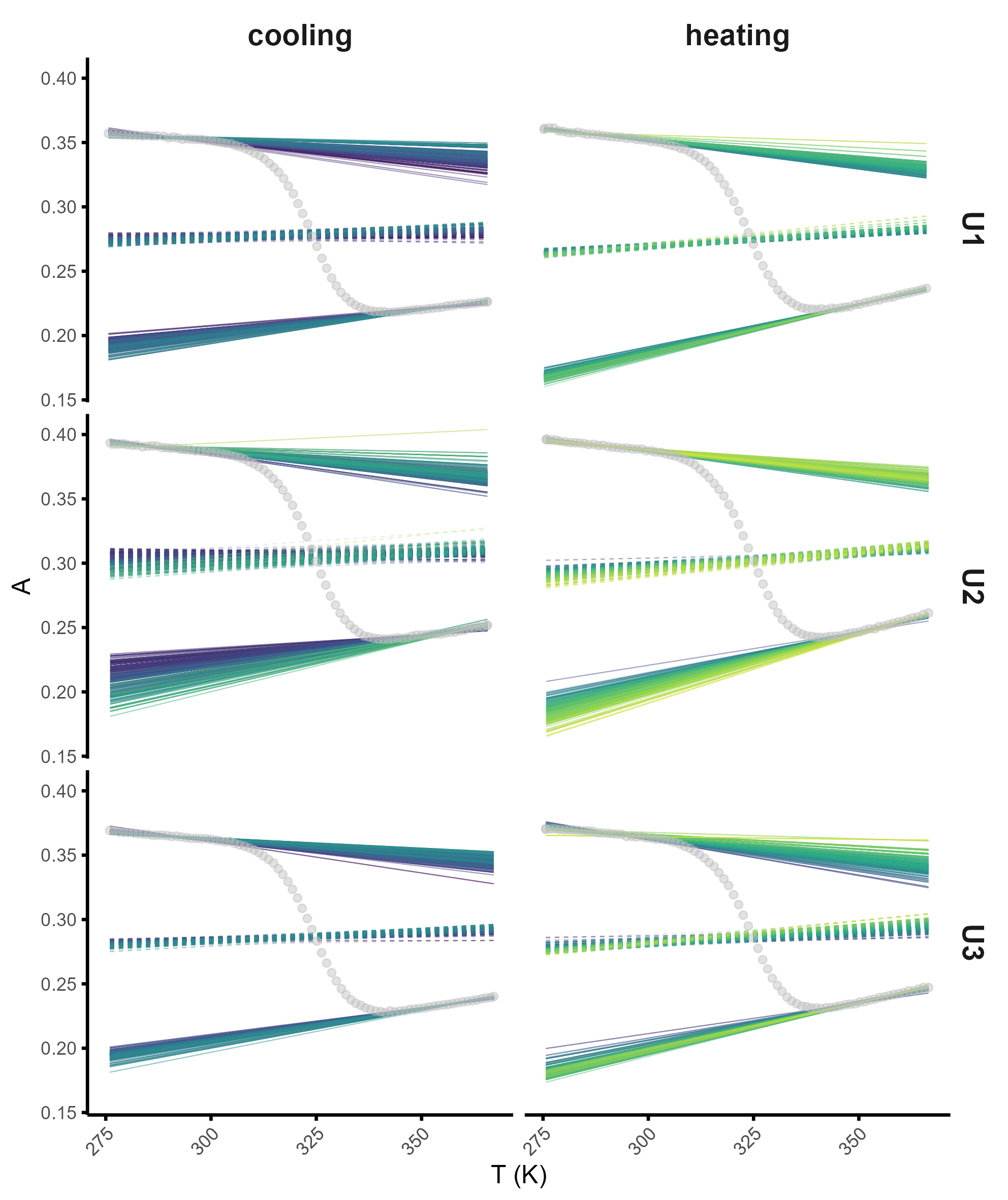
meltR allows high-throughput and robust Tm determination
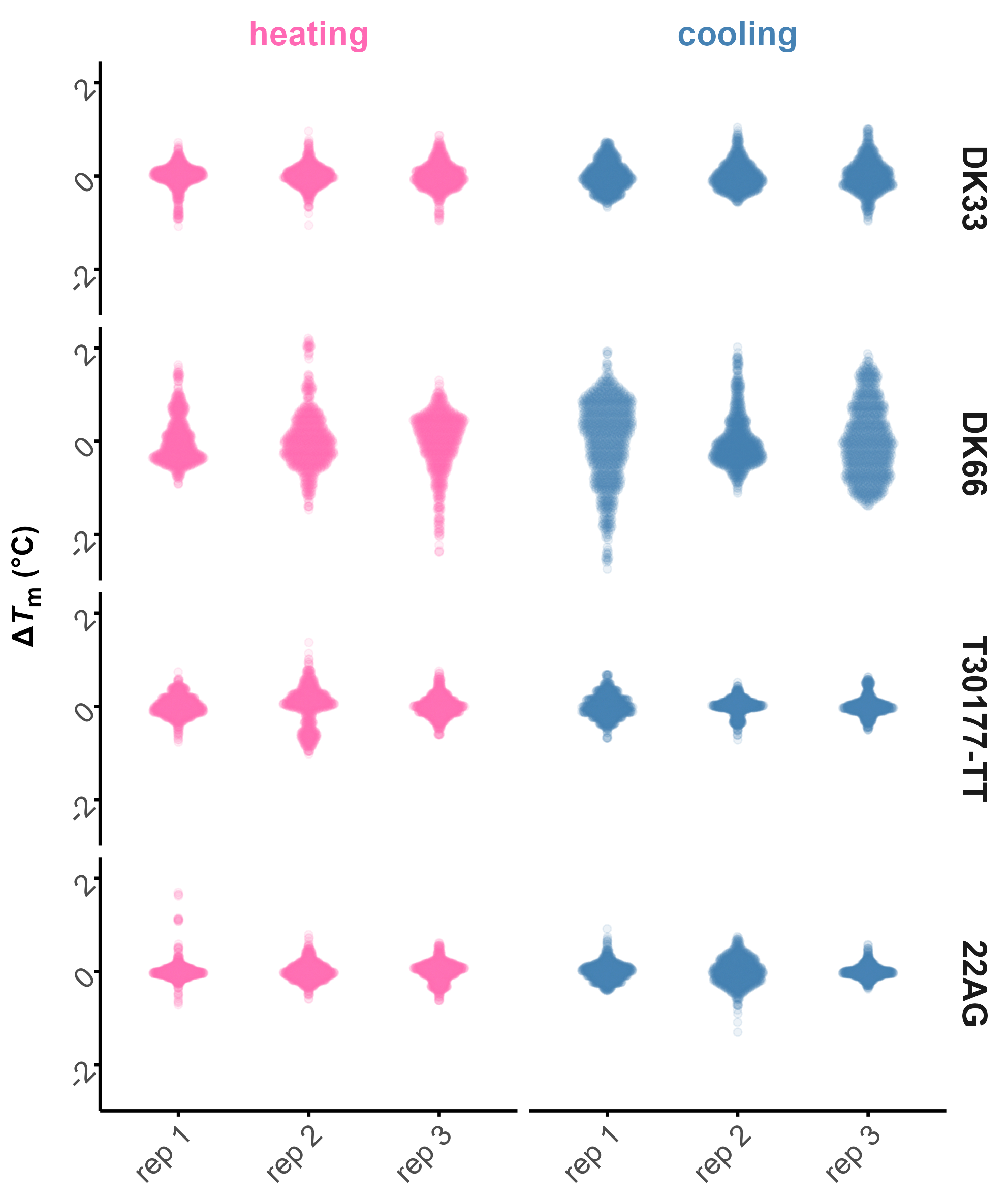
SEC allows robust stoichiometry determination
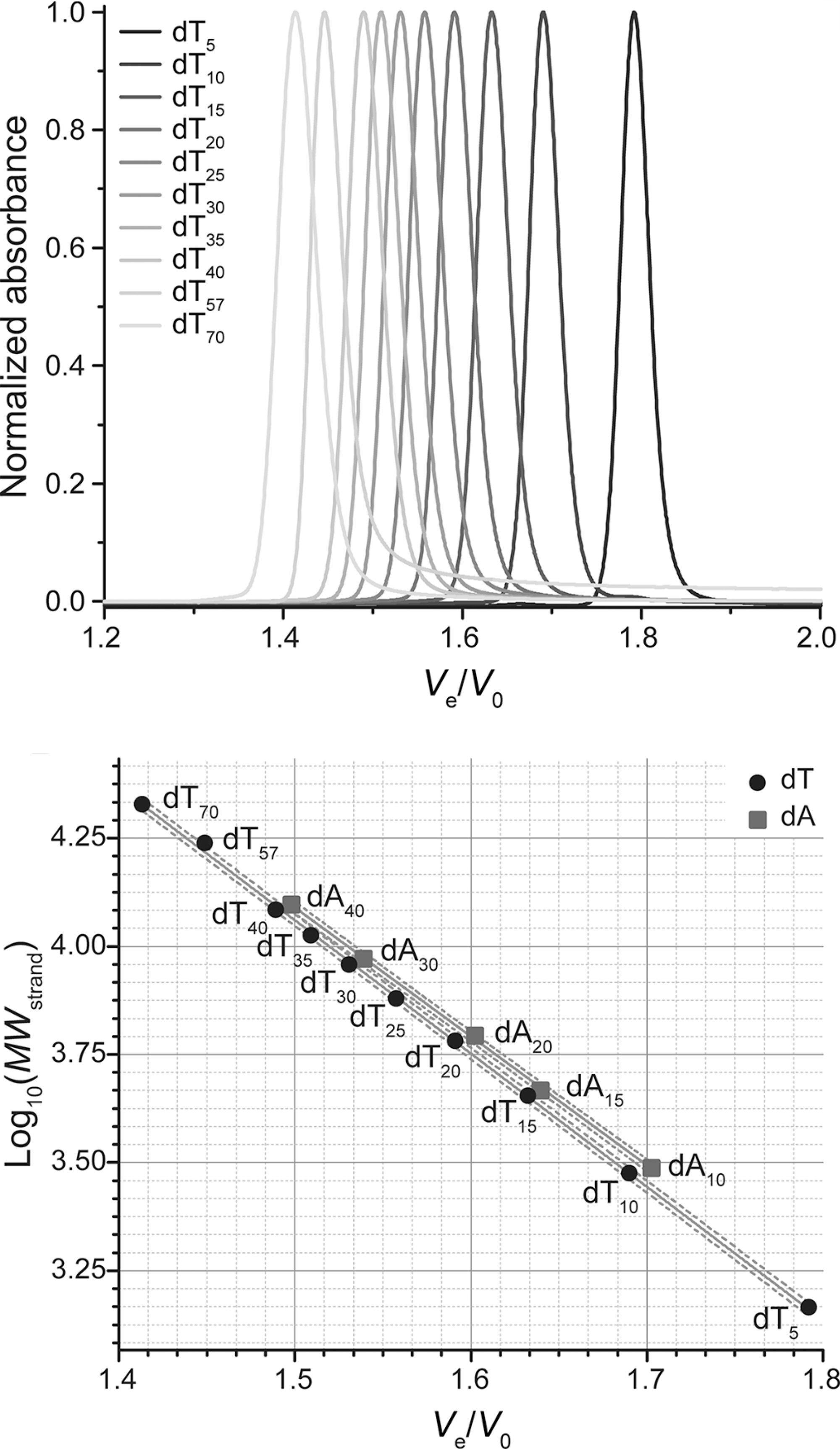
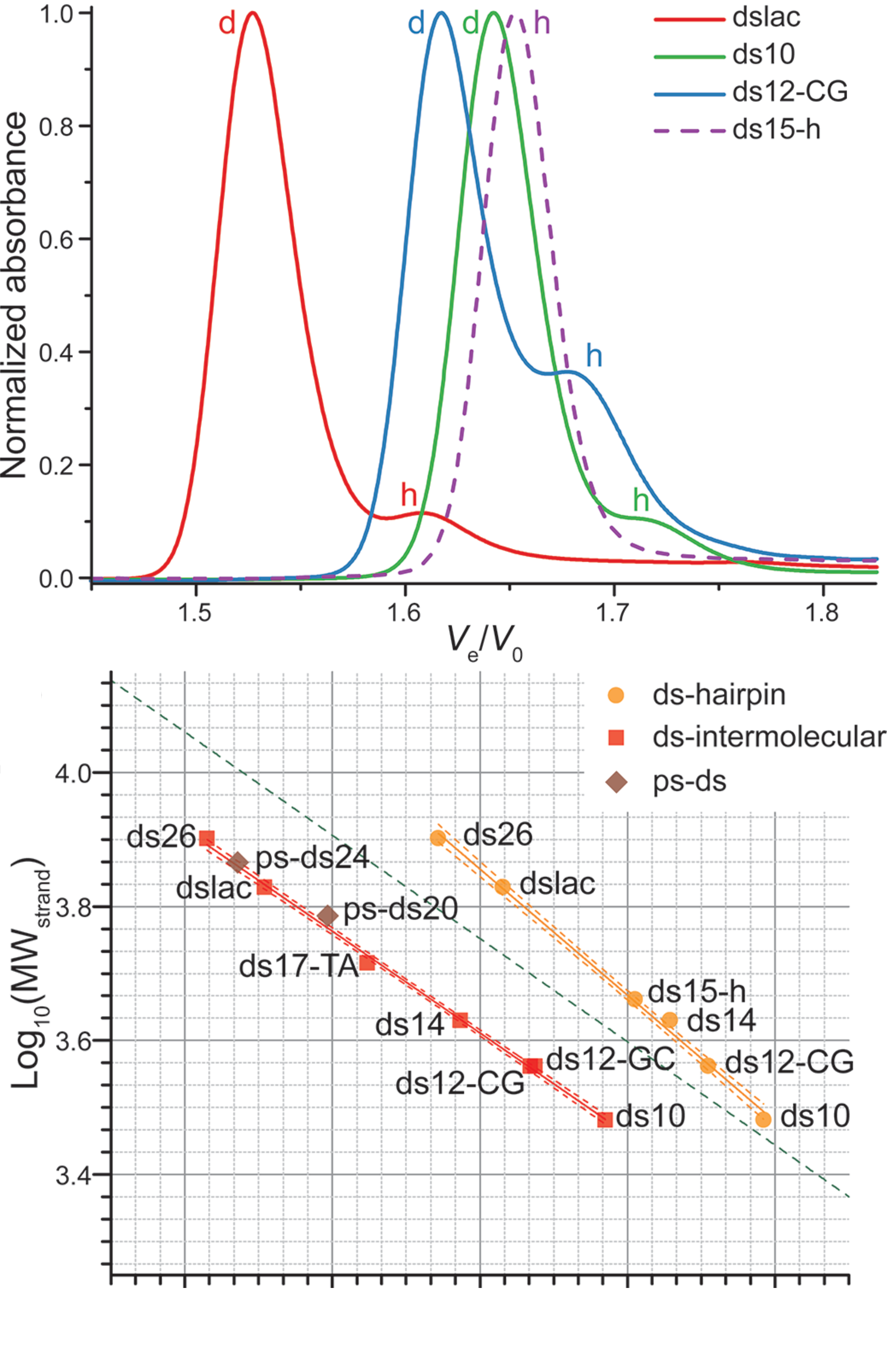
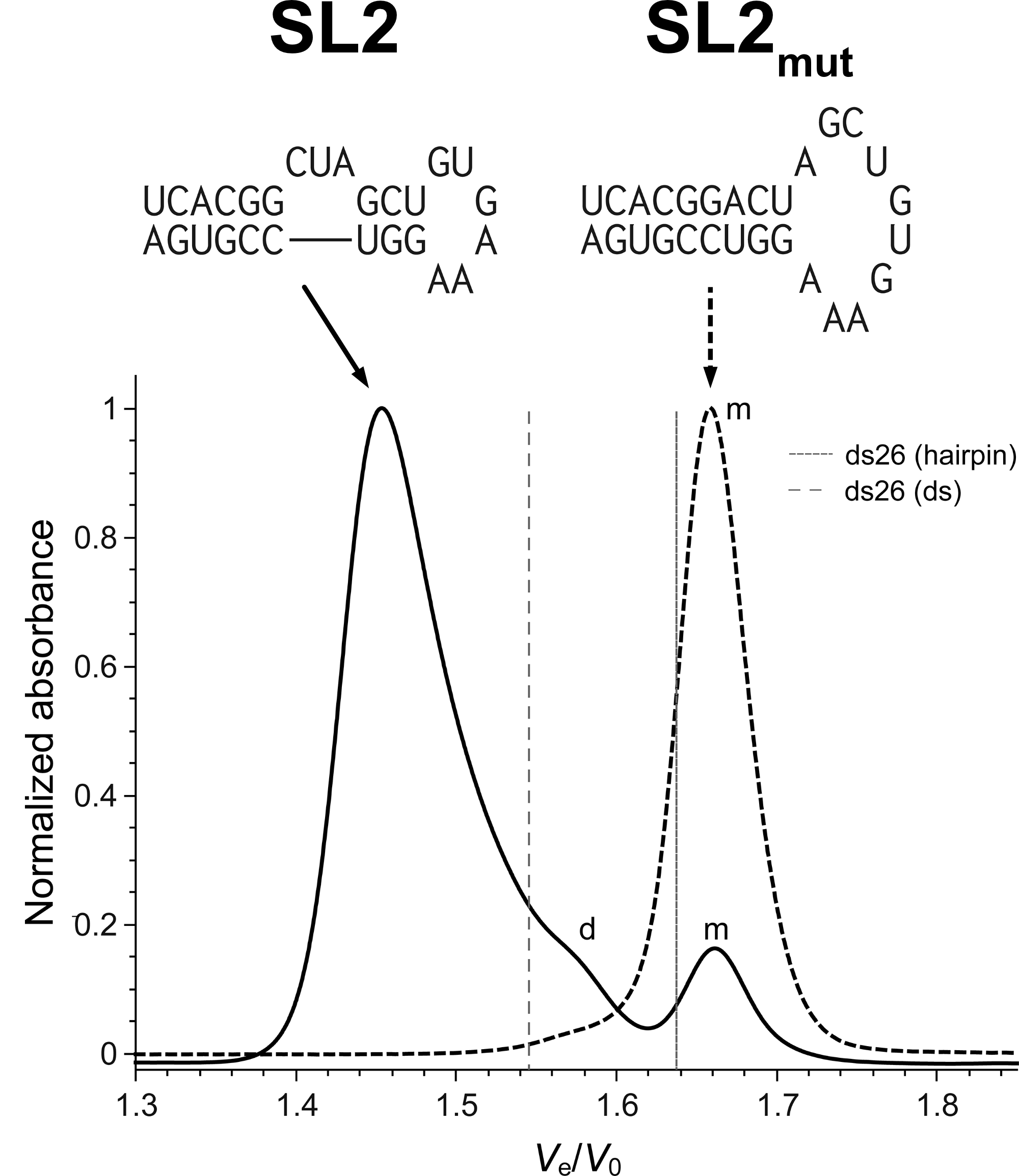

Largy, E. et Mergny, J.-L. Nucleic Acids Res., 2014, 42, e149.
SEC allows robust stoichiometry determination
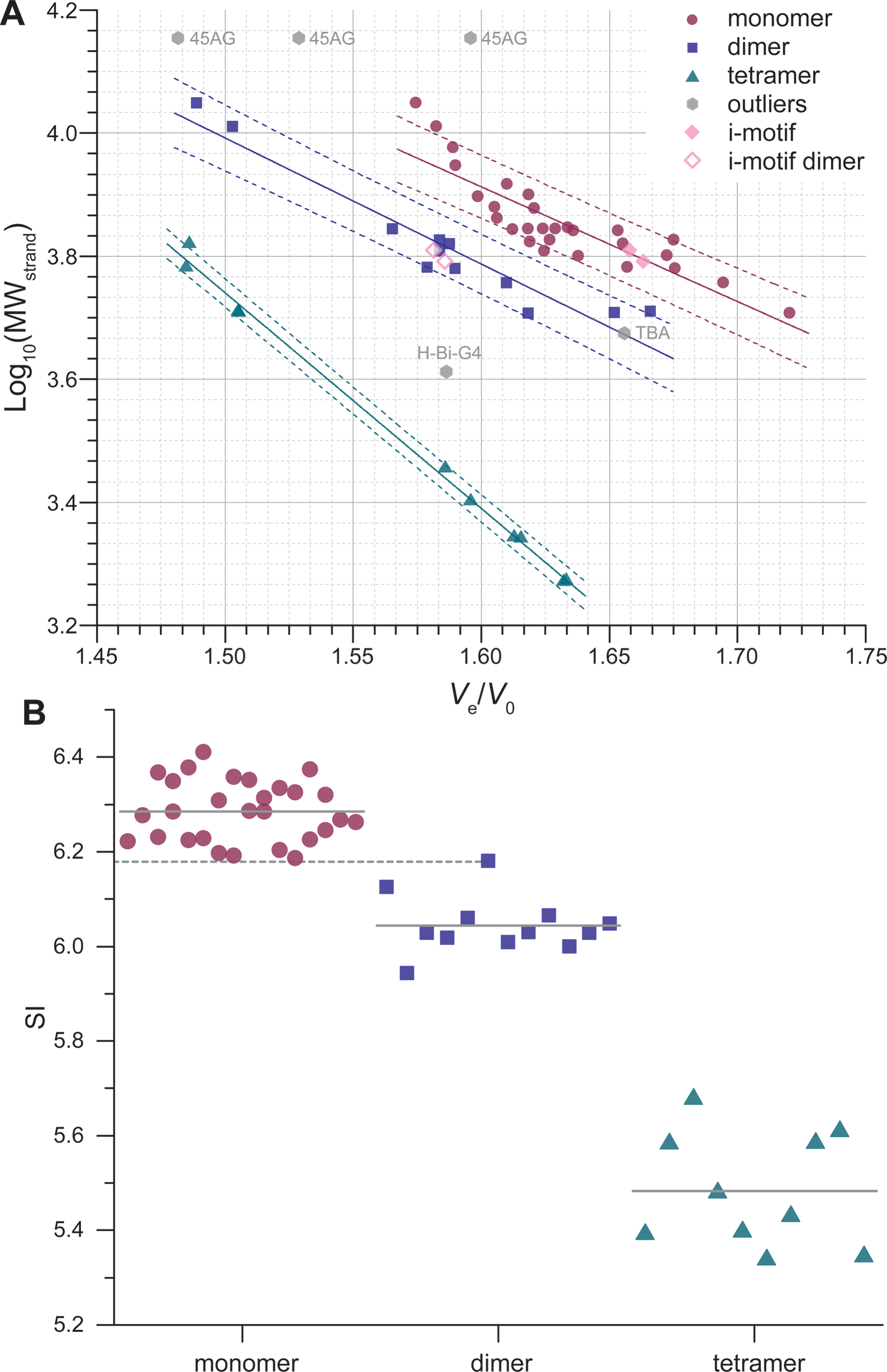
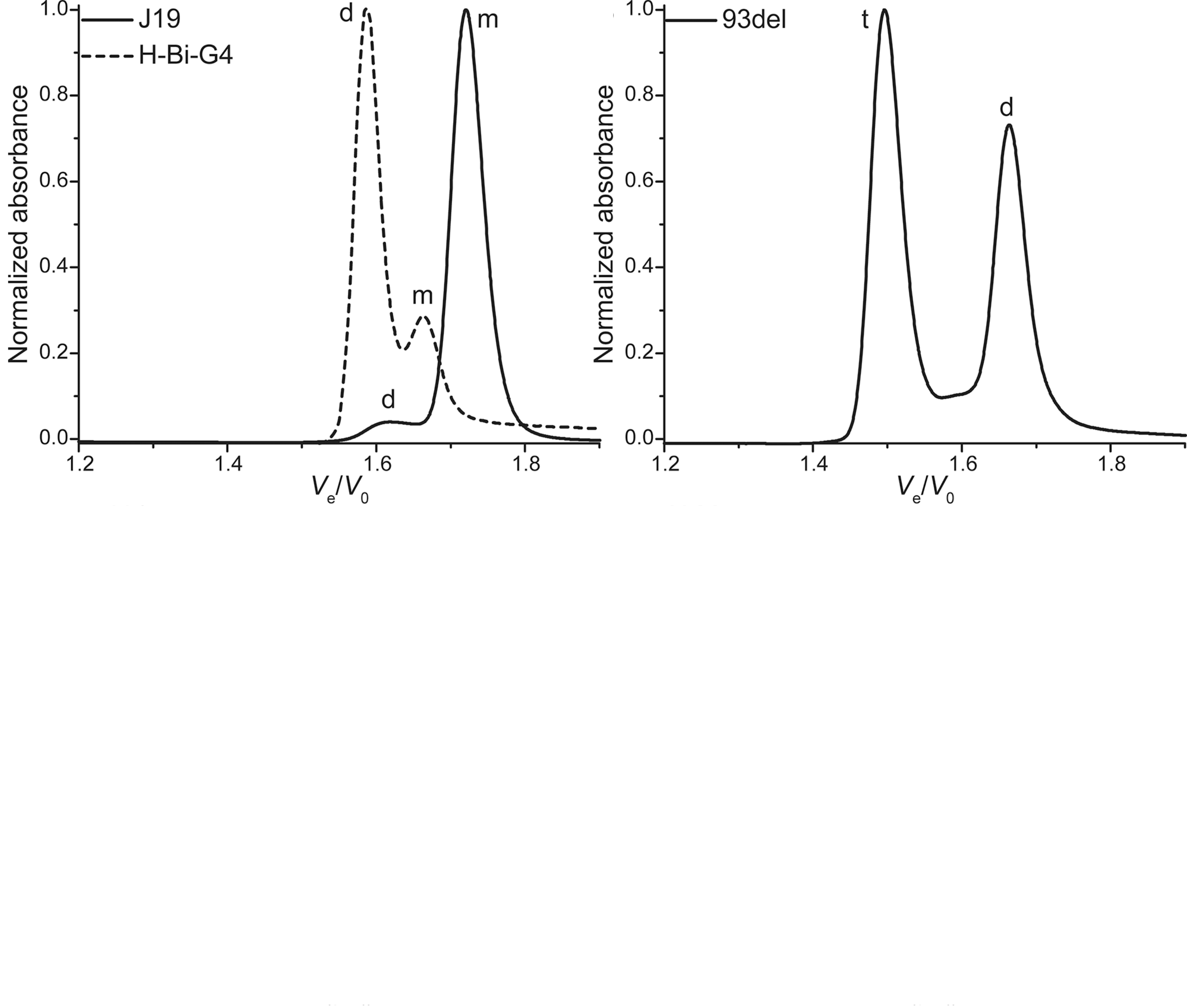
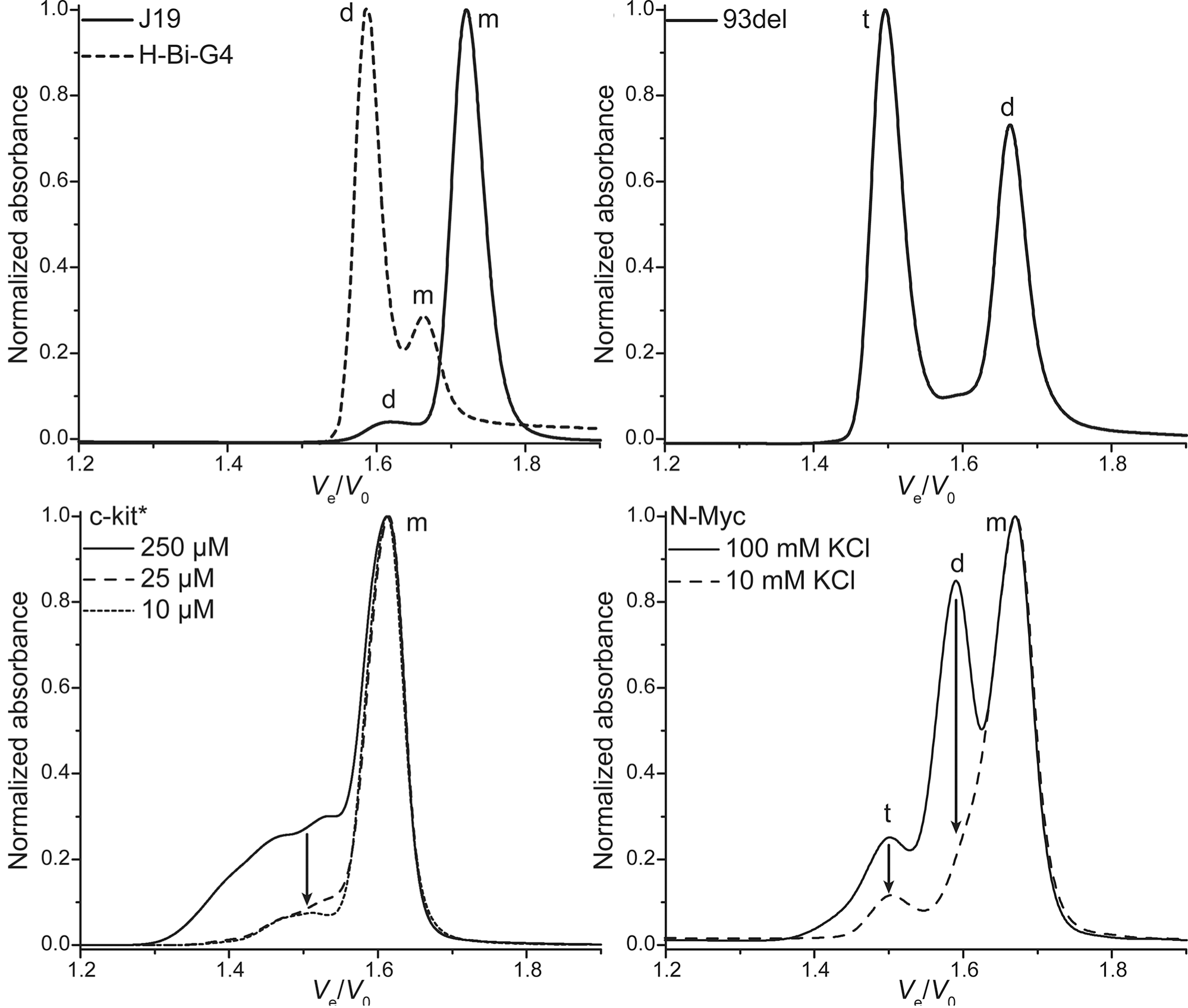
Largy, E. et Mergny, J.-L. Nucleic Acids Res., 2014, 42, e149.

Going further with native MS
Native MS probes non-covalent interactions
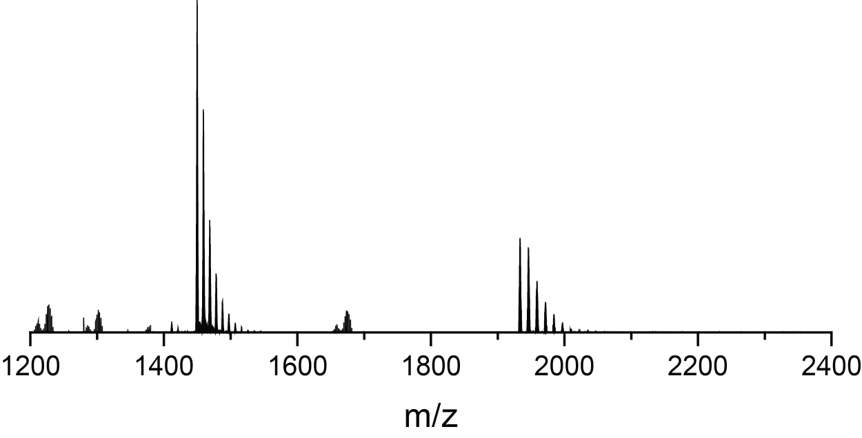
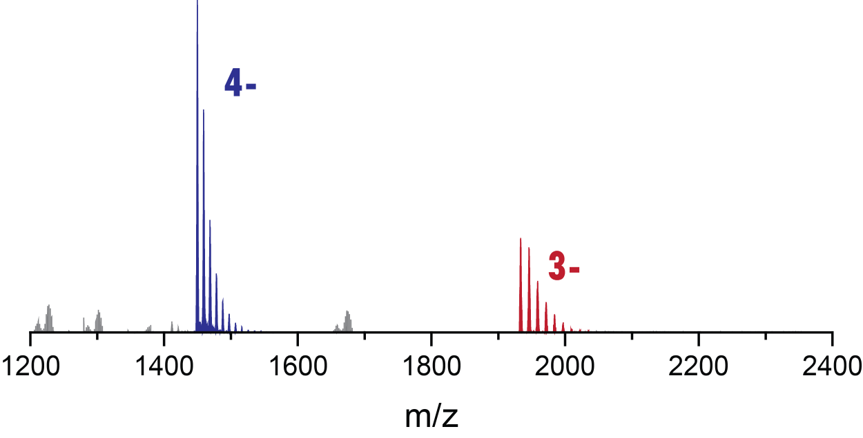
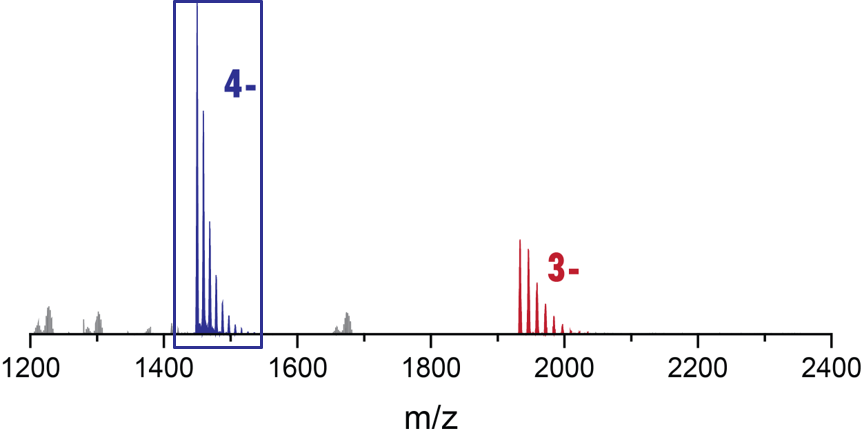
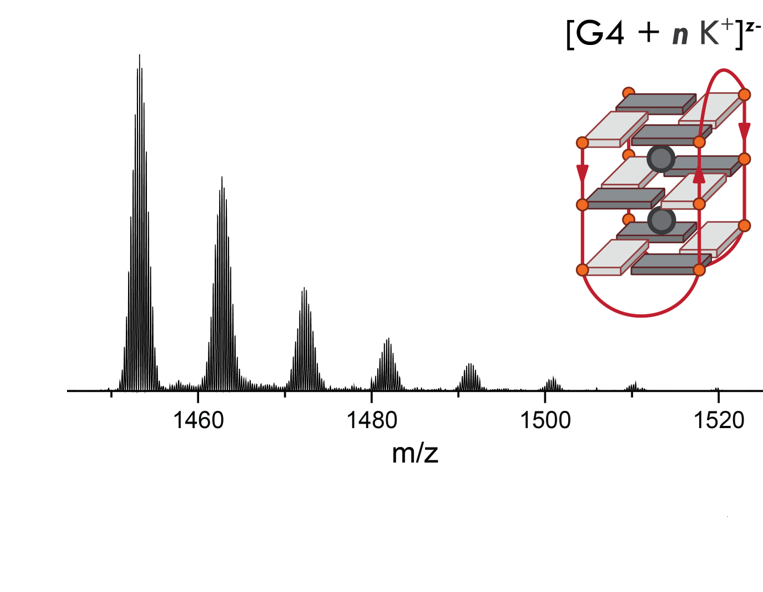
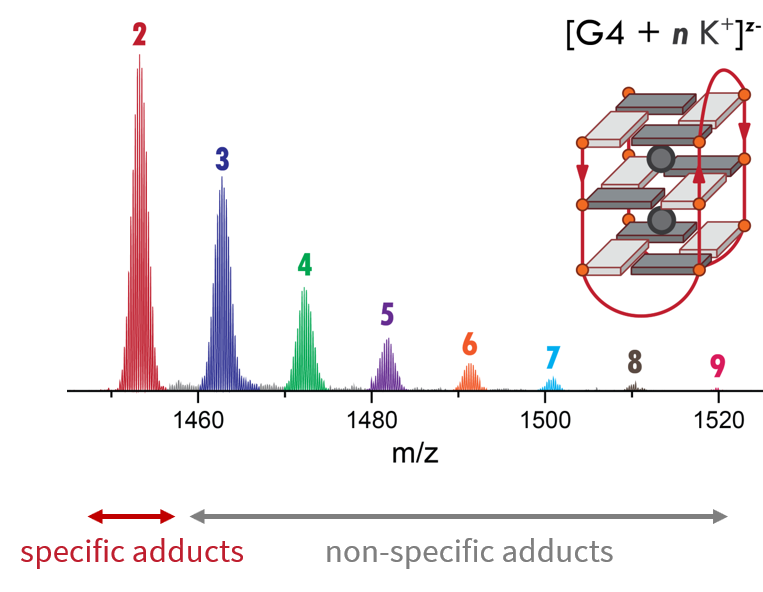
Adapting to a potassium-deficient buffer
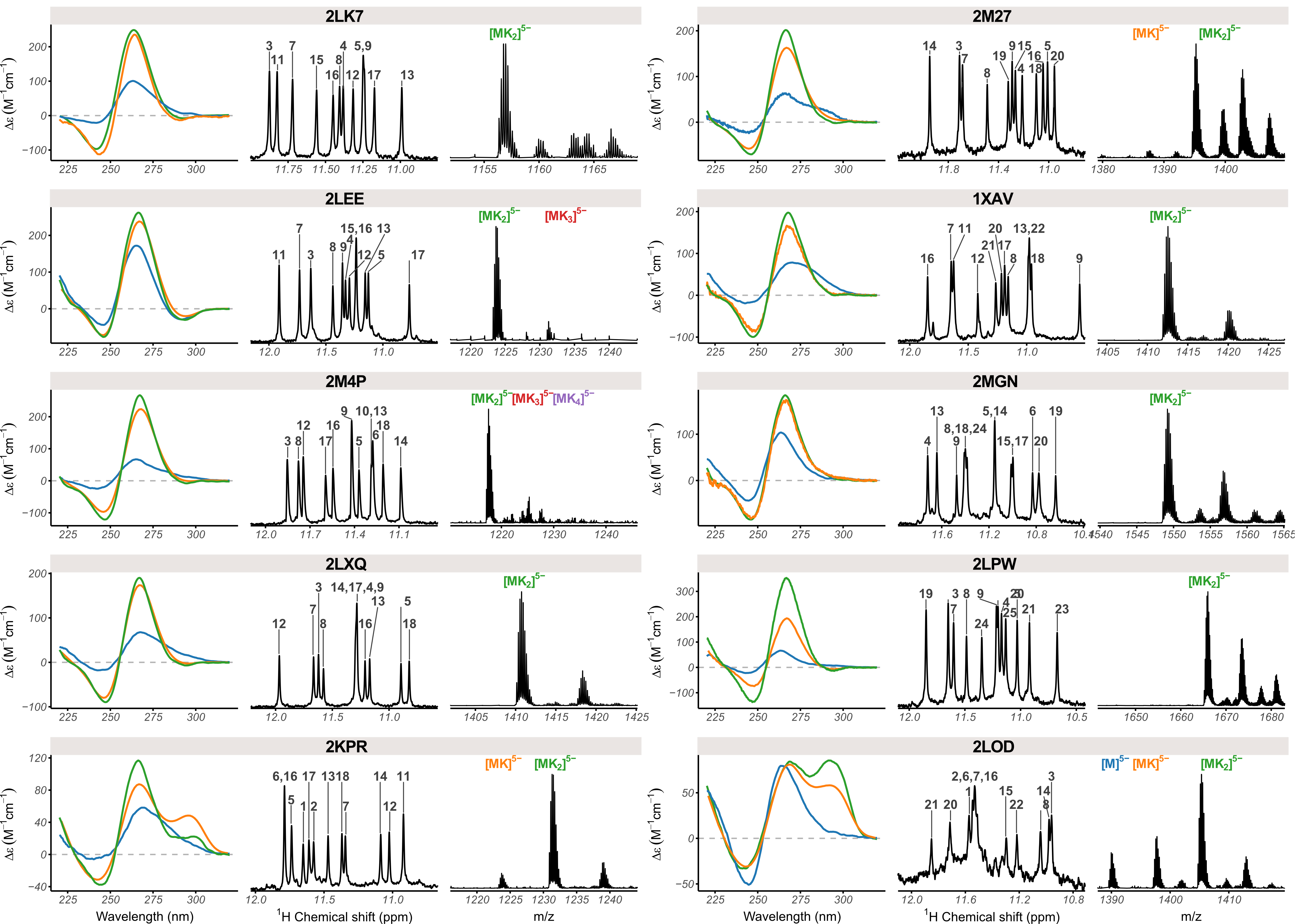

ericlarg4.github.io/G4_database
Ghosh, Largy, Gabelica, Nucleic Acids Res., 2021, 49, 2333
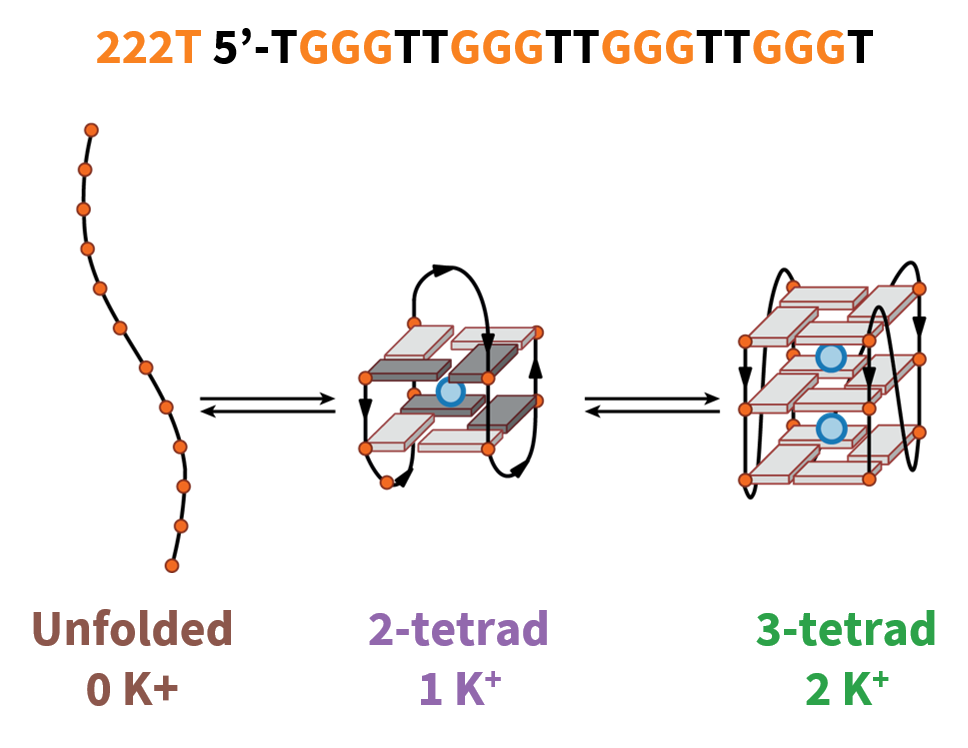
G4 conformations can be tuned with cations
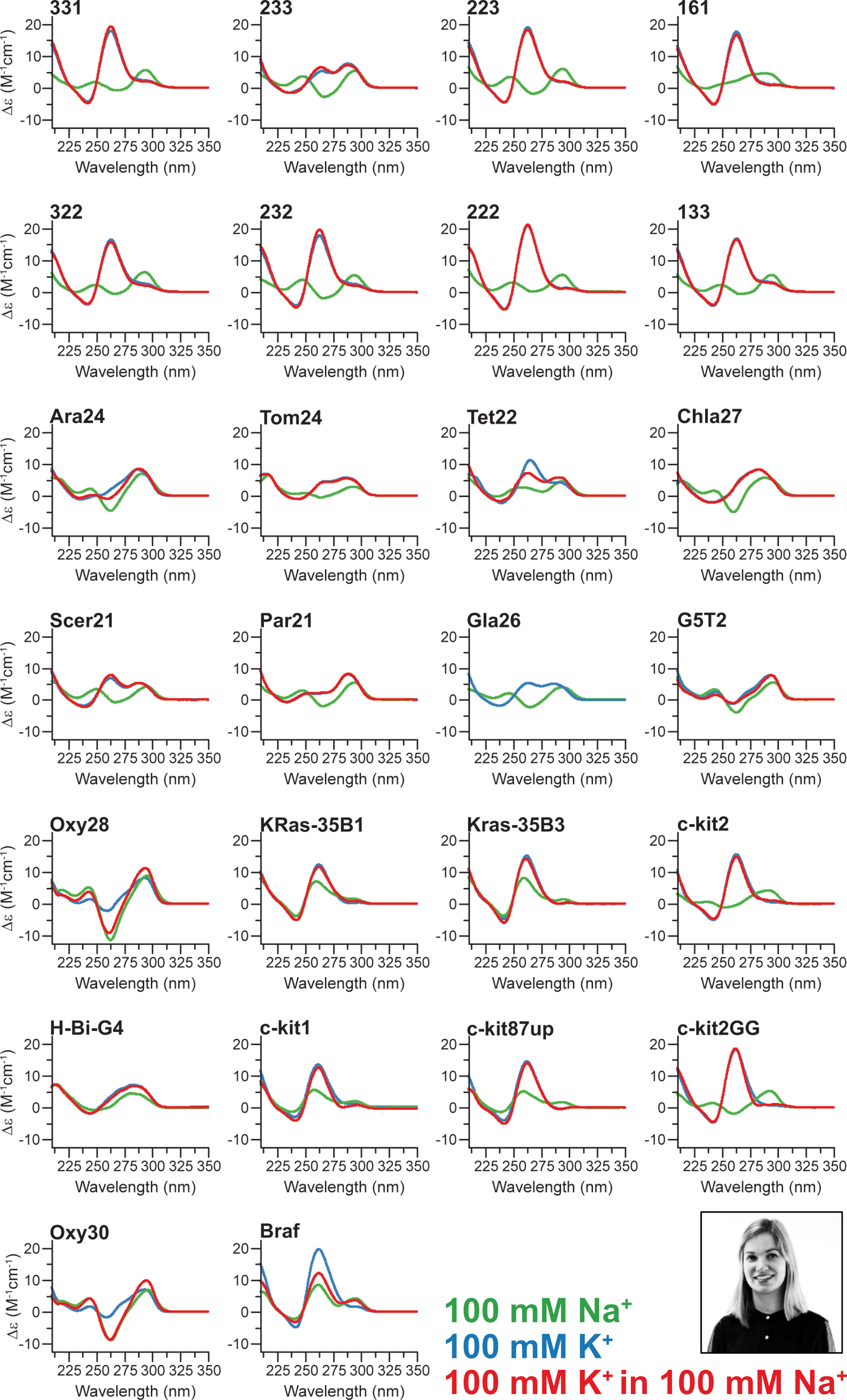
222T: TGGGTTGGGTTGGGTTGGGT

Largy, E., Marchand, A., Amrane, S., Gabelica, V. et Mergny, J.-L. J. Am. Chem. Soc., 2016, 138, 2780
The interconversion occurs at low [KCl]
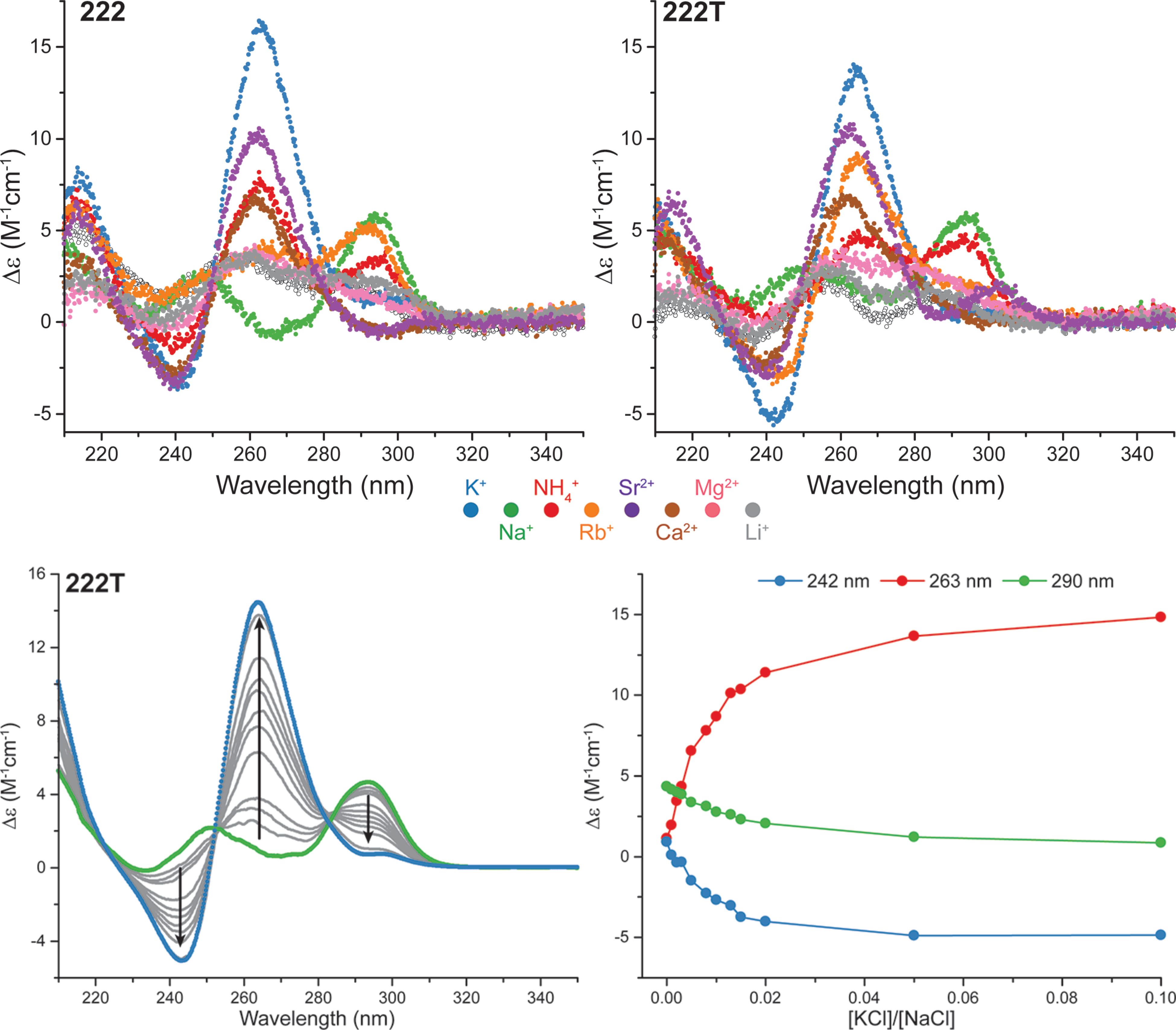
Largy, E., Marchand, A., Amrane, S., Gabelica, V. et Mergny, J.-L. J. Am. Chem. Soc., 2016, 138, 2780
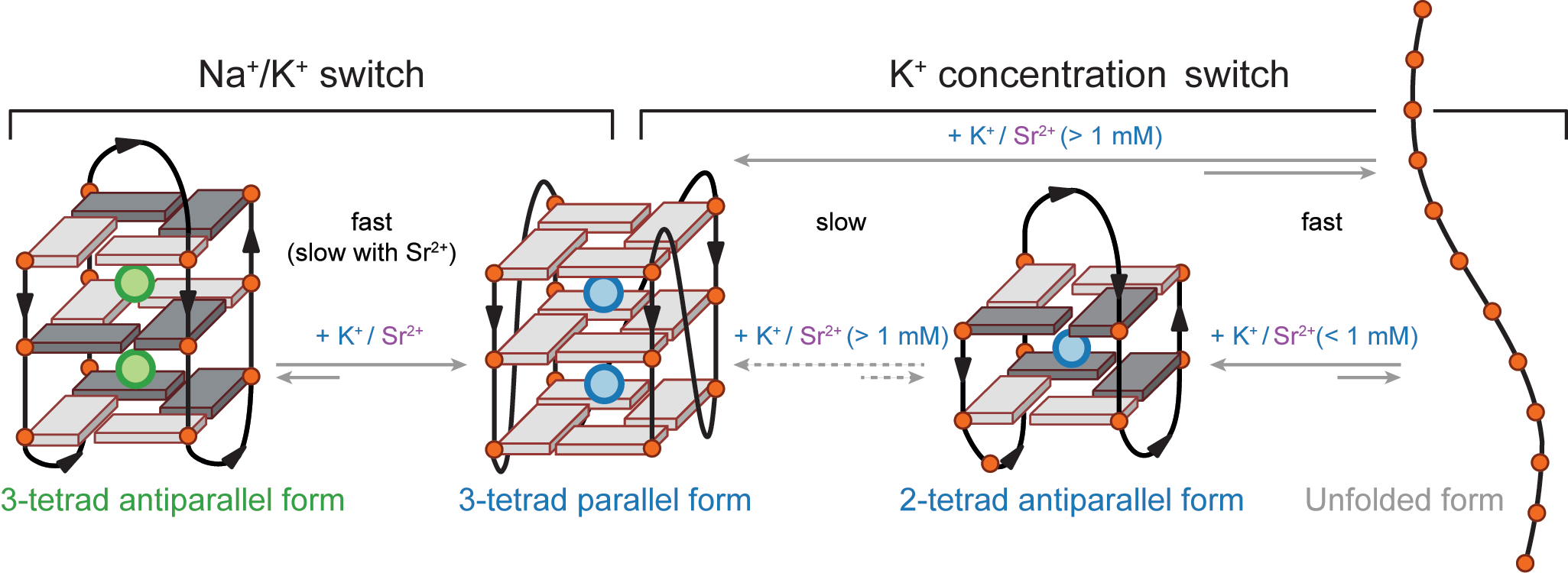
The interconversion is fast
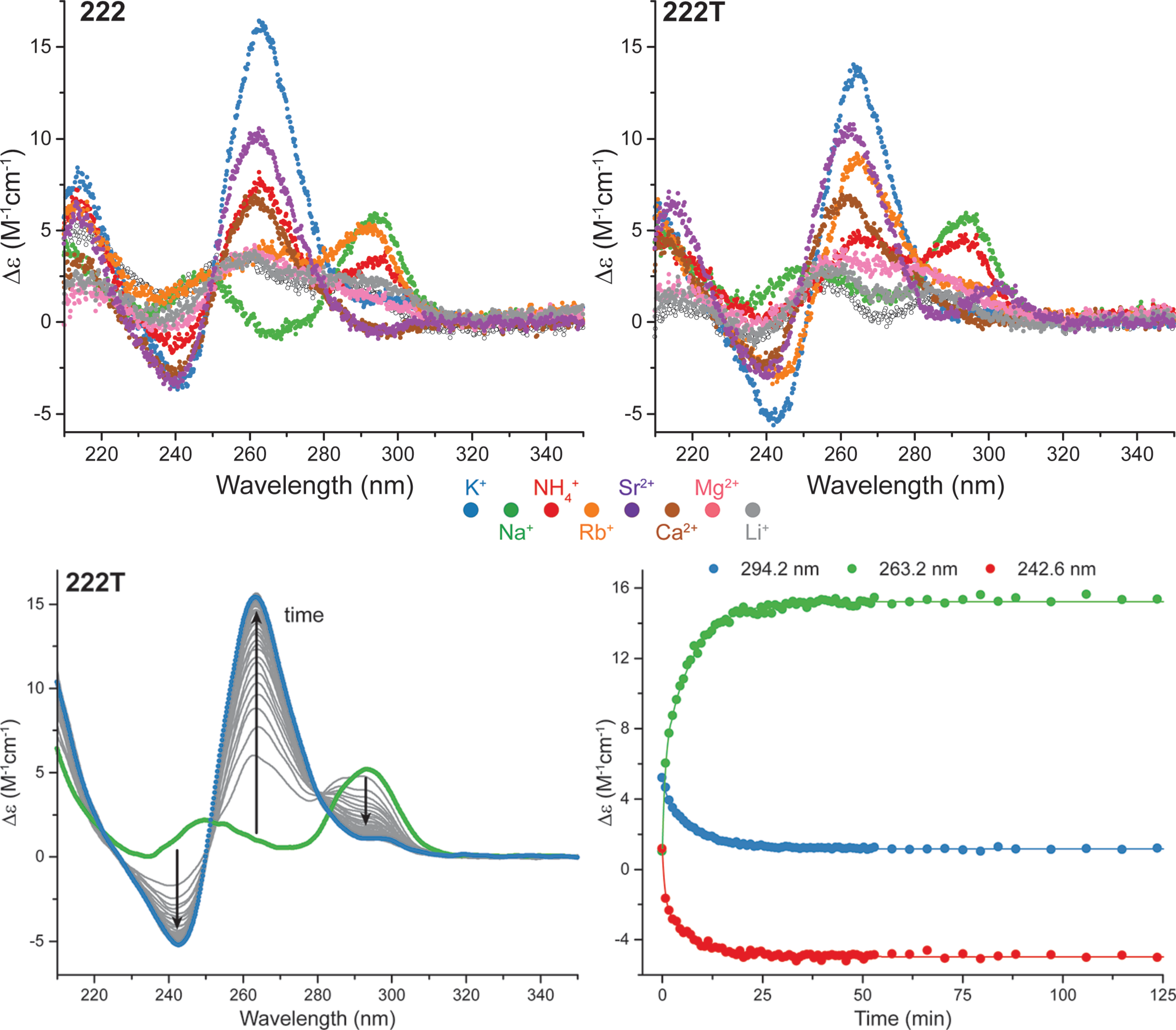
Largy, E., Marchand, A., Amrane, S., Gabelica, V. et Mergny, J.-L. J. Am. Chem. Soc., 2016, 138, 2780
The interconversion arises from large ΔTm
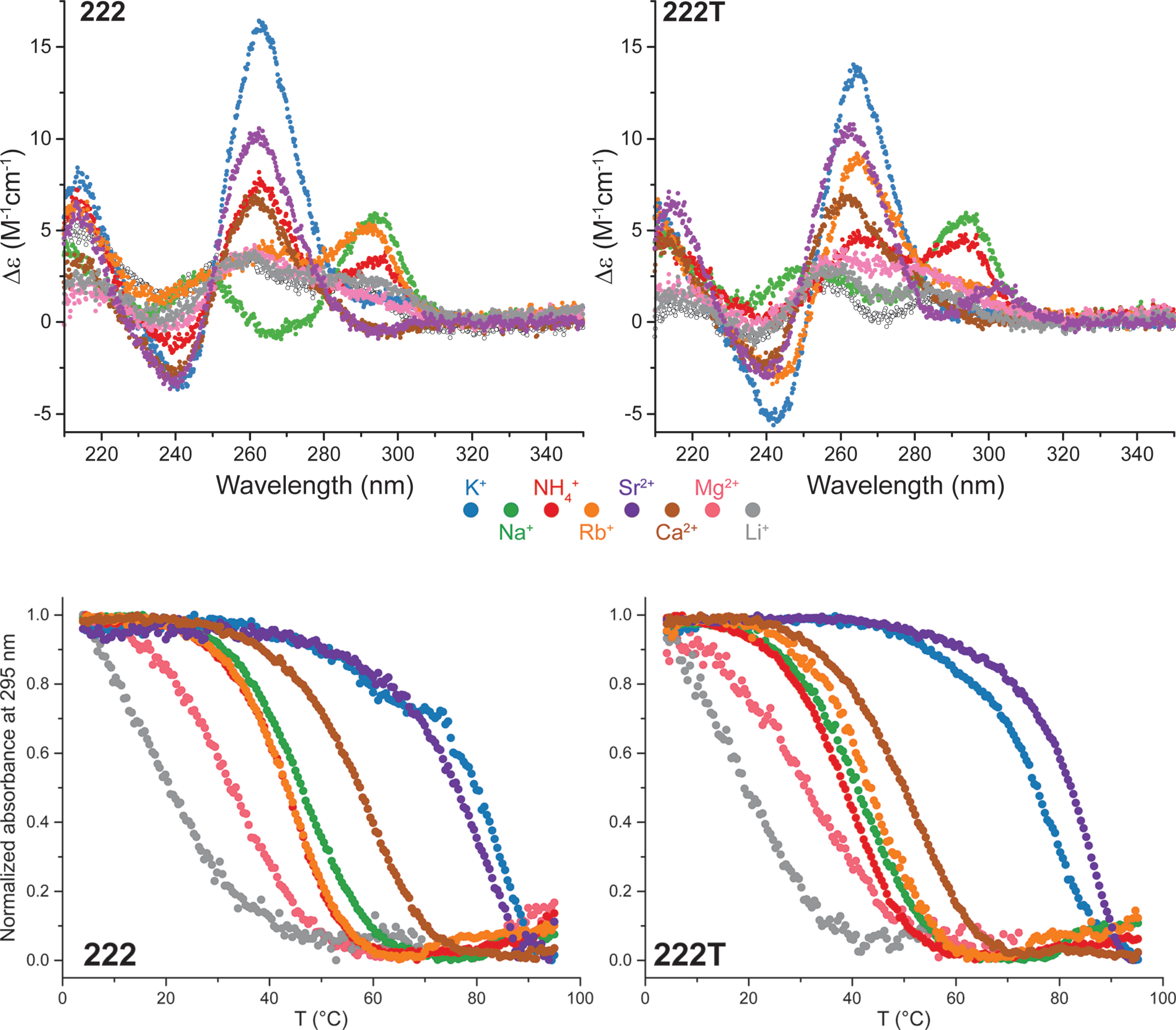
Largy, E., Marchand, A., Amrane, S., Gabelica, V. et Mergny, J.-L. J. Am. Chem. Soc., 2016, 138, 2780
The interconversion can be monitored by UV
Largy, E., Marchand, A., Amrane, S., Gabelica, V. et Mergny, J.-L. J. Am. Chem. Soc., 2016, 138, 2780
The interconversion can be monitored by nMS
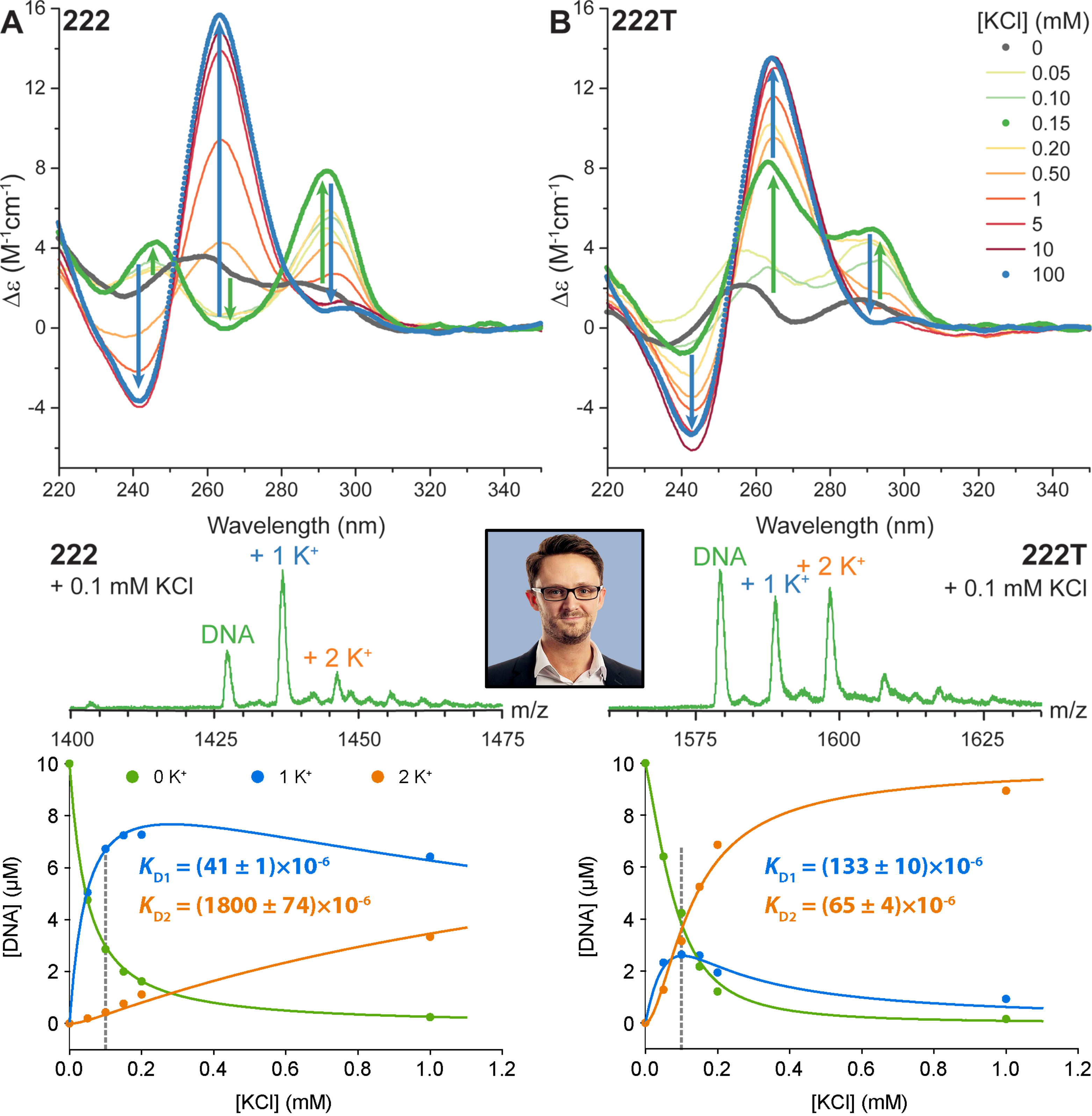

Largy, E., Marchand, A., Amrane, S., Gabelica, V. et Mergny, J.-L. J. Am. Chem. Soc., 2016, 138, 2780
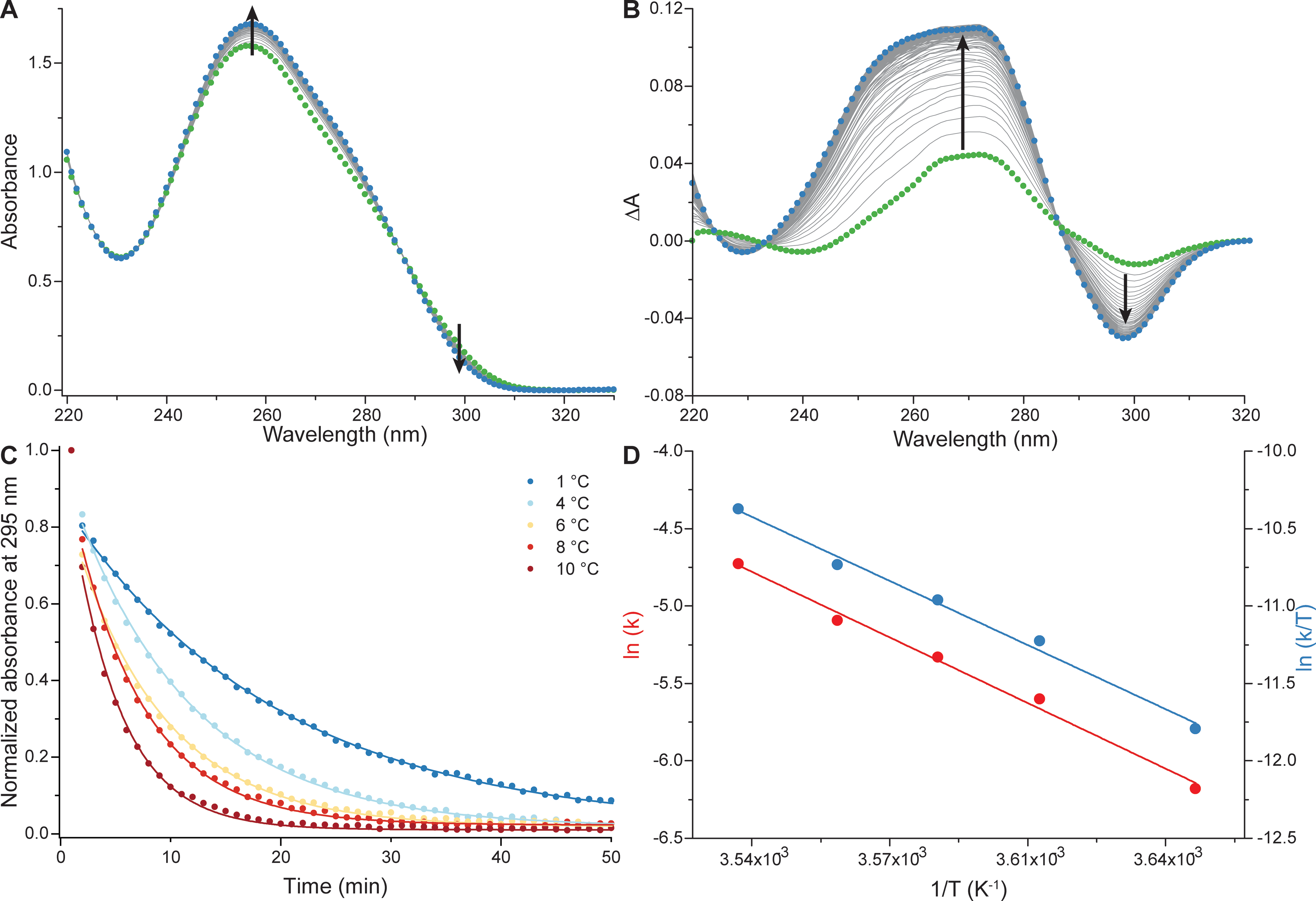
The combination of methods allows thermodynamic
and kinetic characterization of complex system
Largy, E., Marchand, A., Amrane, S., Gabelica, V. et Mergny, J.-L. J. Am. Chem. Soc., 2016, 138, 2780
Combining methods to decipher ligand interactions
G4 can be targeted by small molecules

Most G4 ligands are \(\pi\)-stacking on external tetrads
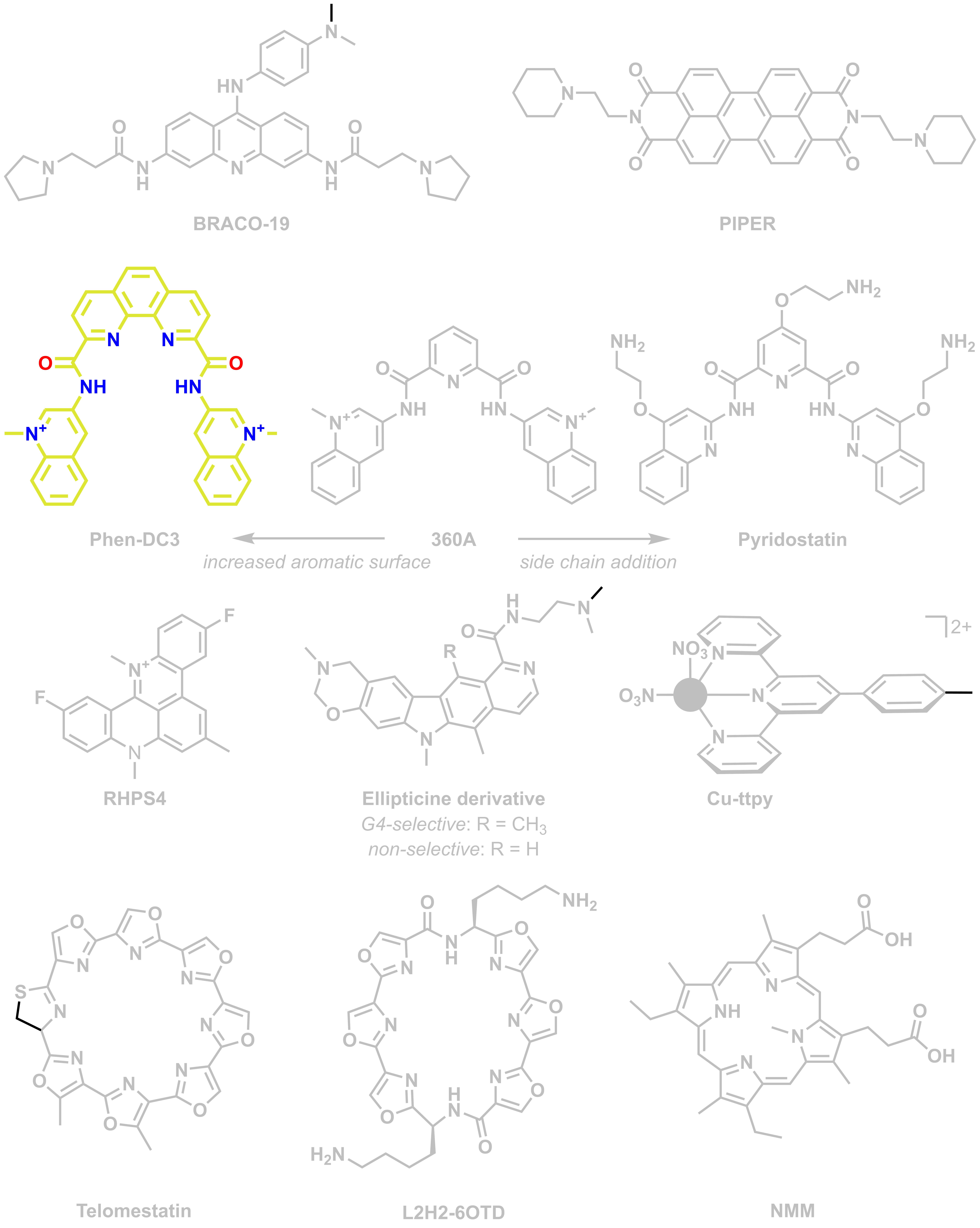
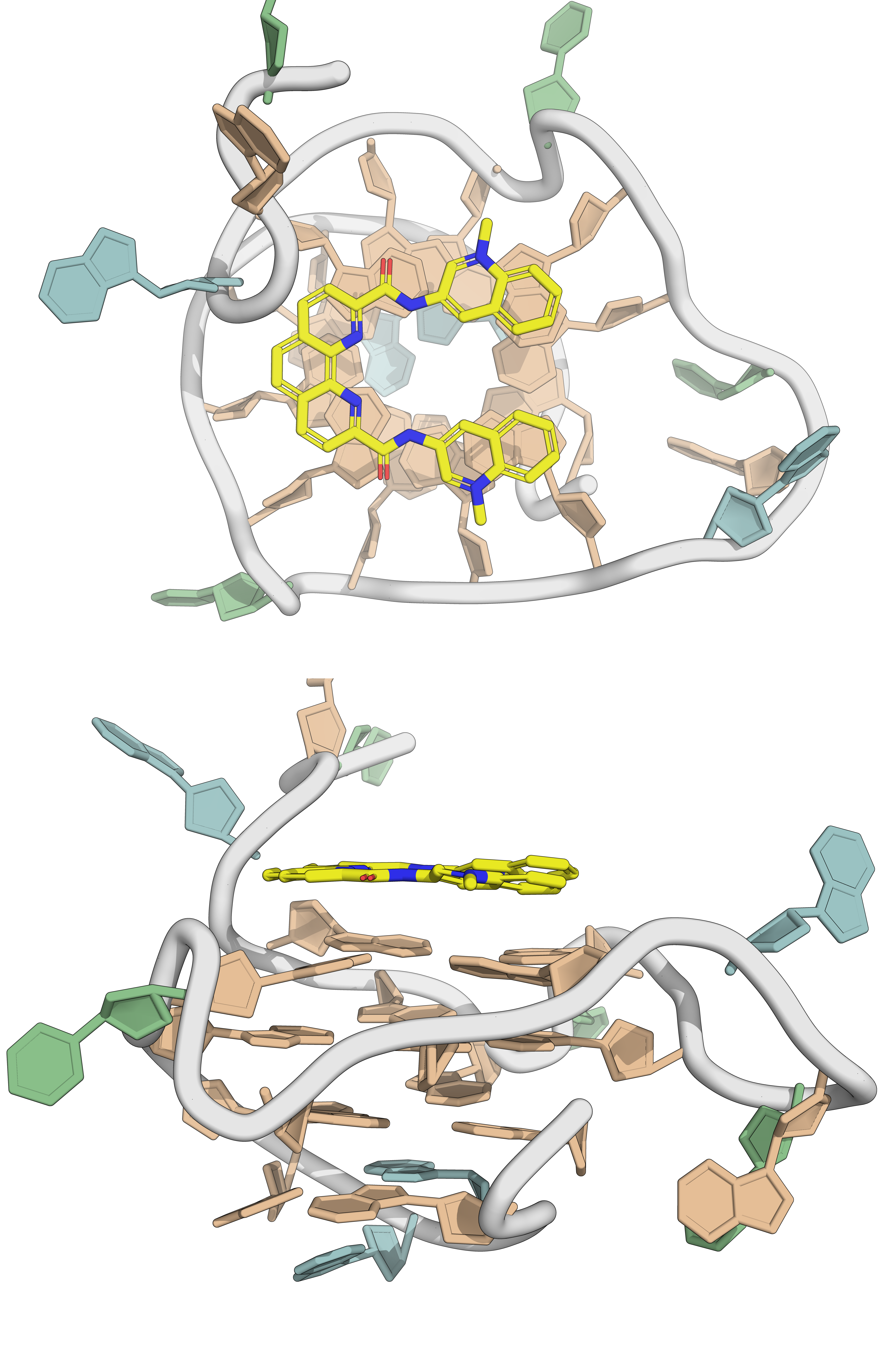
Largy et al., Chem. Rev. 2022, 122, 8, 7720
Sometimes luck does better than reason

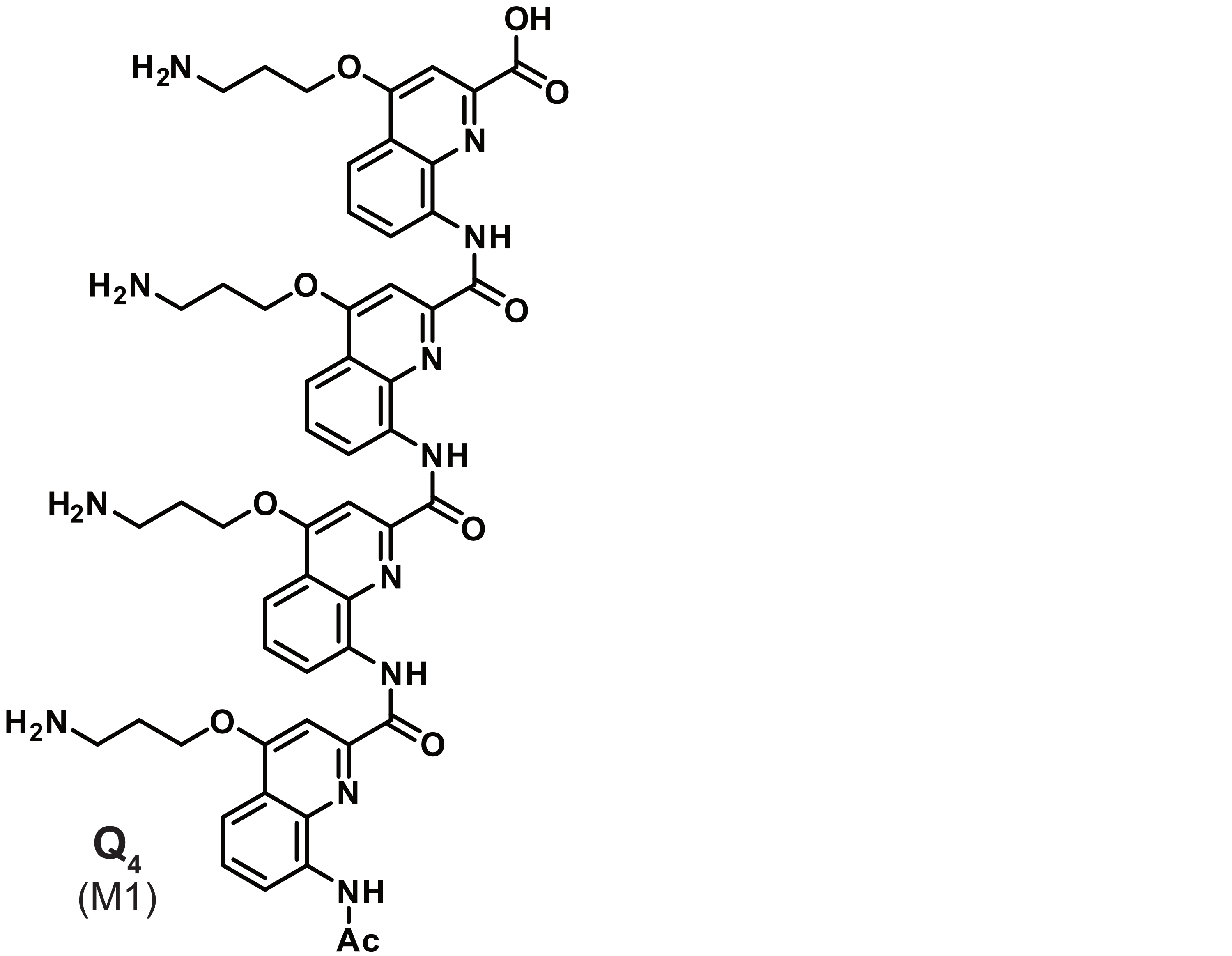
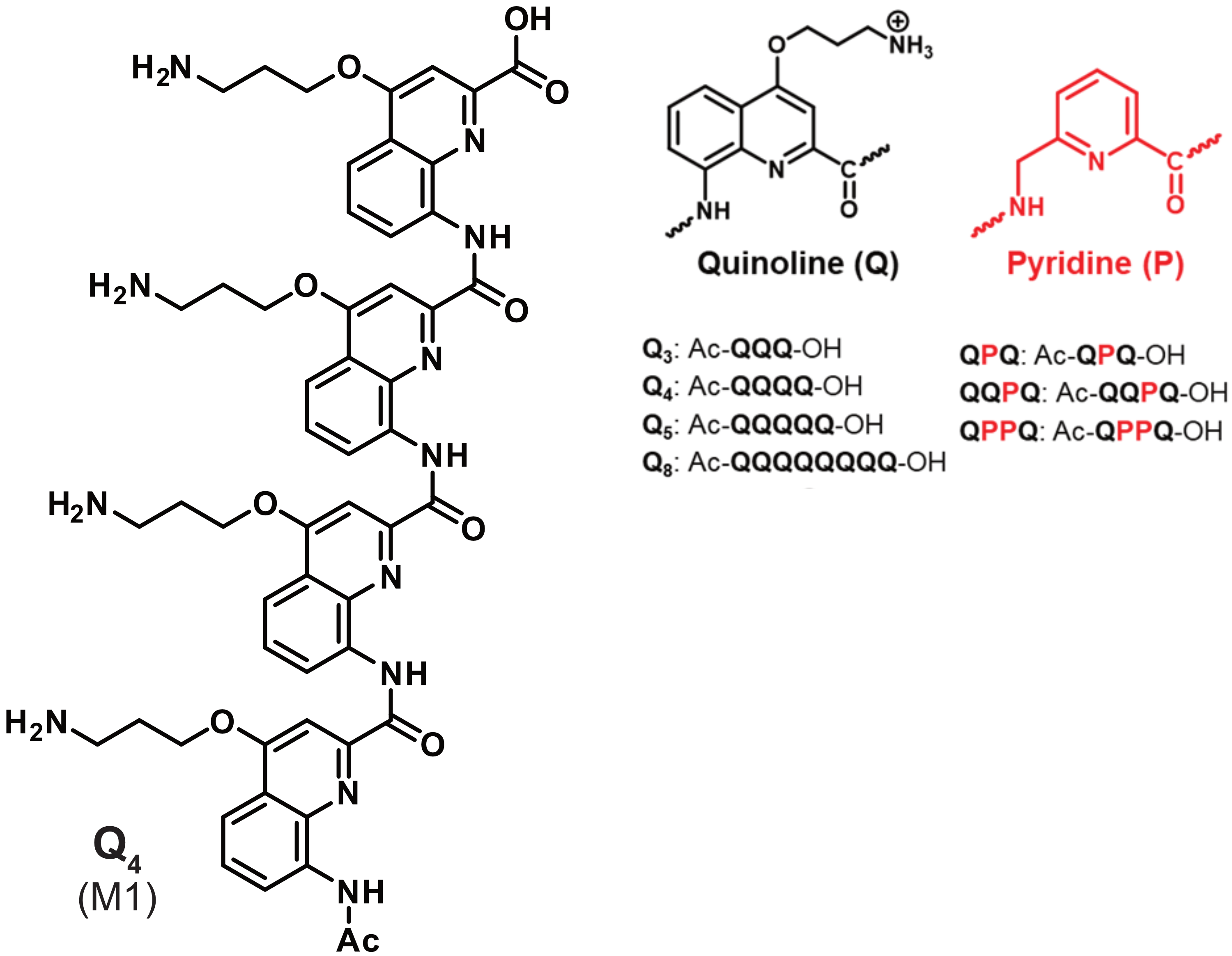
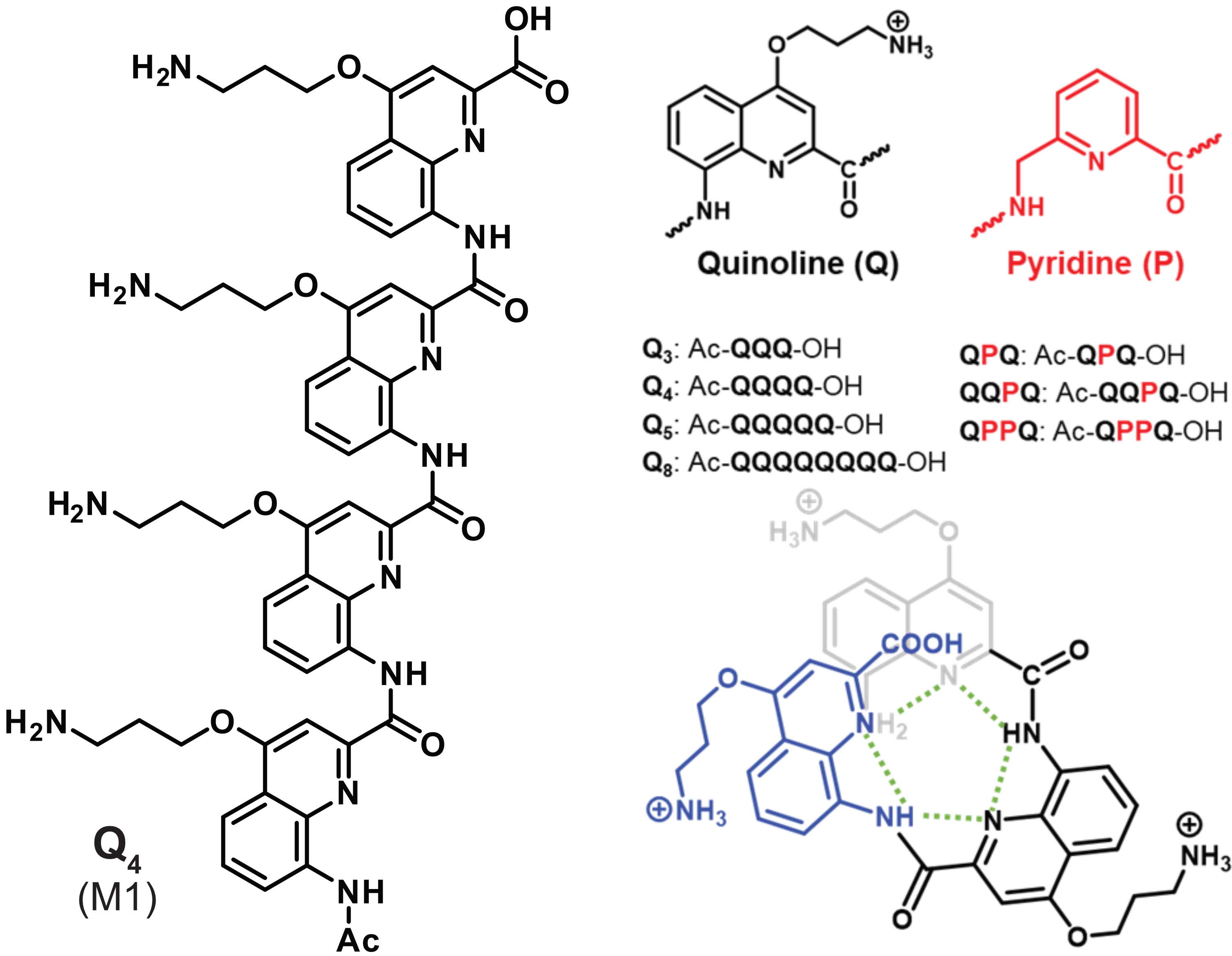
König, A., Laffilé, V., Thore, S., Mackereth, C., Yatsunyk, L., Ferrand, Y., Largy, E., Gabelica, V., Nucleic Acids Res., 2025, gkaf1365
Binding preference to parallel topology
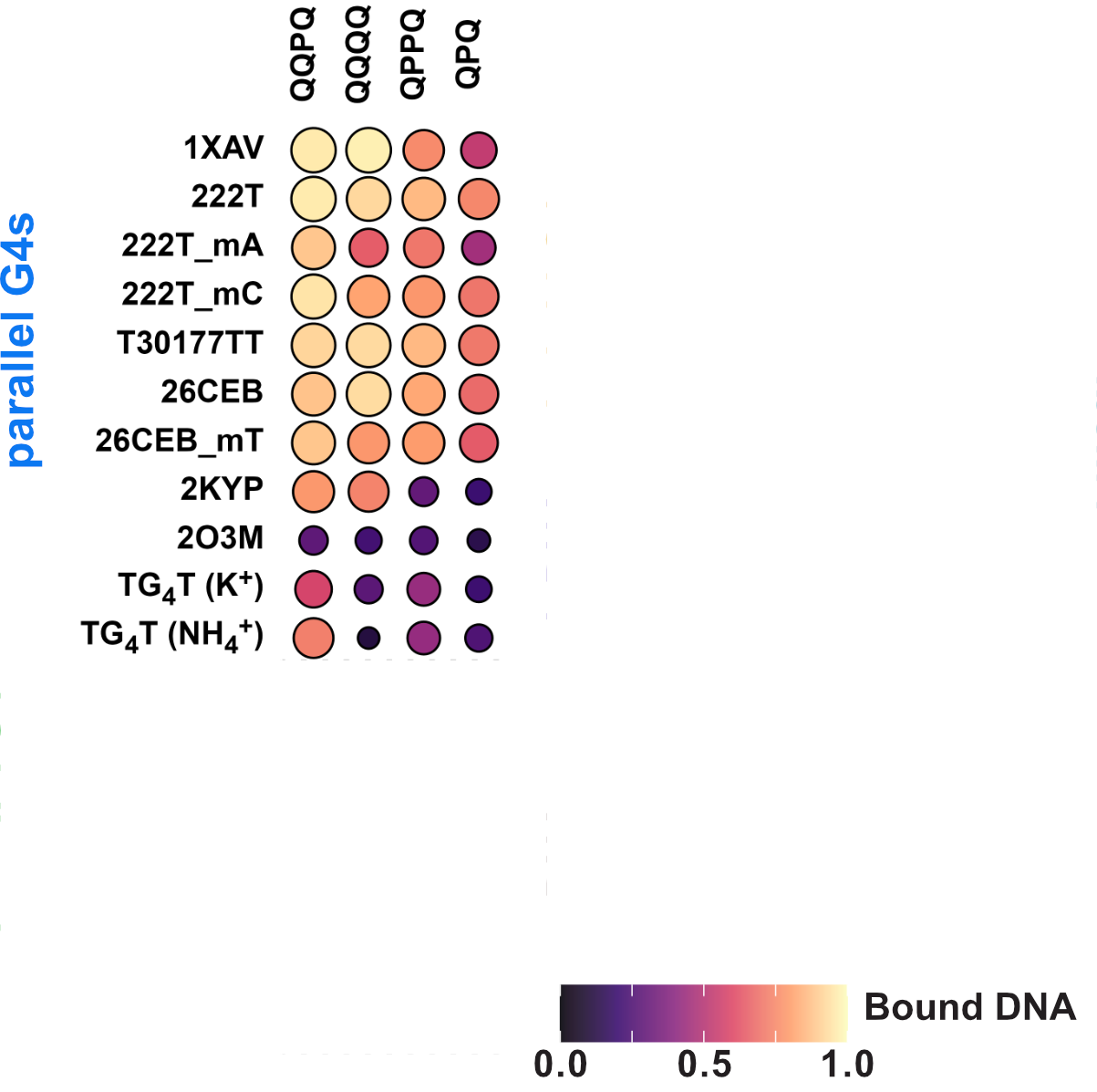
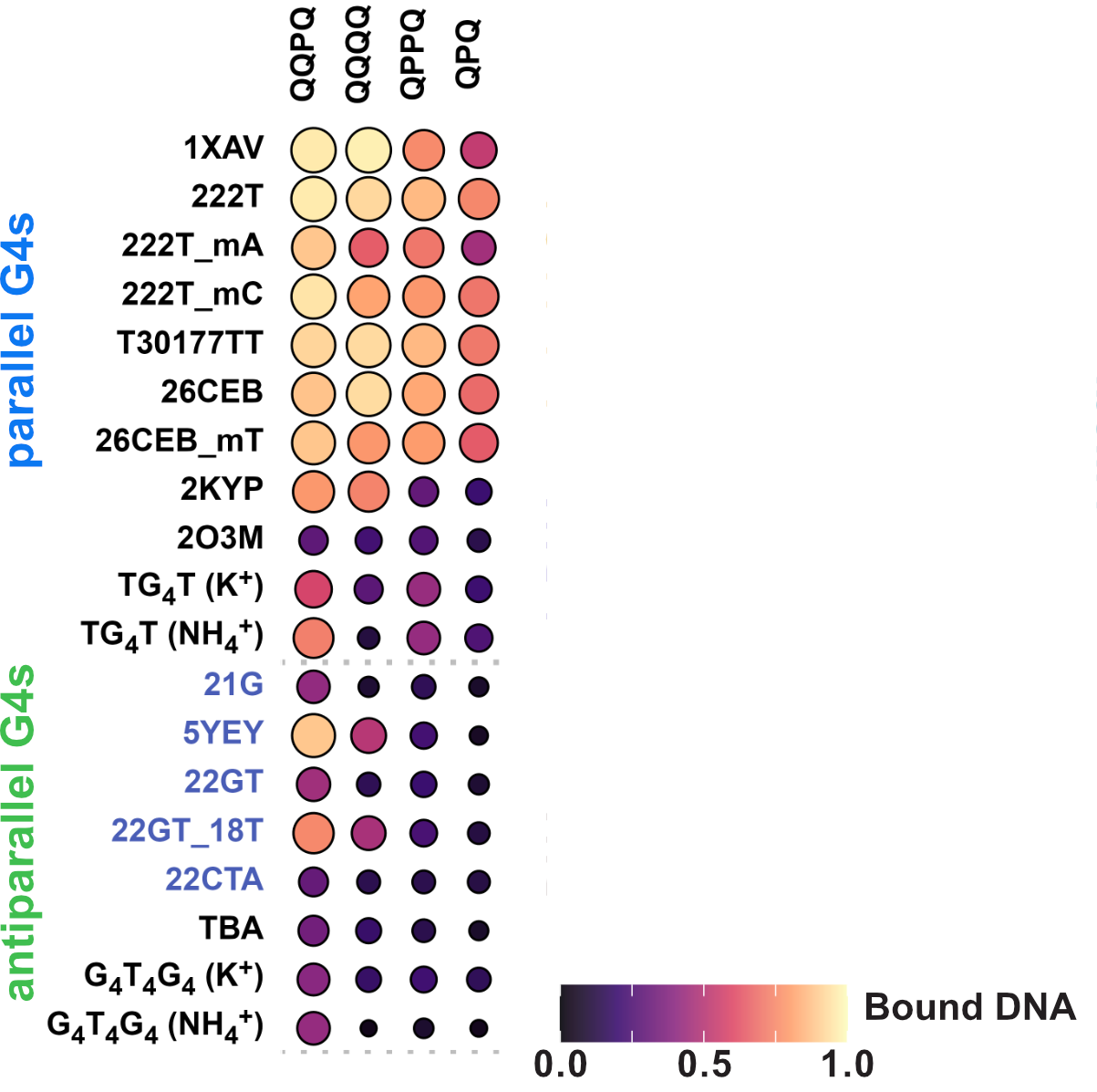
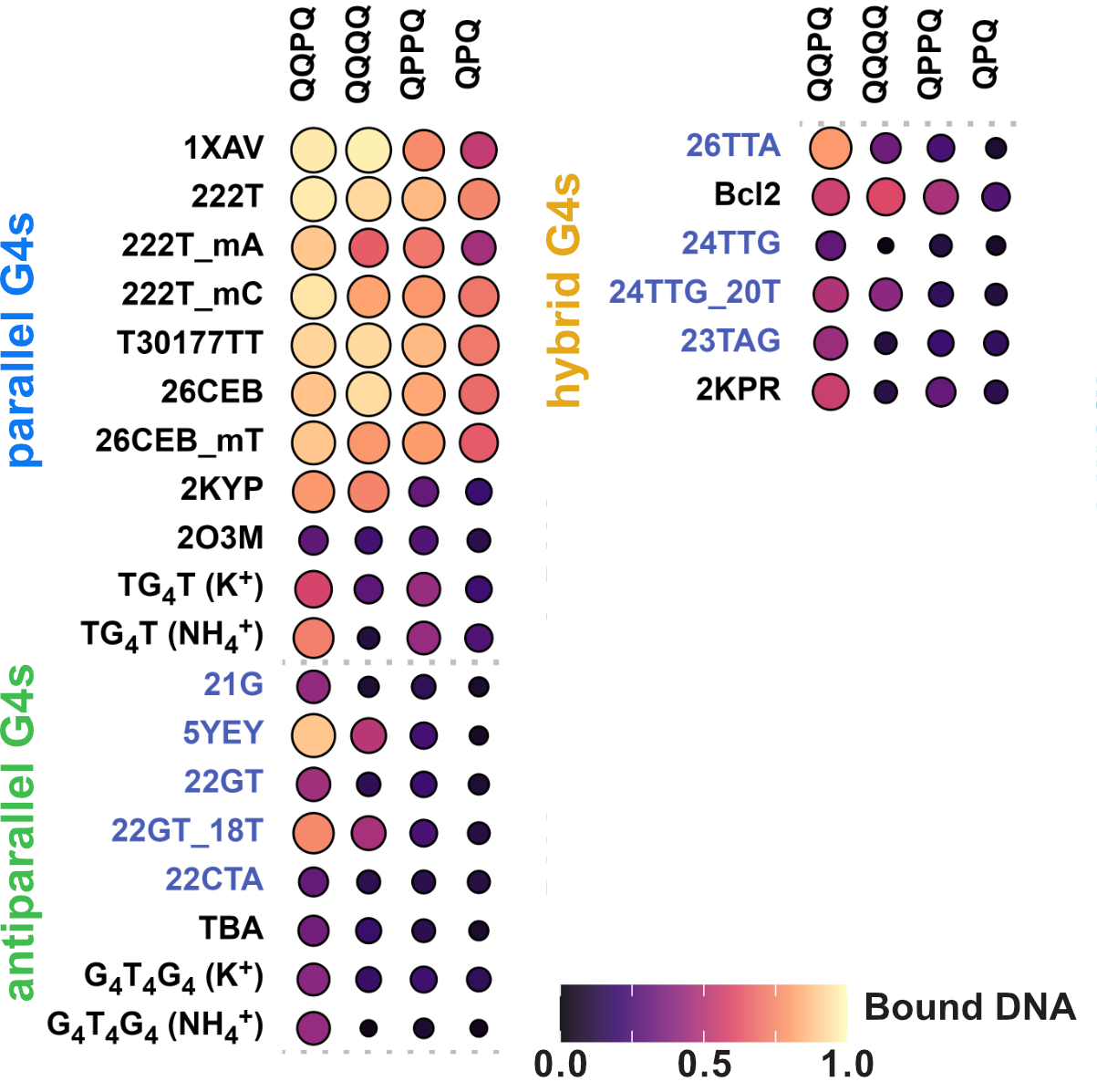
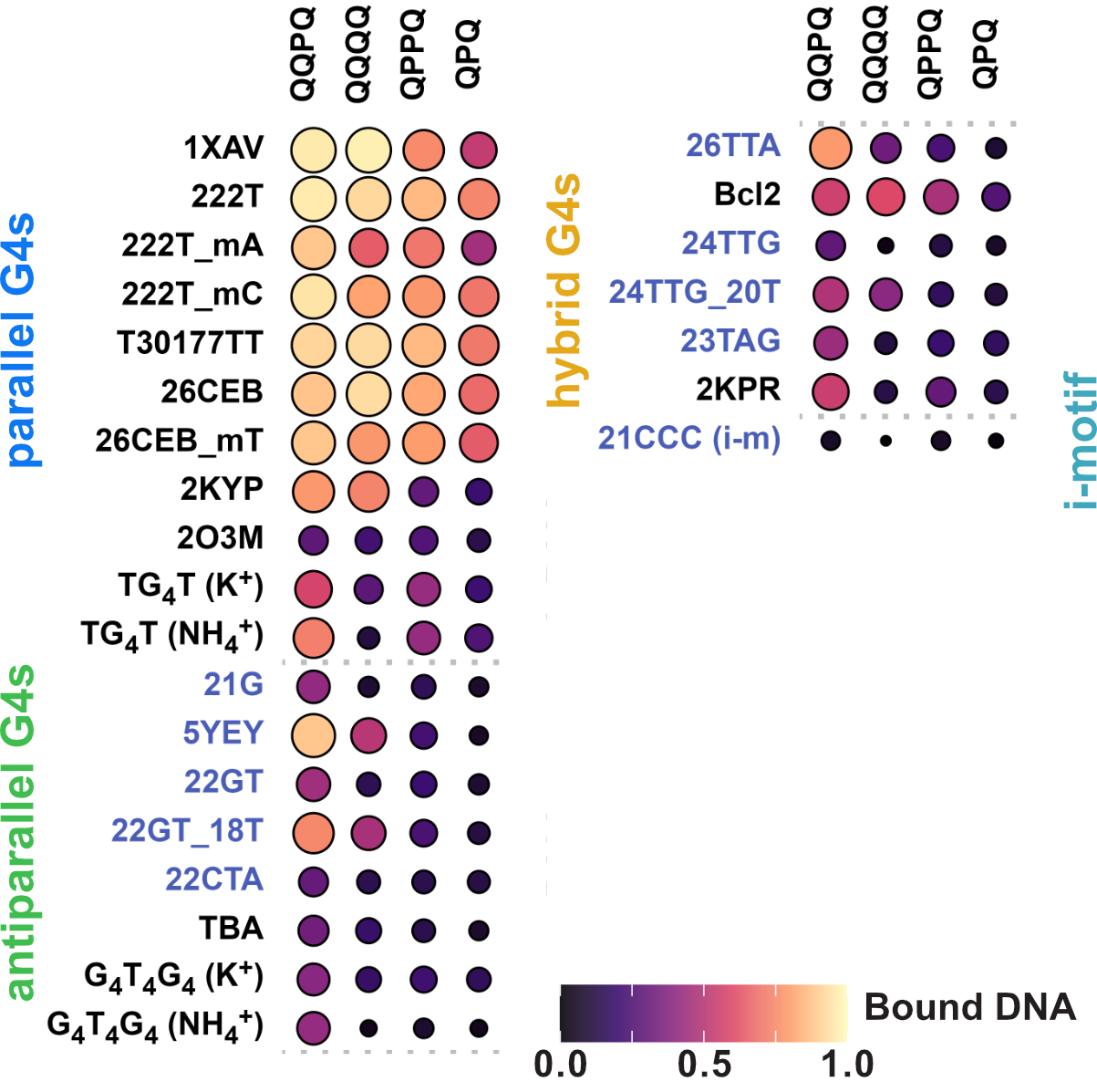
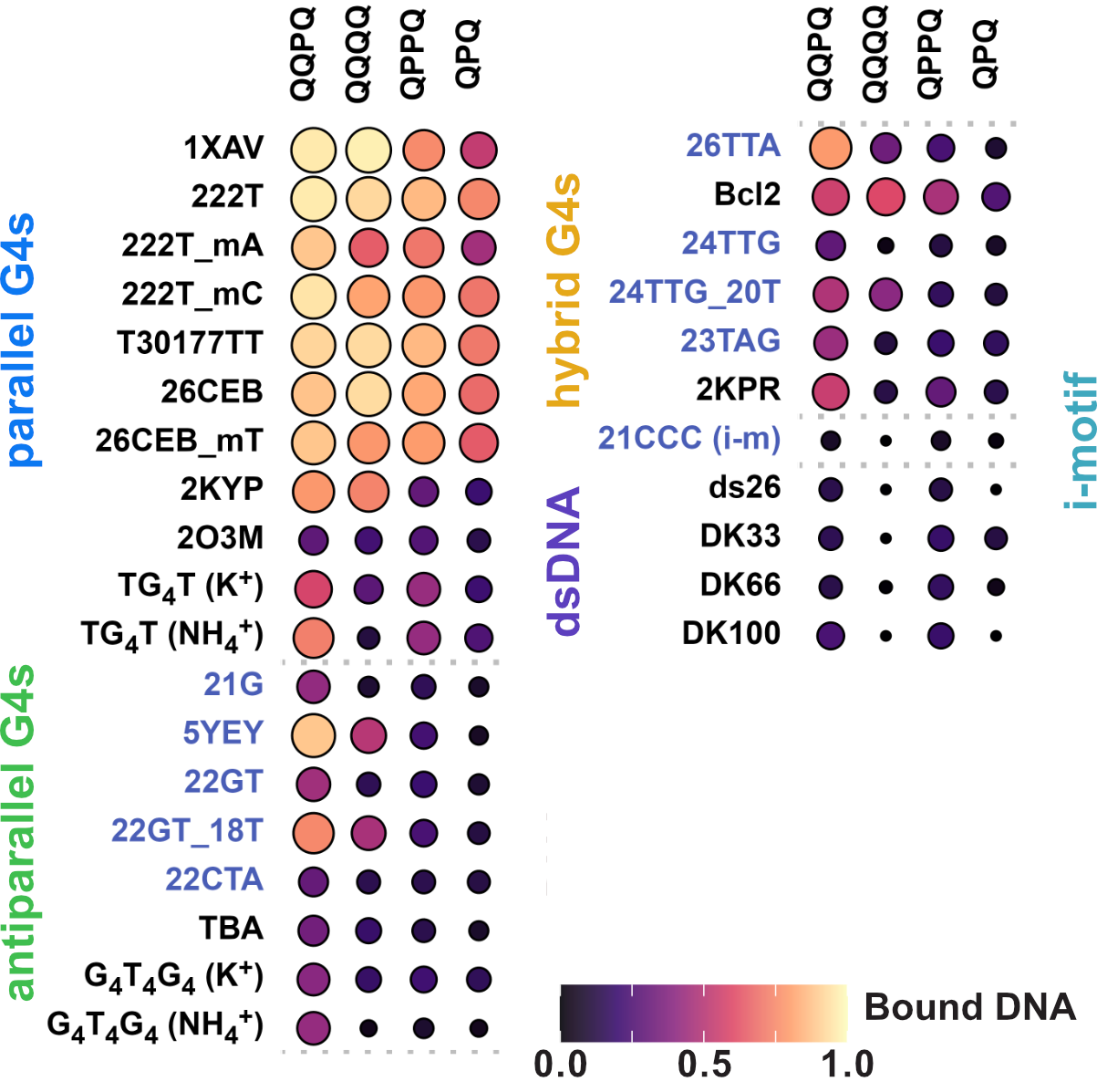
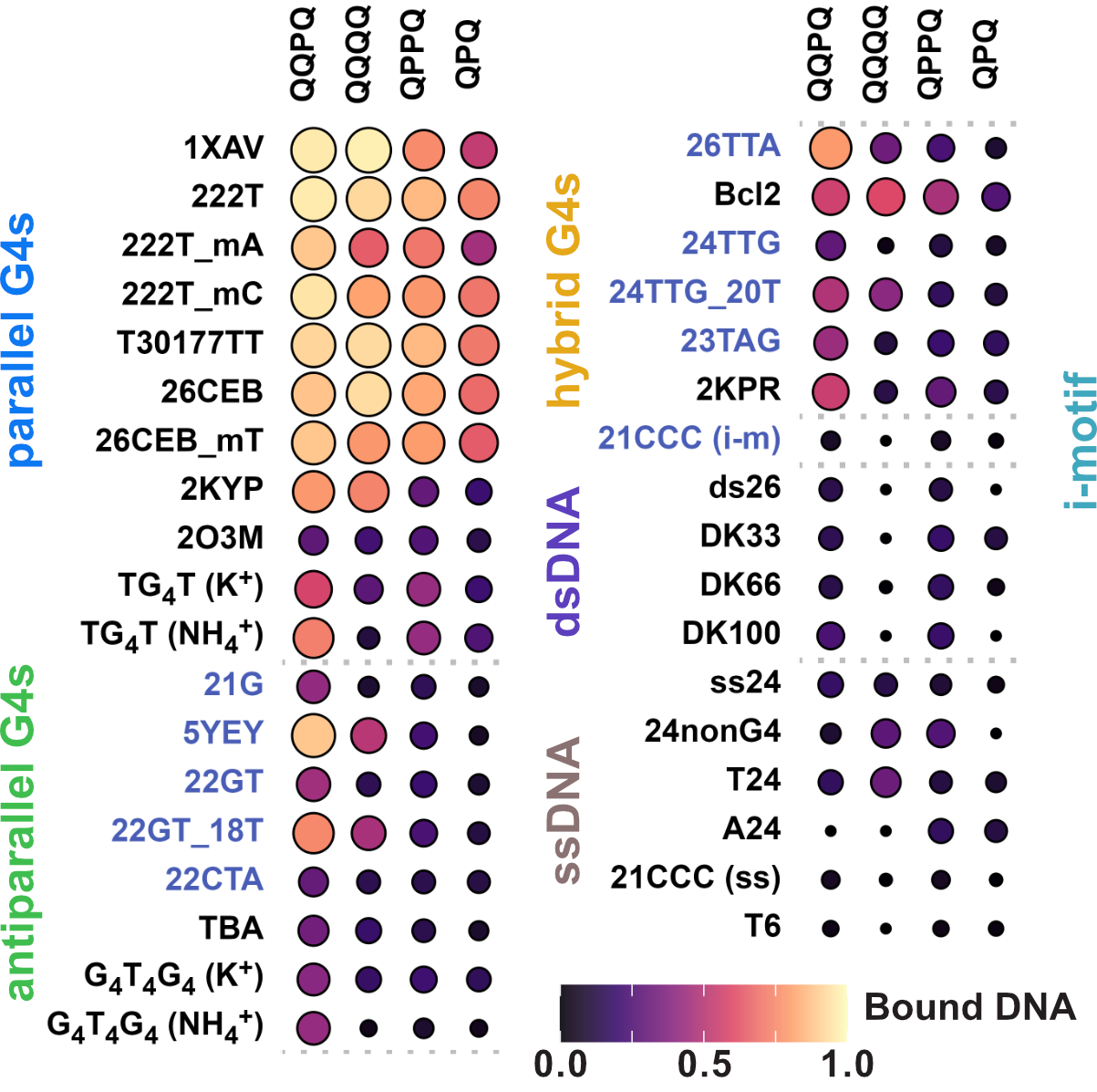
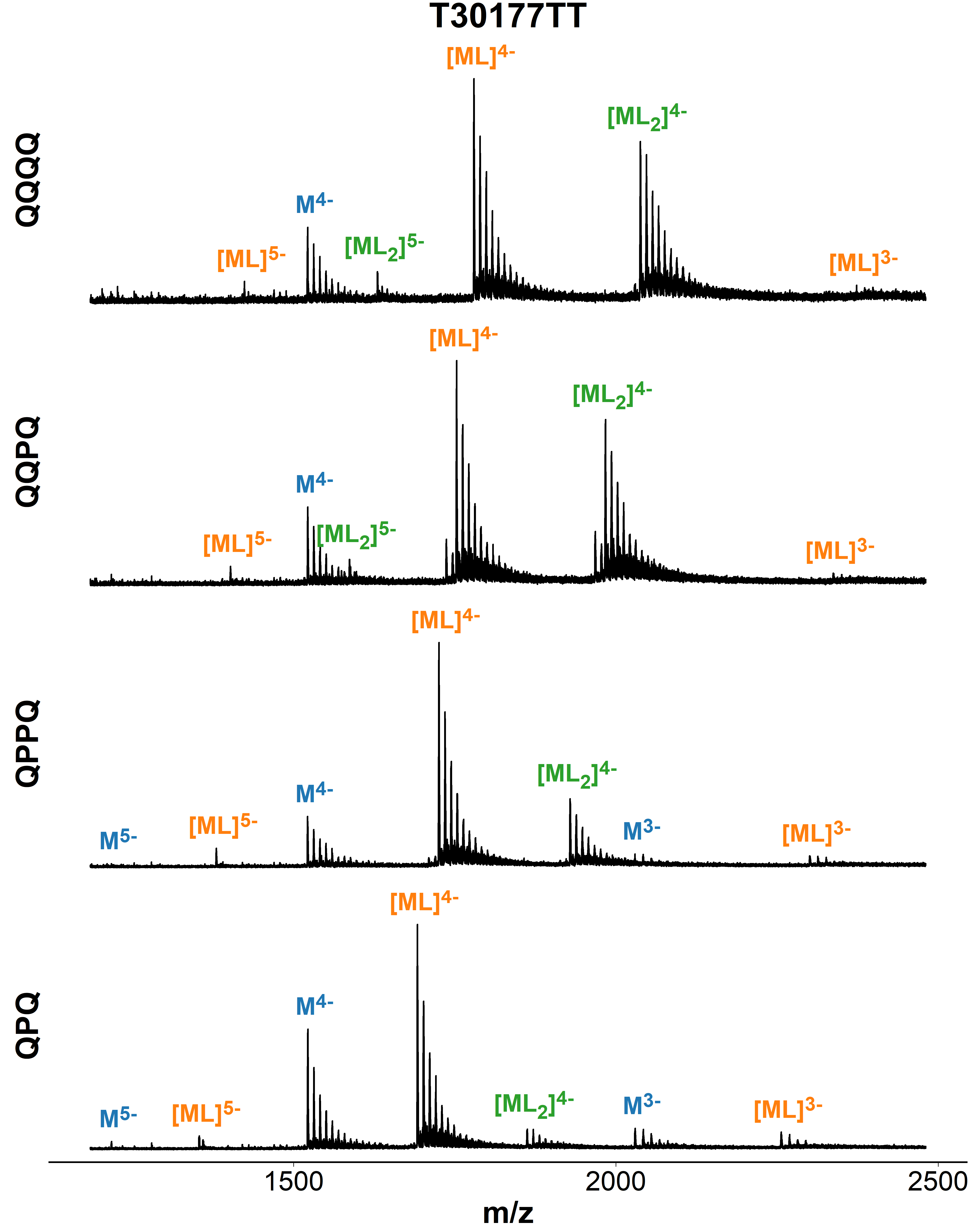
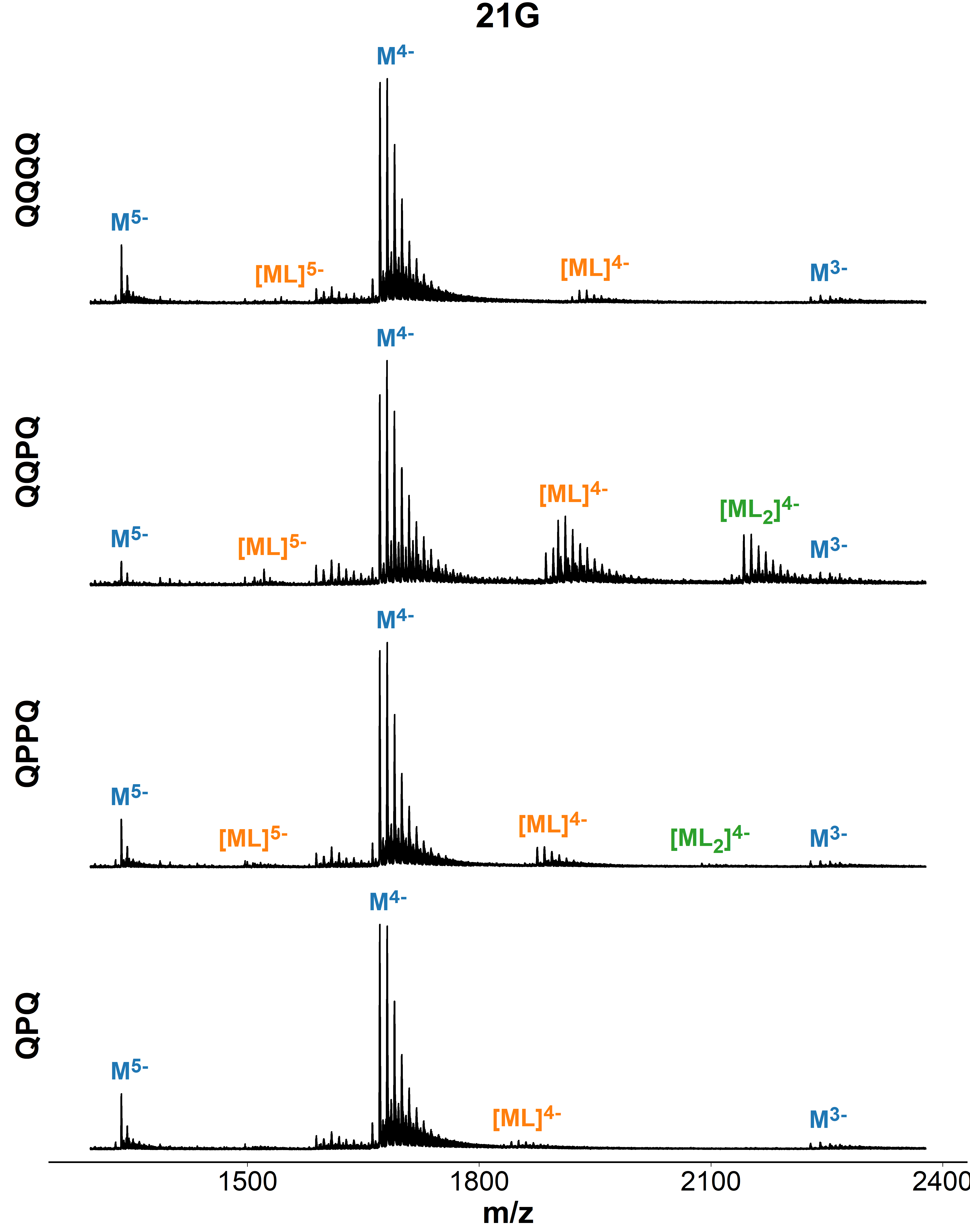
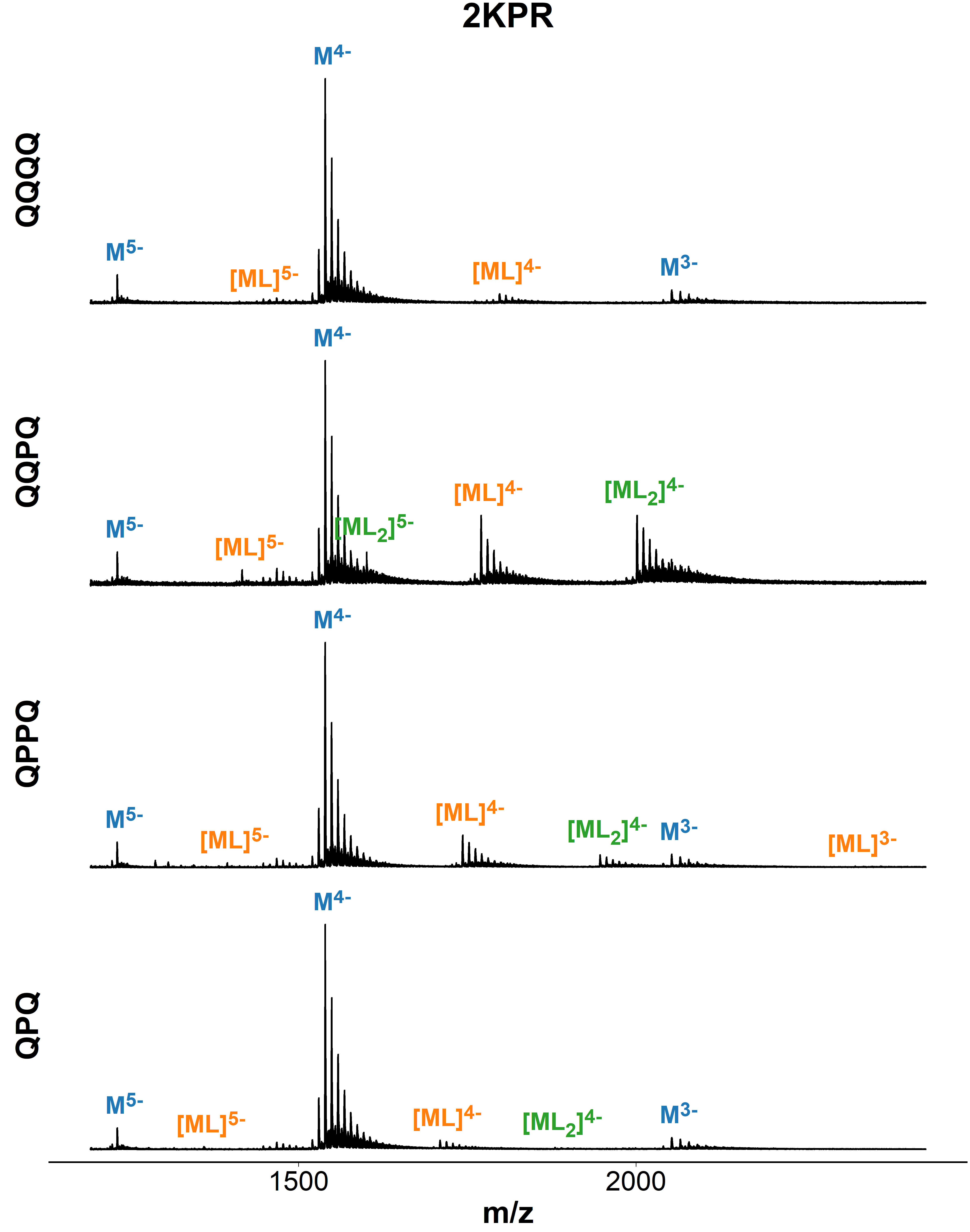
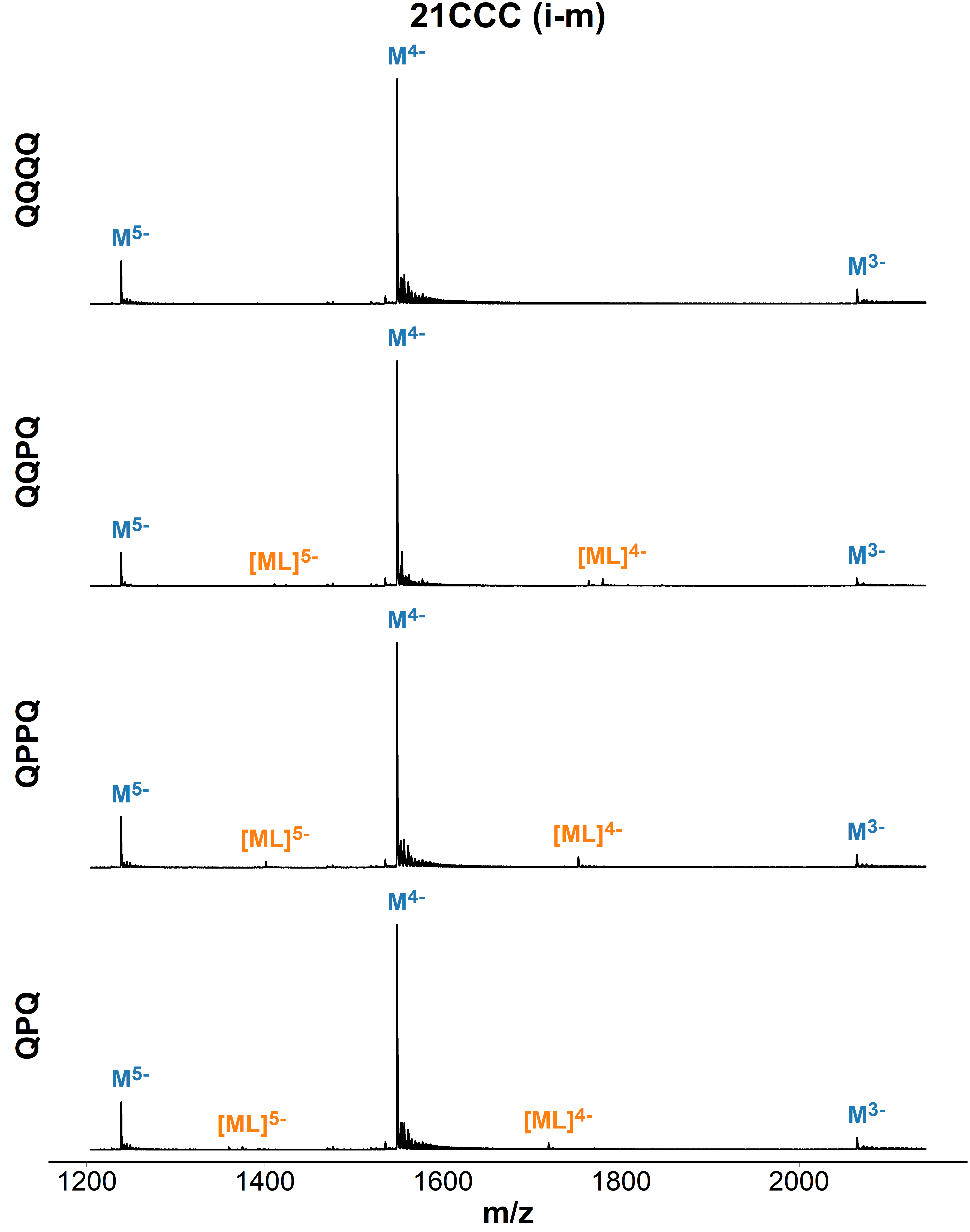
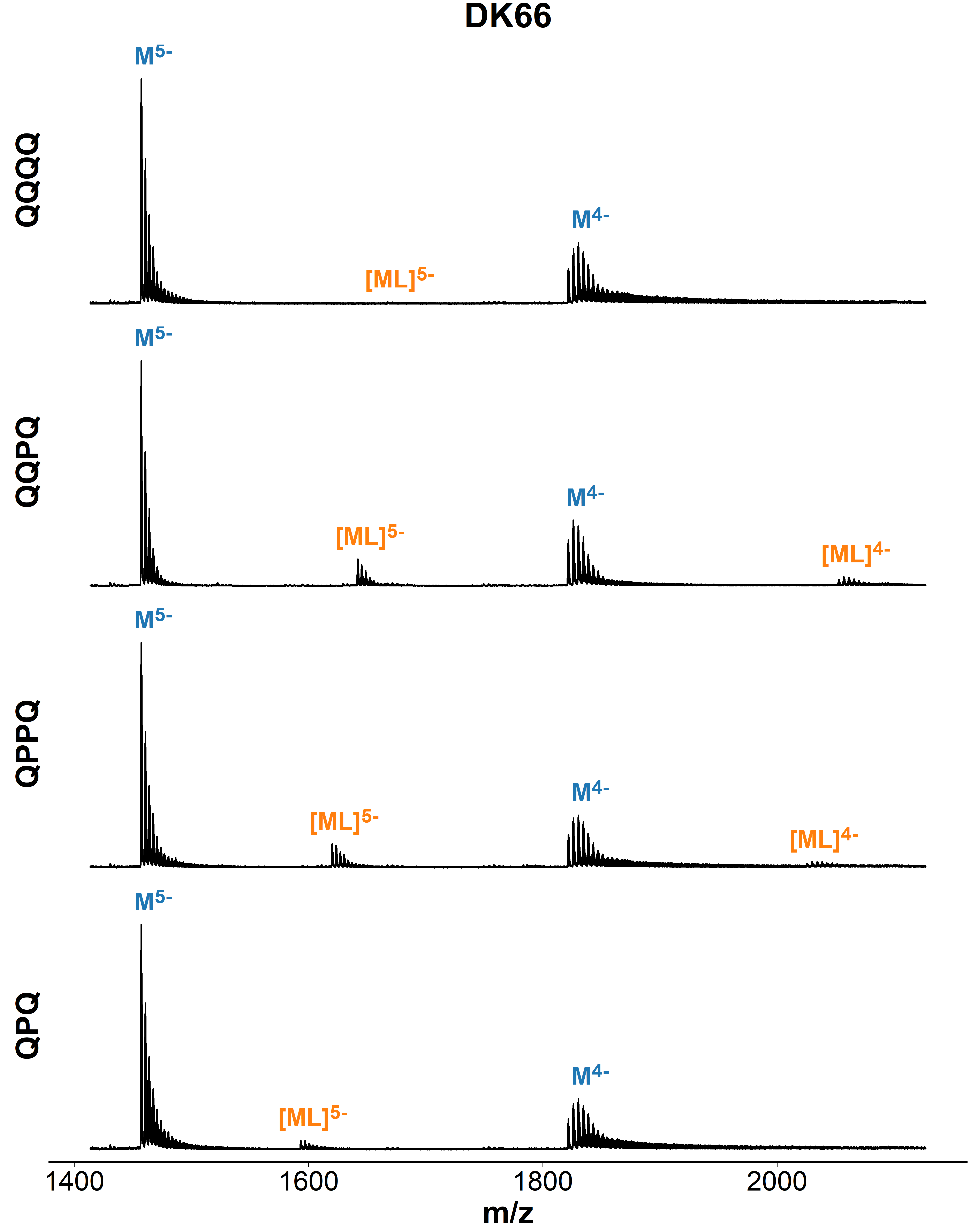
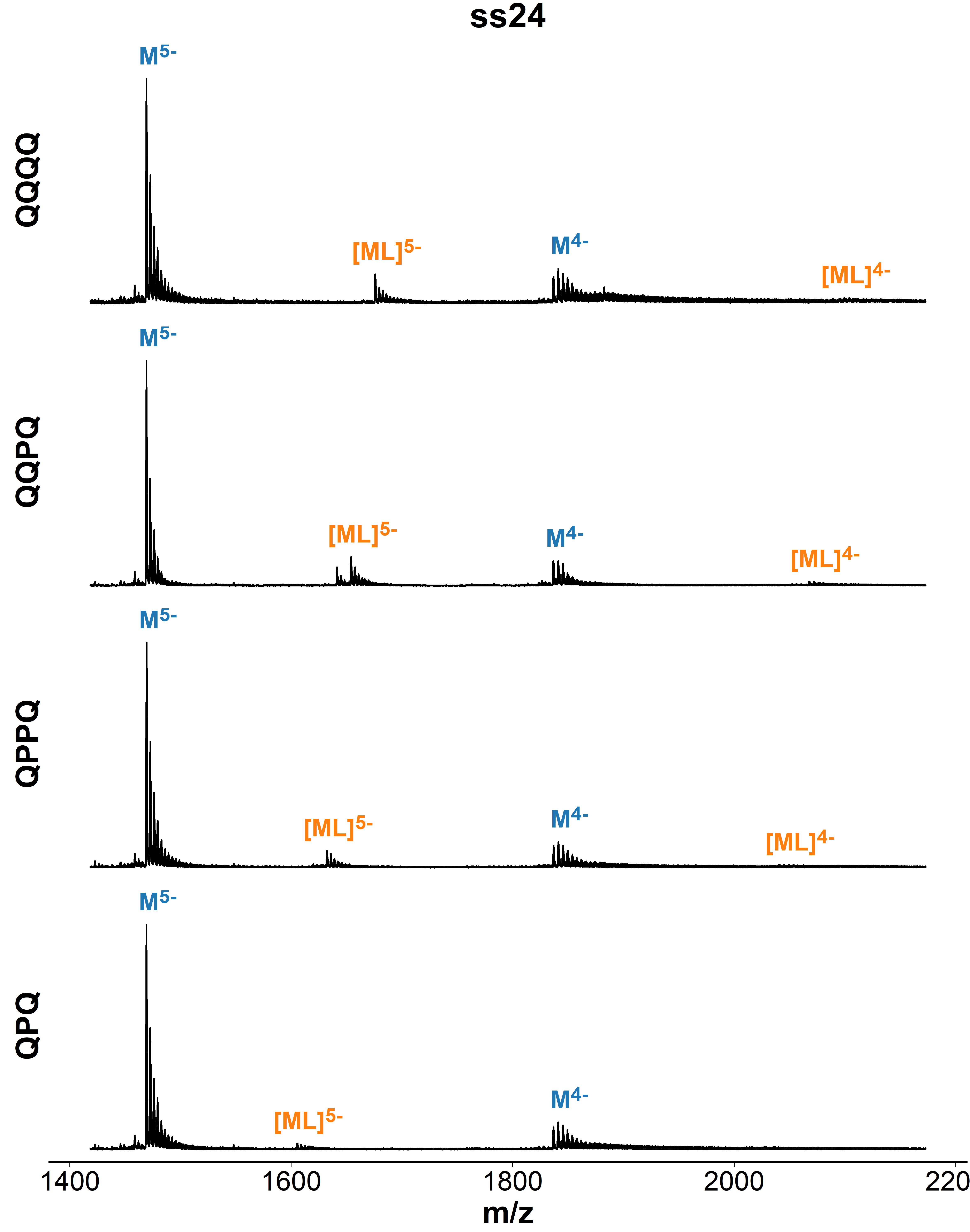

36 oligonucleotides × 4 foldamers = 144 spectra
König, A., Laffilé, V., Thore, S., Mackereth, C., Yatsunyk, L., Ferrand, Y., Largy, E., Gabelica, V., Nucleic Acids Res., 2025, gkaf1365
Selectivity confirmed by high-throughput nMS K titrations
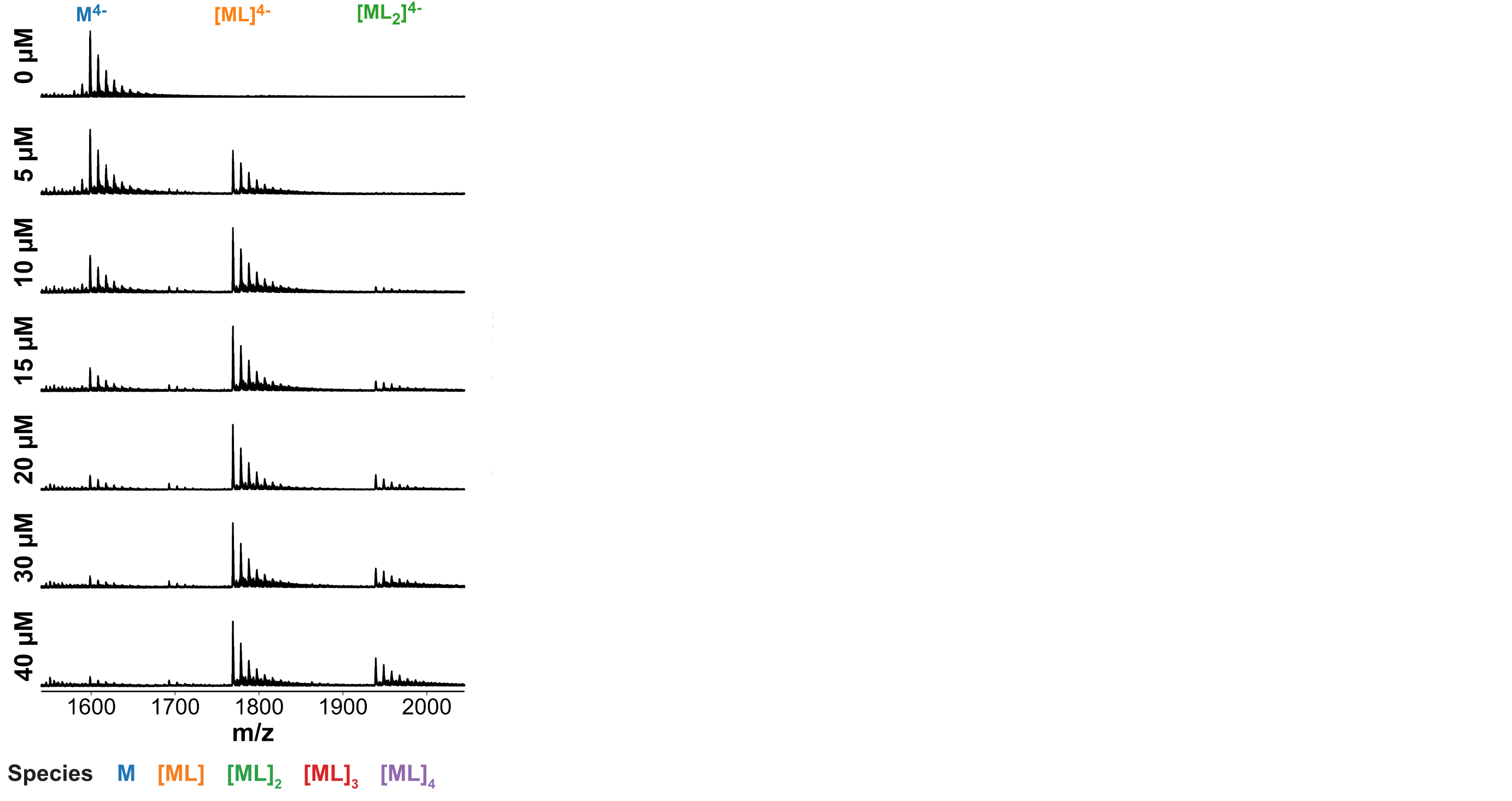
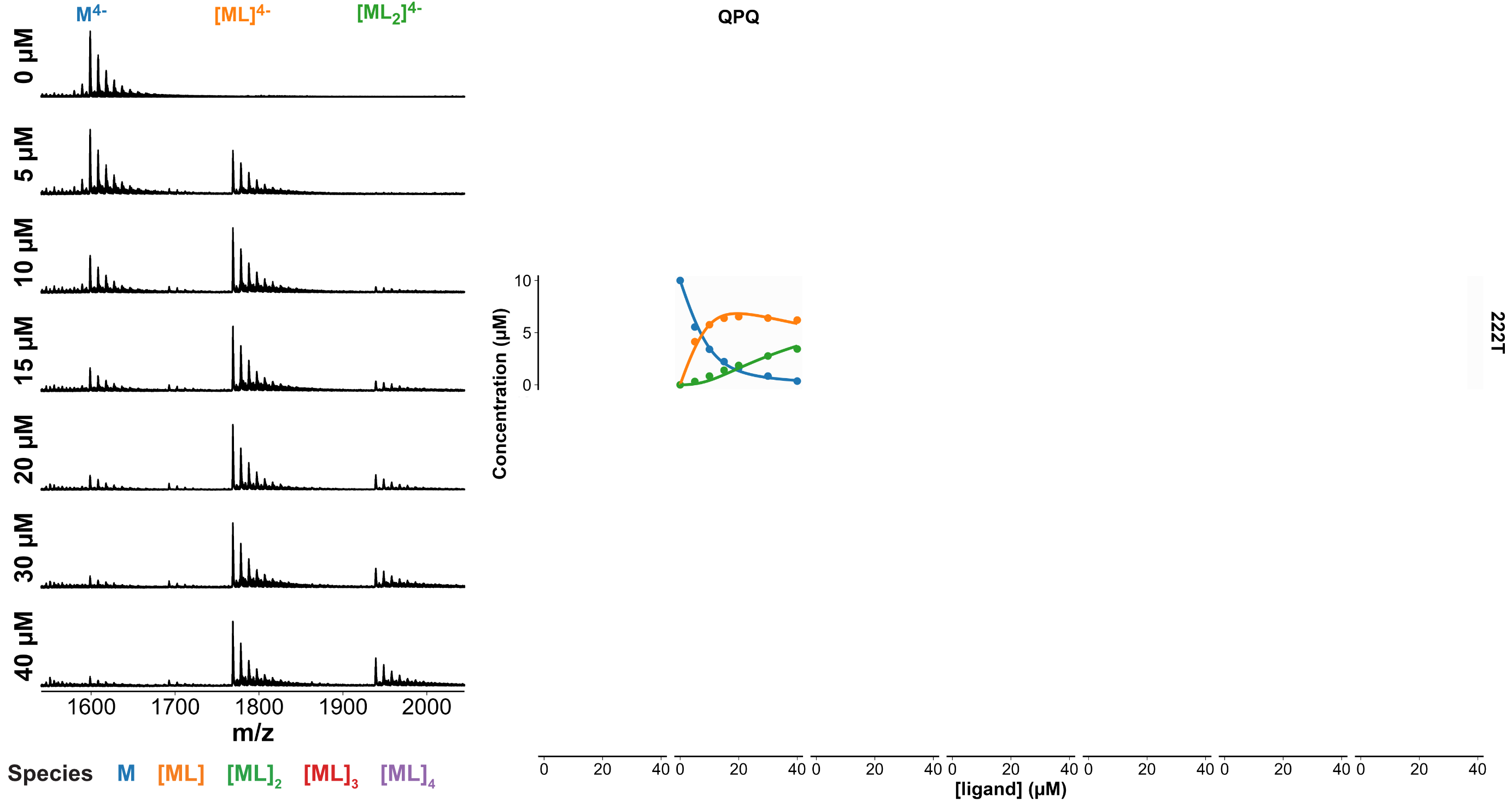
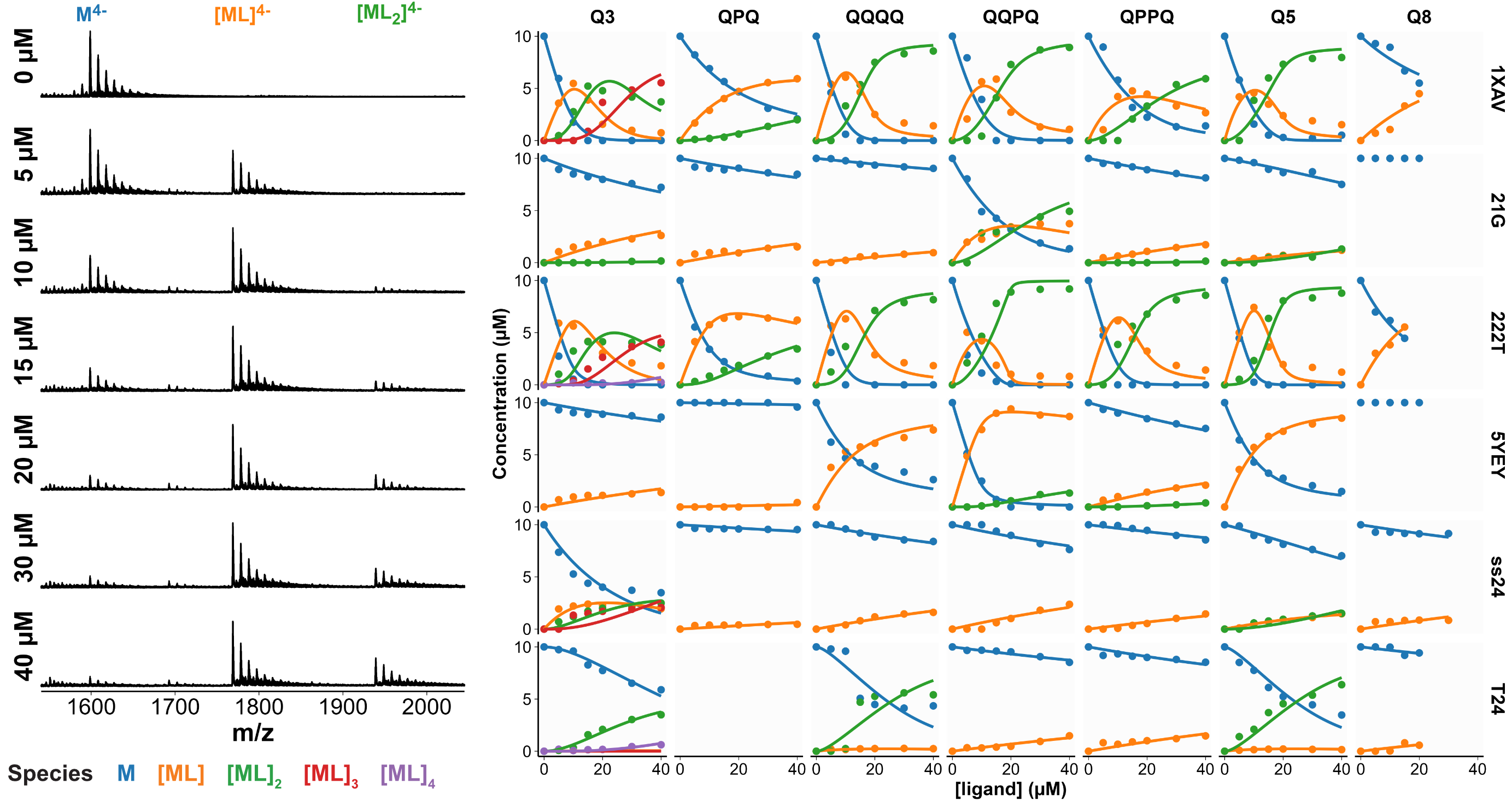
6 oligonucleotides × 7 foldamers × 7 foldamer concentrations = 294 spectra
König, A., Laffilé, V., Thore, S., Mackereth, C., Yatsunyk, L., Ferrand, Y., Largy, E., Gabelica, V., Nucleic Acids Res., 2025, gkaf1365
Not completely crystal clear binding
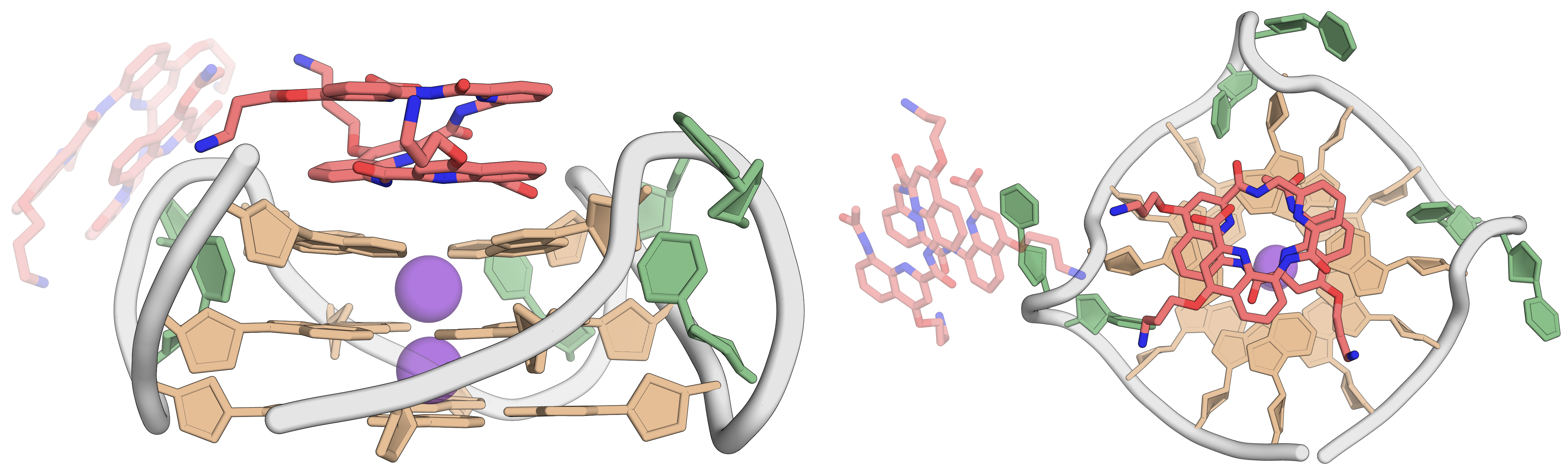
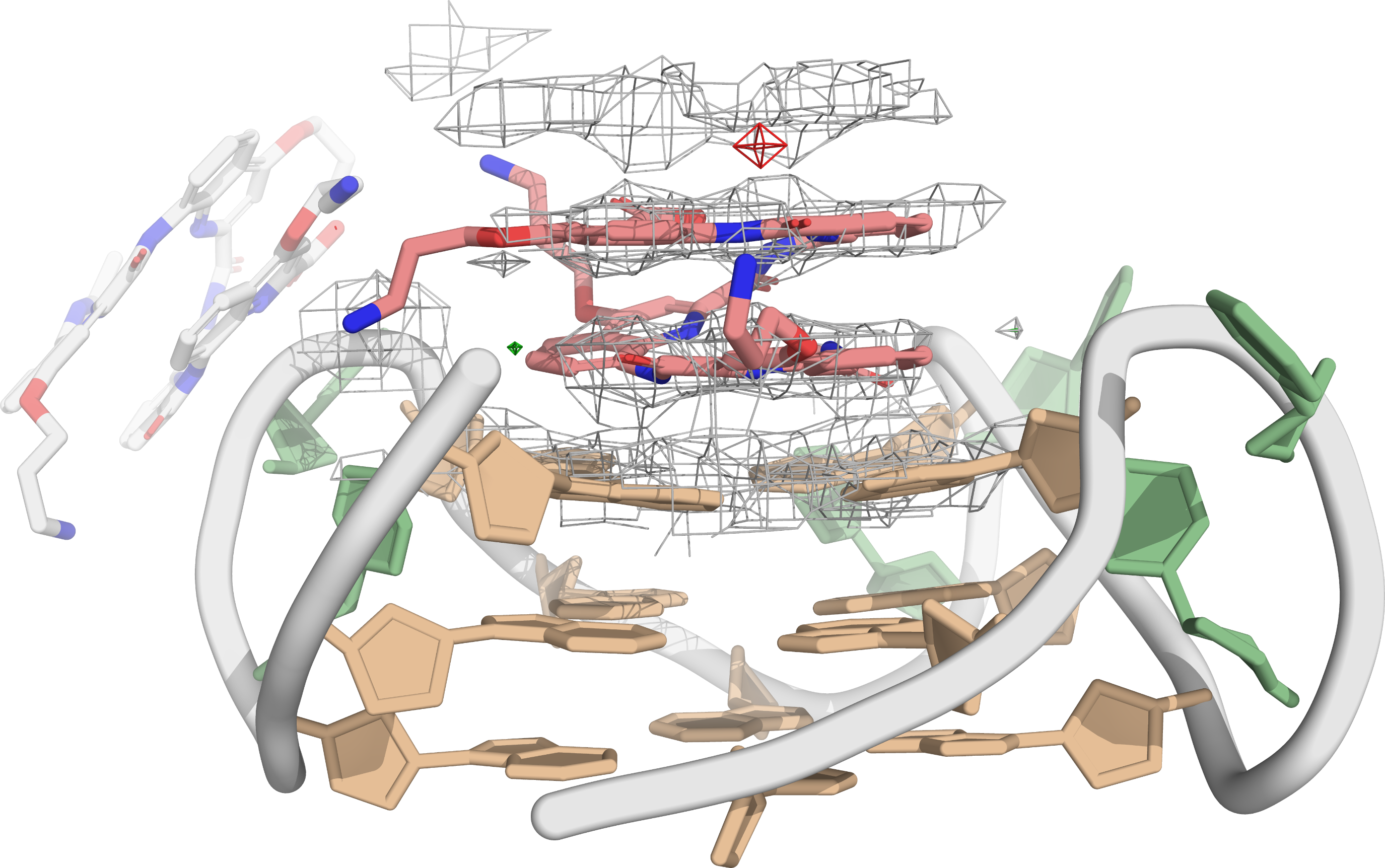

QQPQ complexed to 222T adopting a parallel topology. PDB: 8QN2.
König, A., Laffilé, V., Thore, S., Mackereth, C., Yatsunyk, L., Ferrand, Y., Largy, E., Gabelica, V., Nucleic Acids Res., 2025, gkaf1365
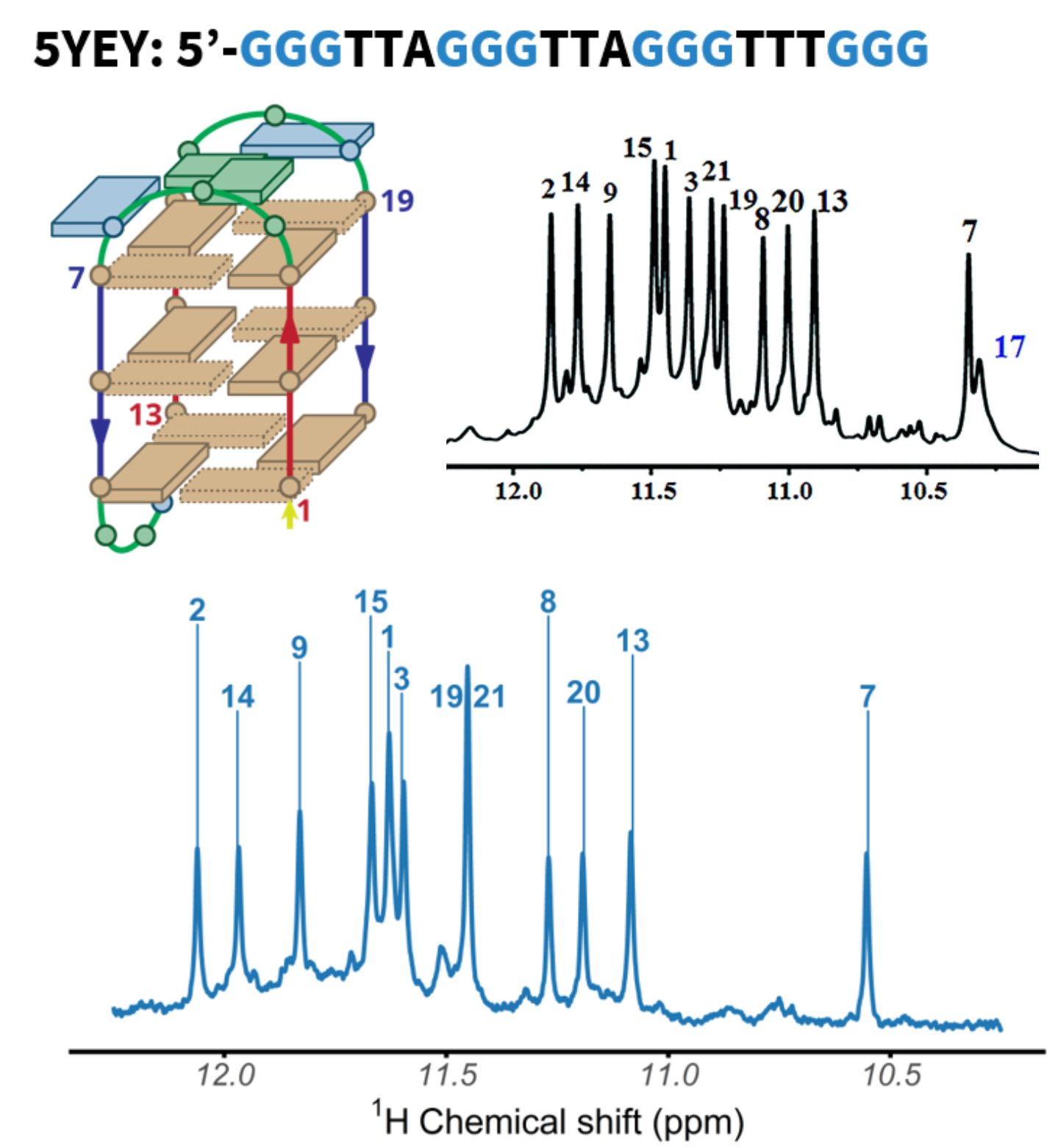
NMR confirms the stacking on both ends
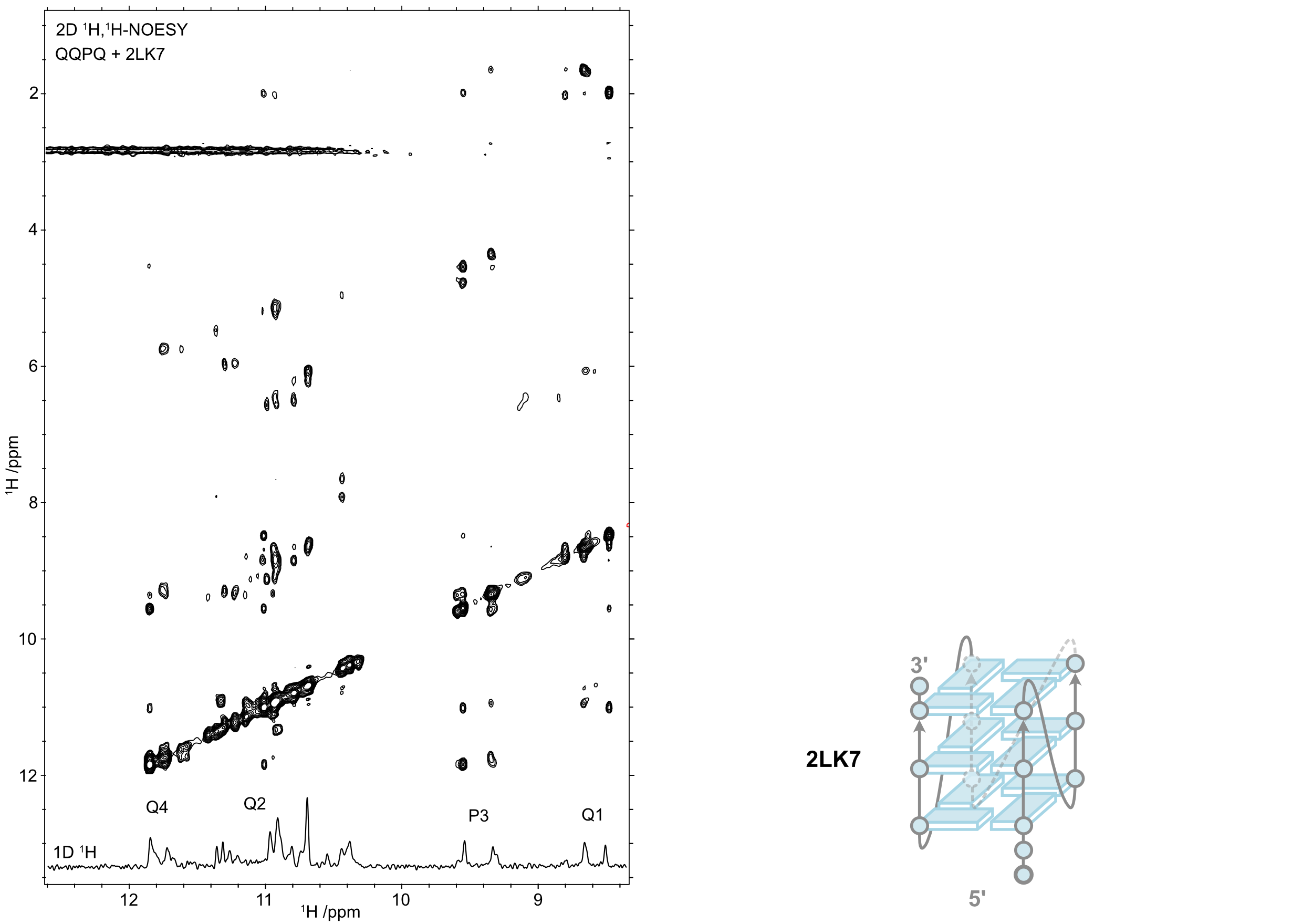
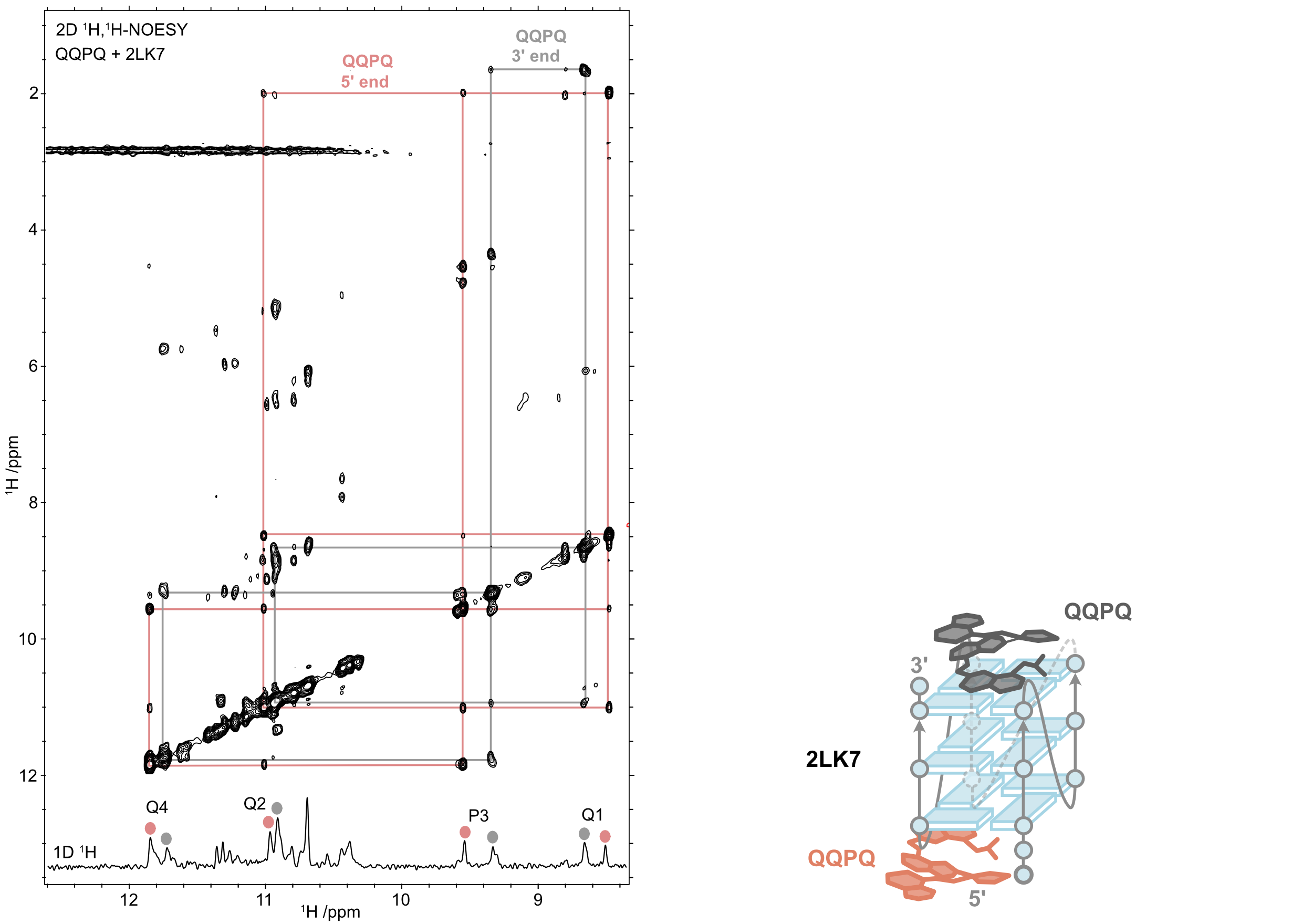
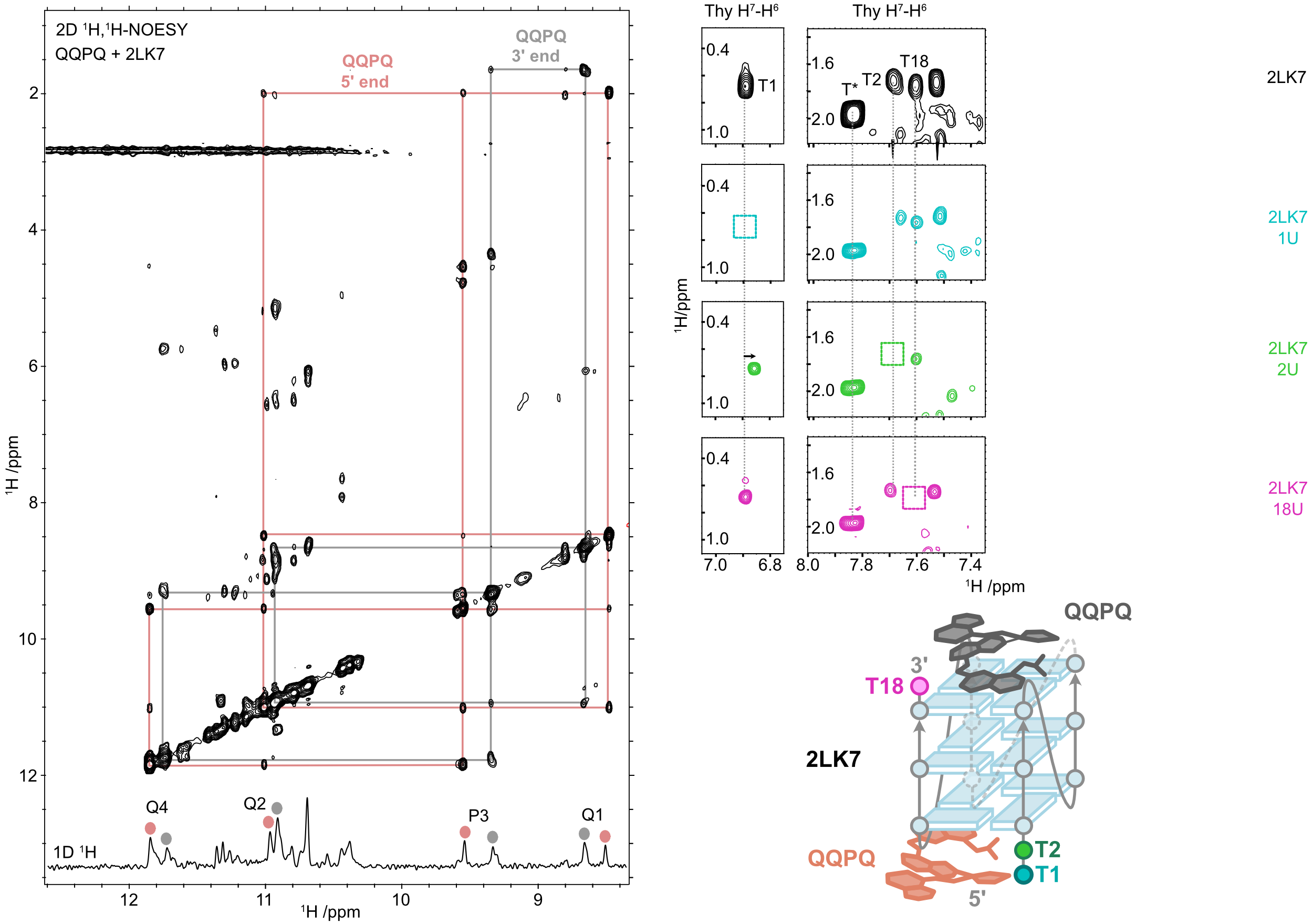
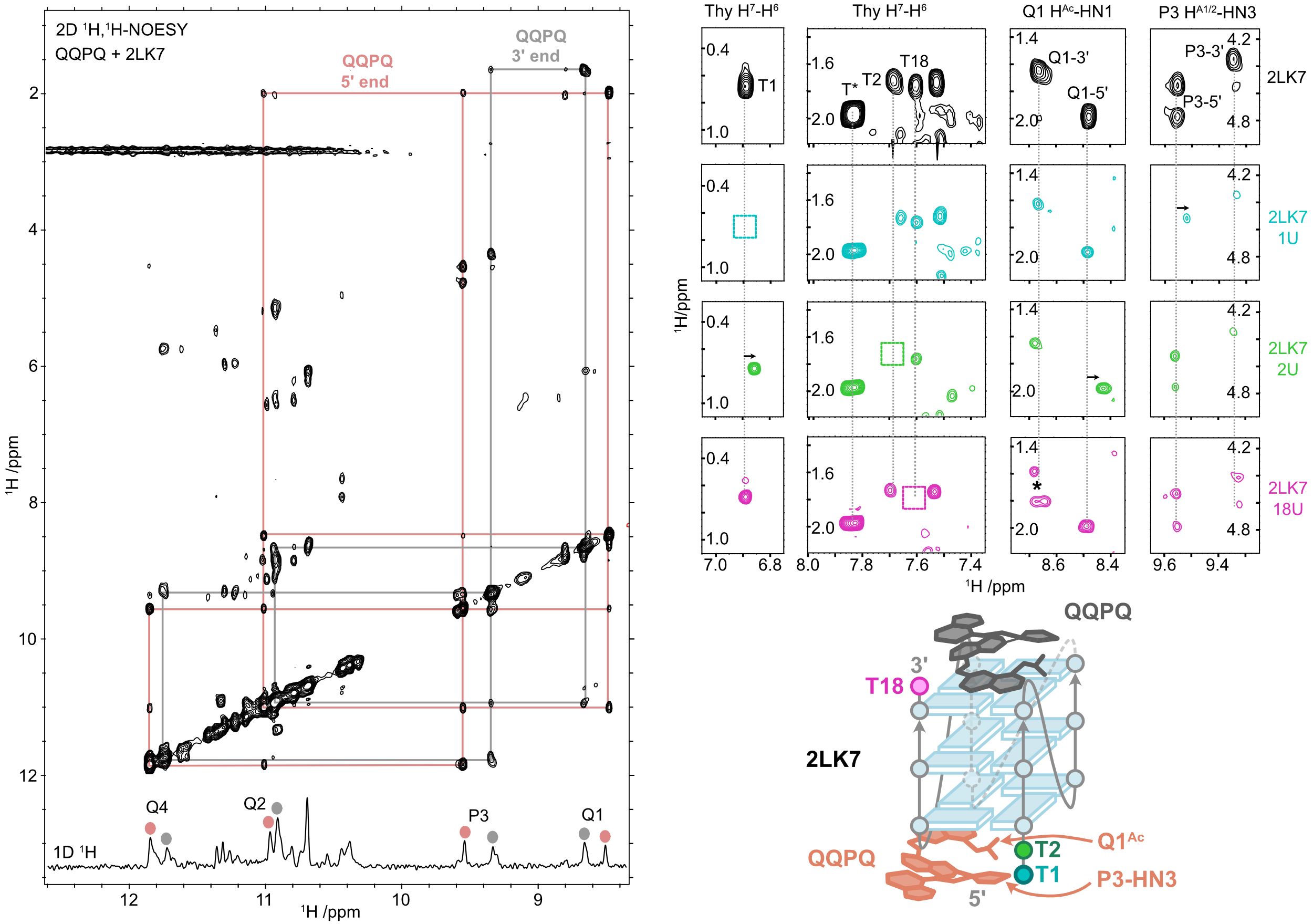

QQPQ complexed to 2LK7 (and not 222T) and T → U variants (missing H7, see boxes).
König, A., Laffilé, V., Thore, S., Mackereth, C., Yatsunyk, L., Ferrand, Y., Largy, E., Gabelica, V., Nucleic Acids Res., 2025, gkaf1365
Final models supported by MD simulations
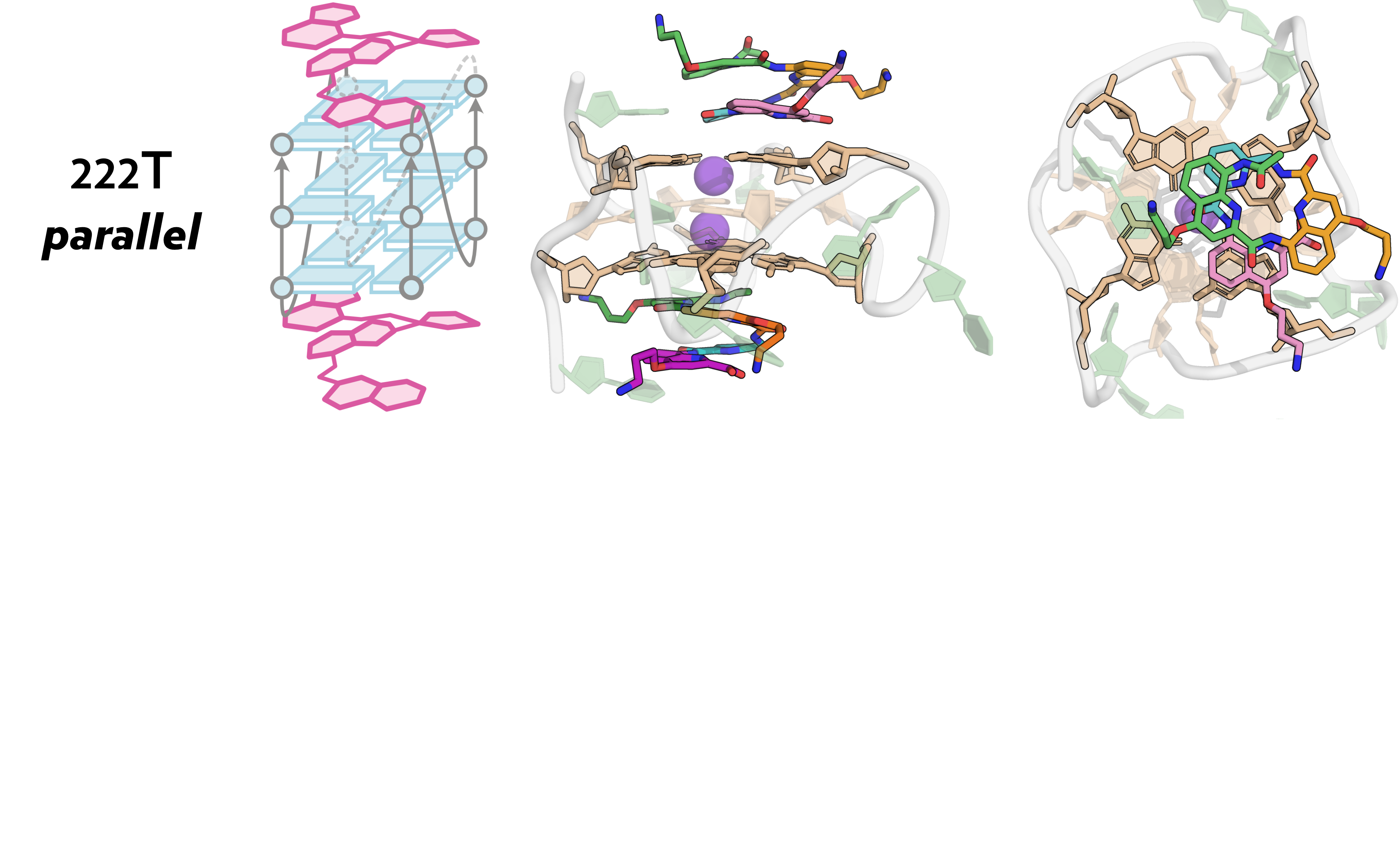
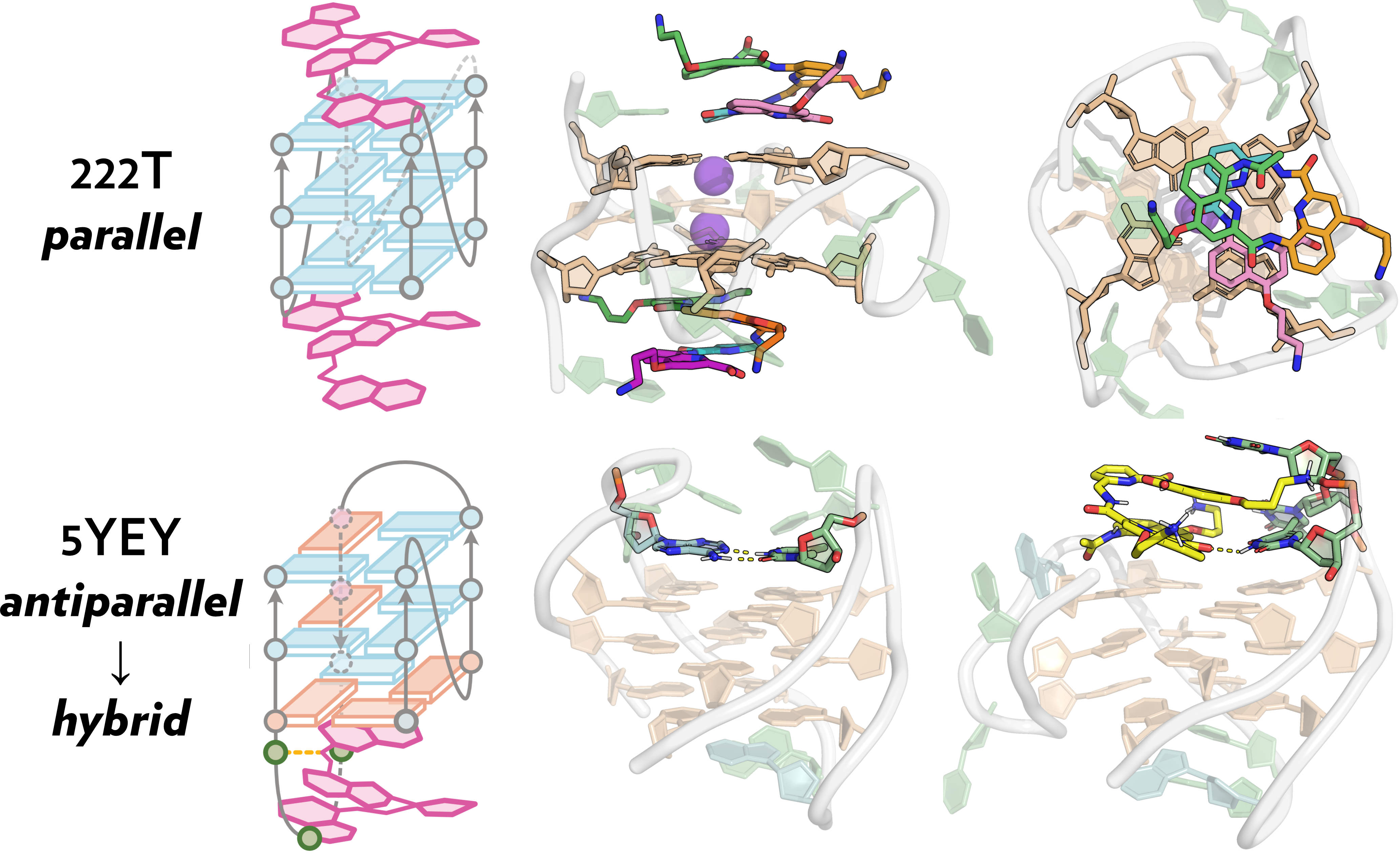
König, A., Laffilé, V., Thore, S., Mackereth, C., Yatsunyk, L., Ferrand, Y., Largy, E., Gabelica, V., Nucleic Acids Res., 2025, gkaf1365
Foldamers allow precisal structural control
of artifical protein assemblies
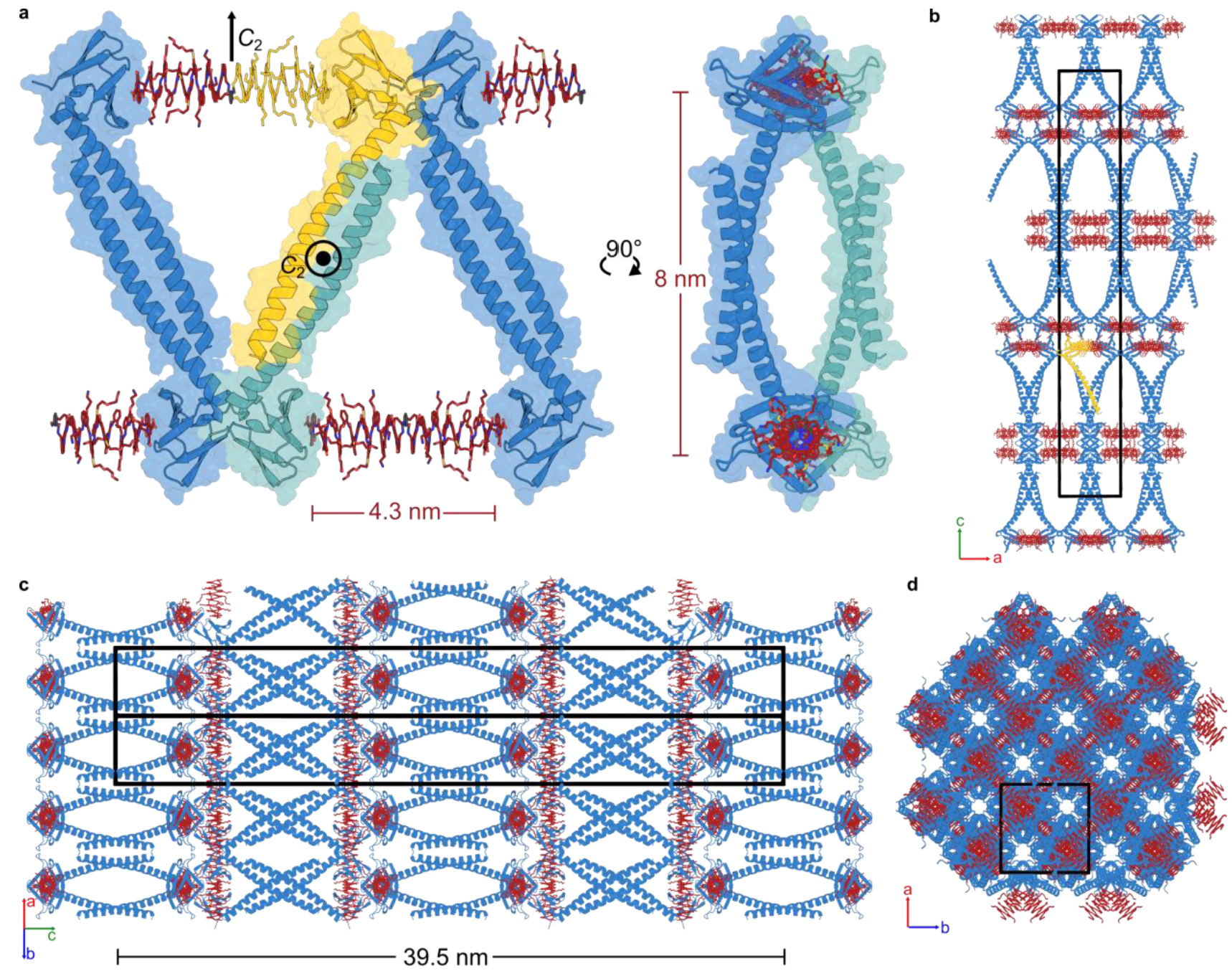
Sigl, J. et al. Nat. Chem., accepted

SEC and nMS are methods of choice
to determine stoichiometries
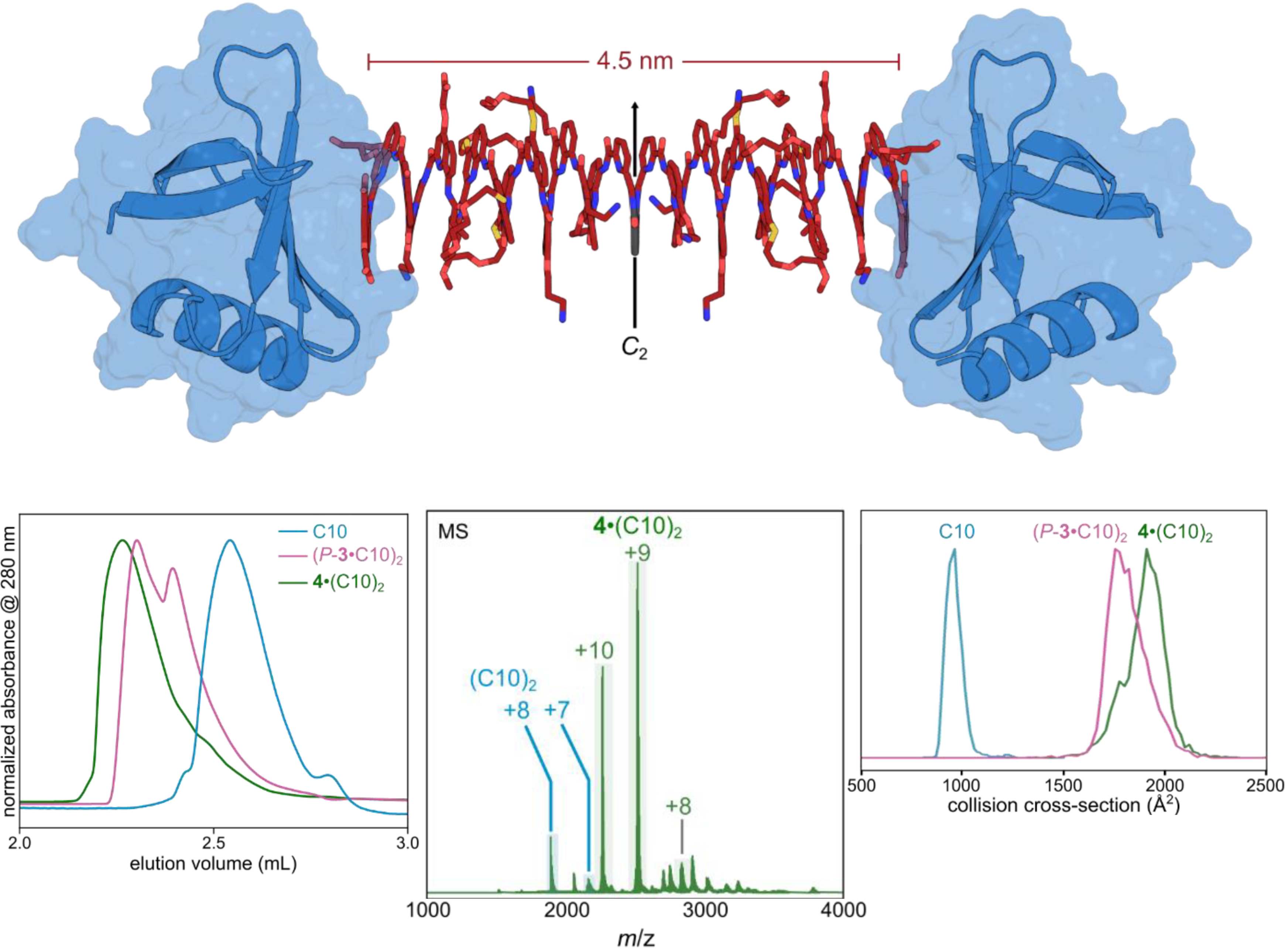
Sigl, J. et al. Nat. Chem., accepted
Foldamers also bind proteins in unexpected ways
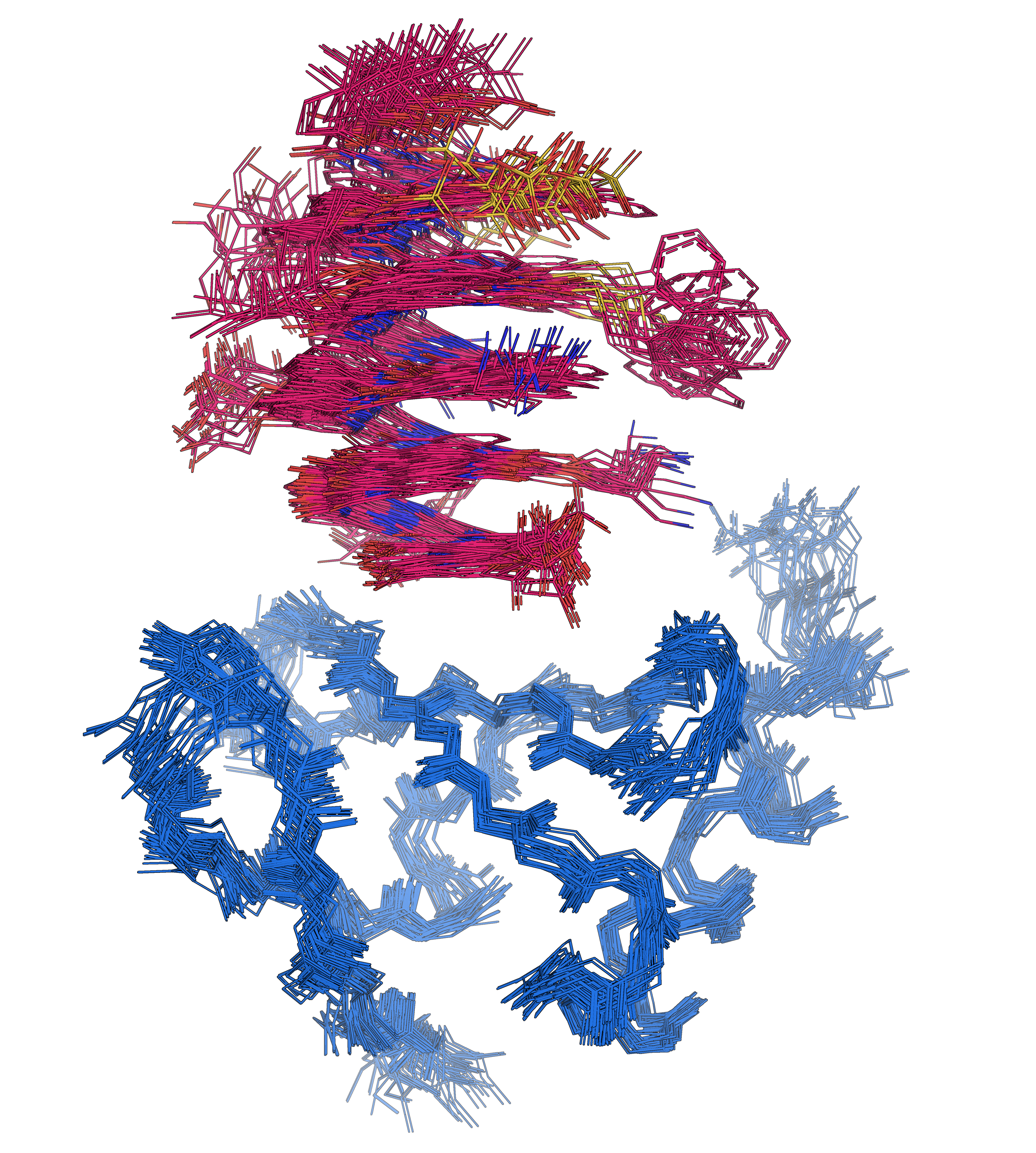
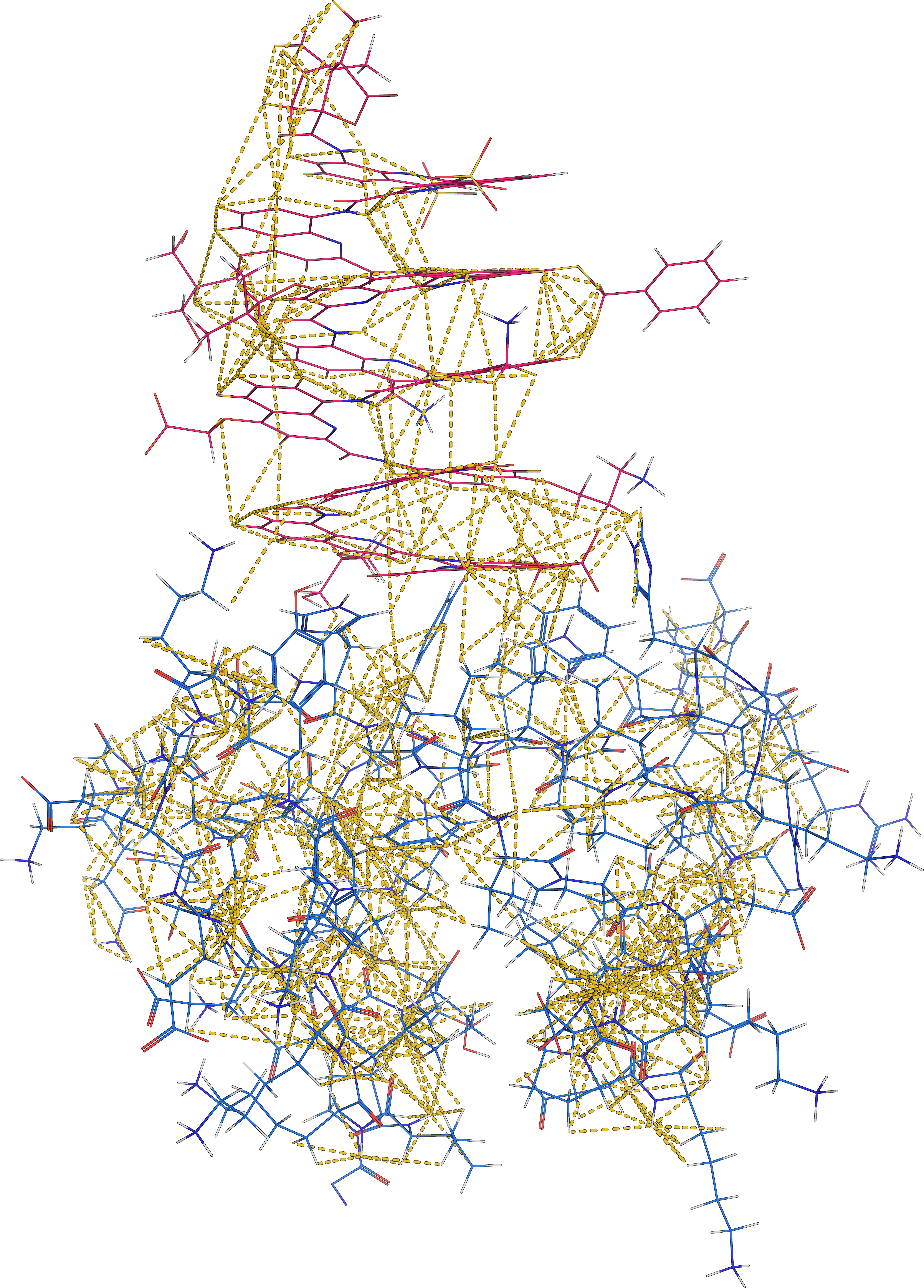

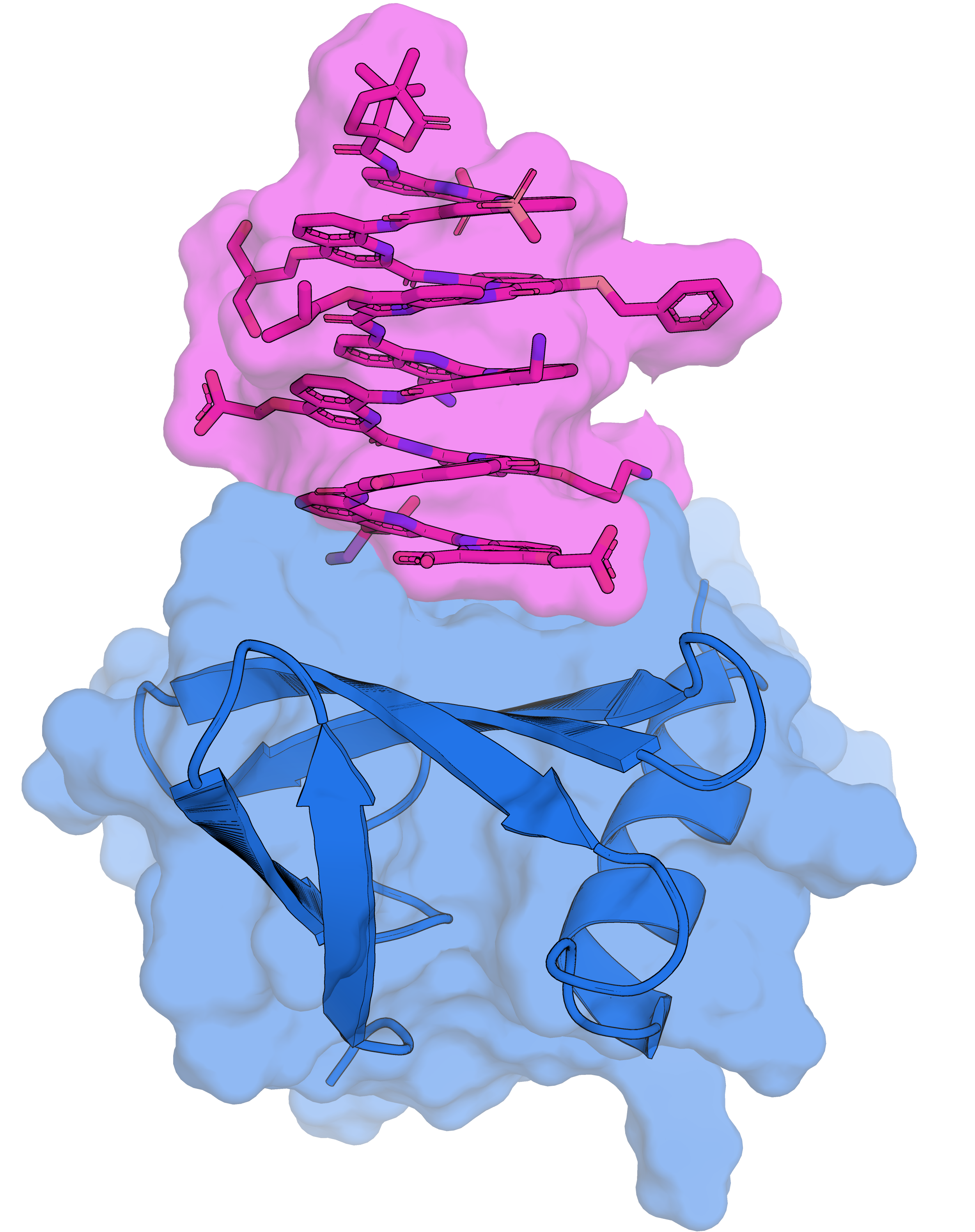
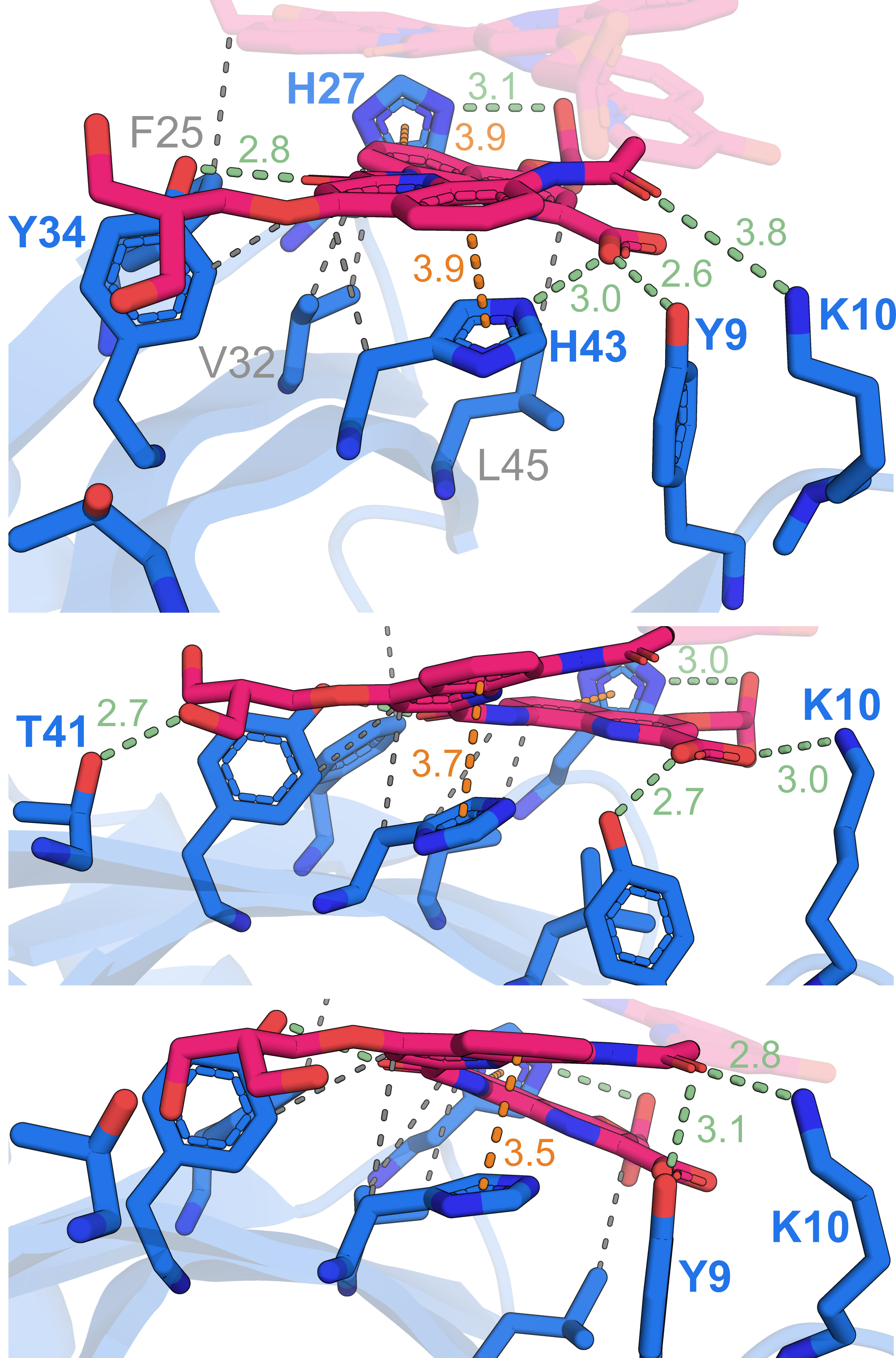
Solution structure of Nanofitin C10 in complex with a 12unit aromatic oligoamide foldamer | PDB: 9T0O | Sigl, J. et al. Nat. Chem., accepted
737 non-ambiguous distance + 114 dihedral angle restraints

Native HDX/MS
from proteins to nucleic acids
HDX/MS gives insights
into protein structural dynamics
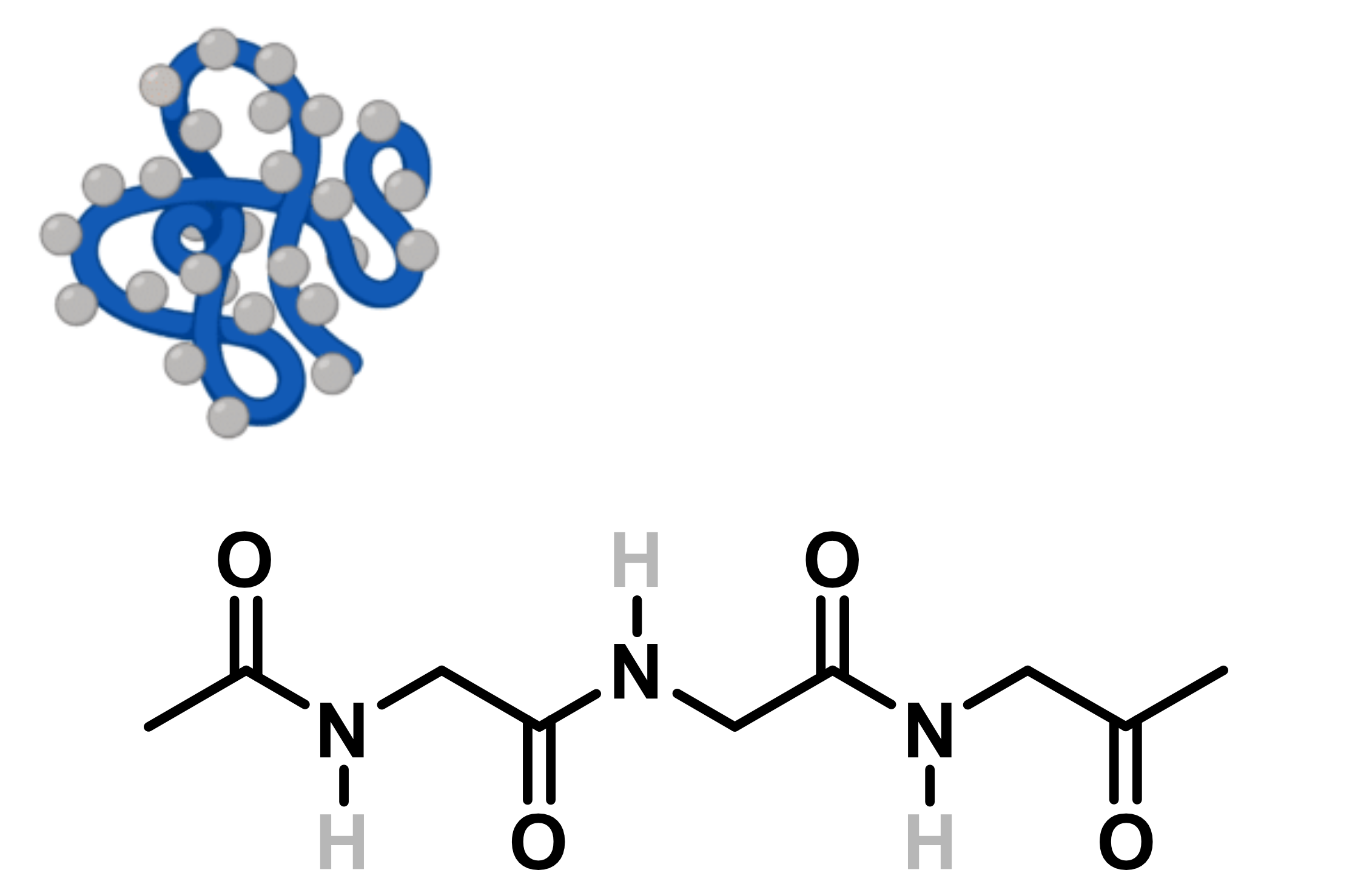
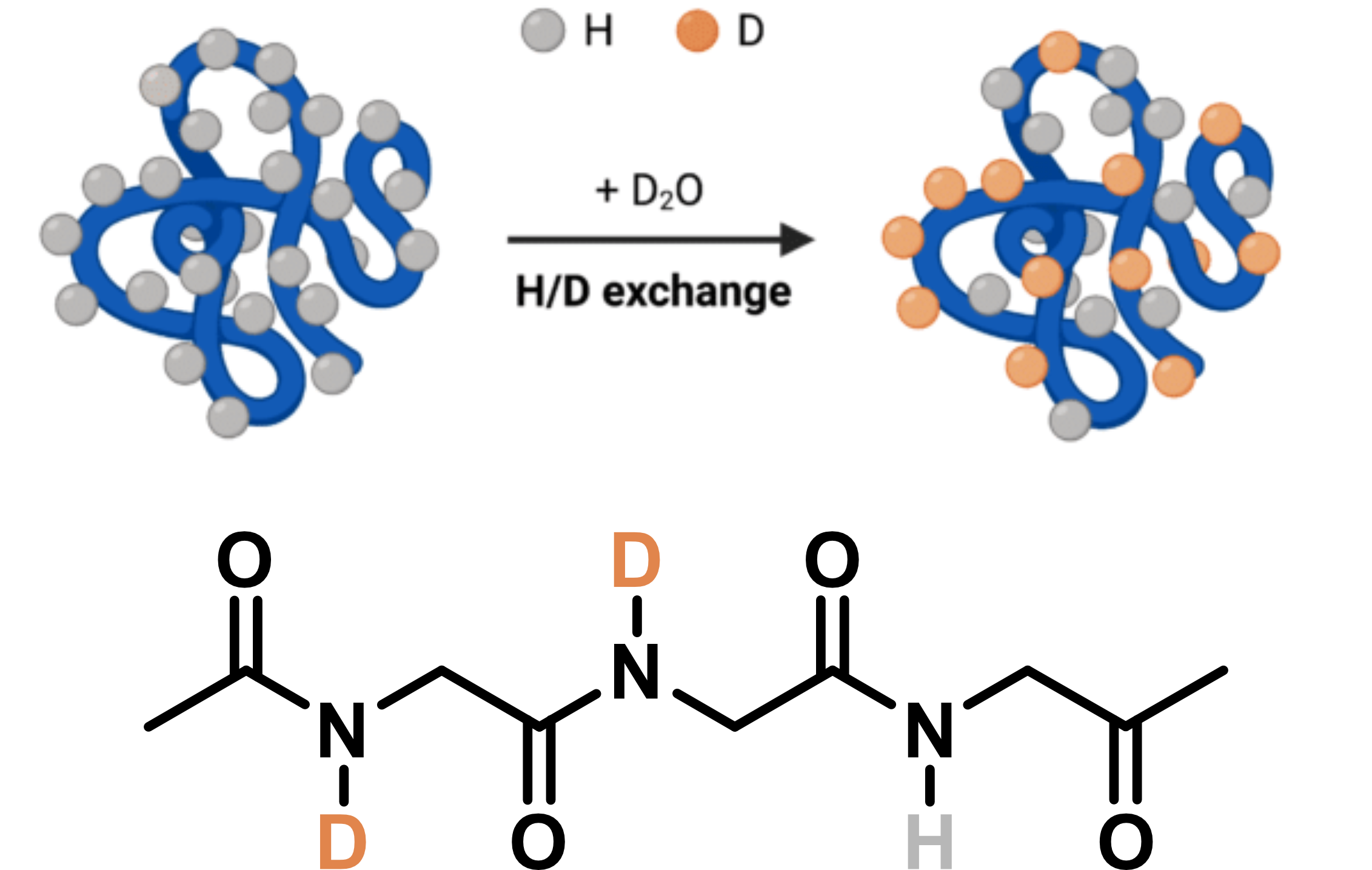
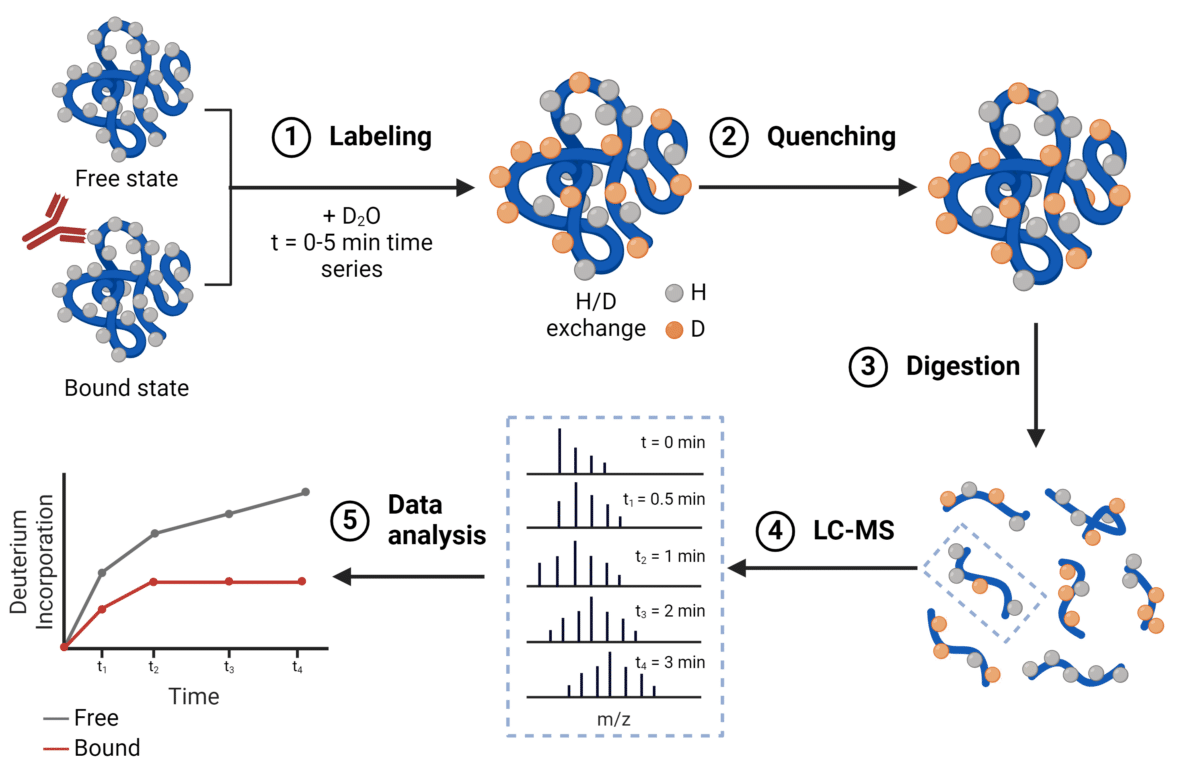

Adapted from Wang, Y. © 2024 Rapid Novor, Inc.
HDX/MS allows epitope mapping
in regulated environments
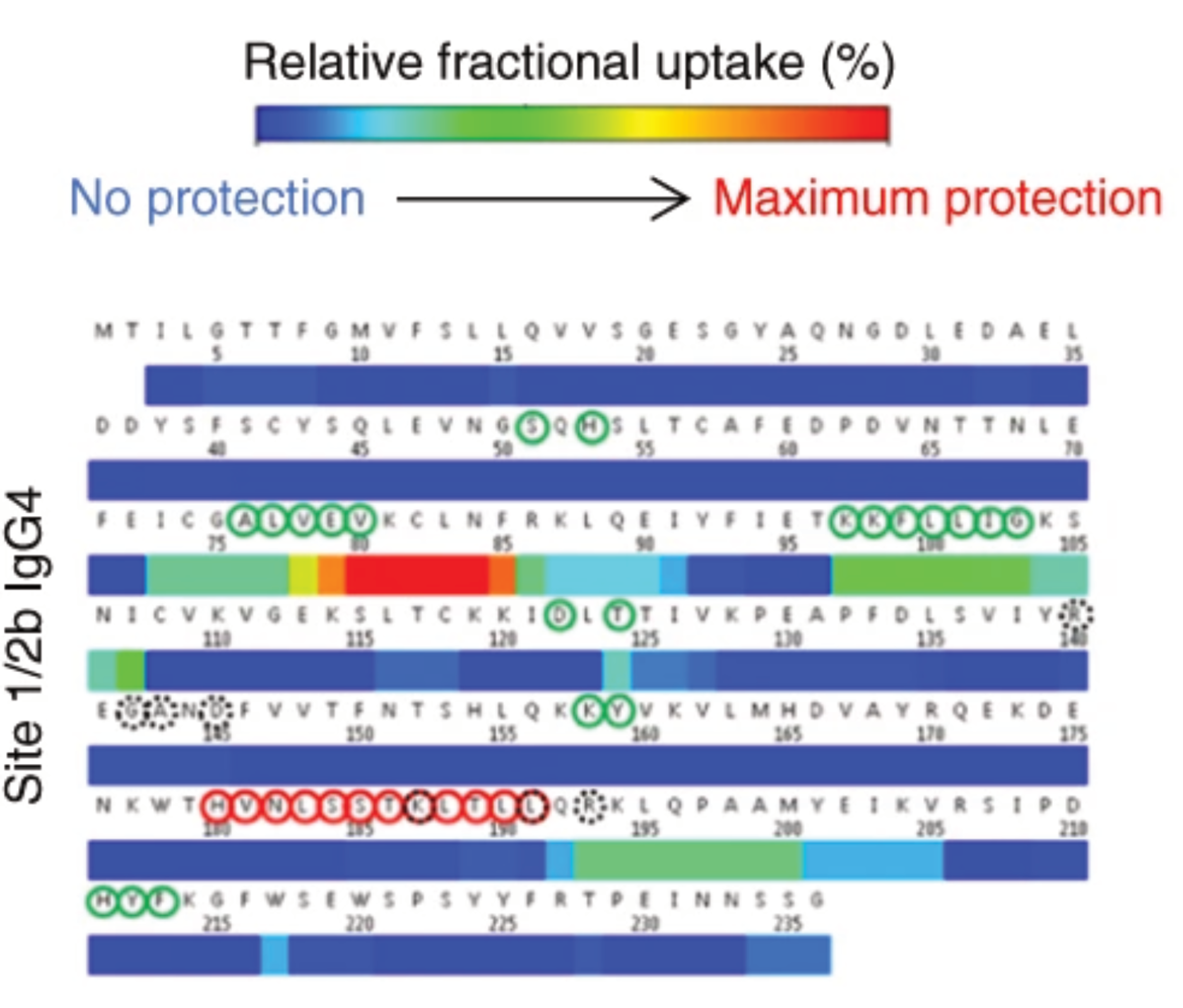
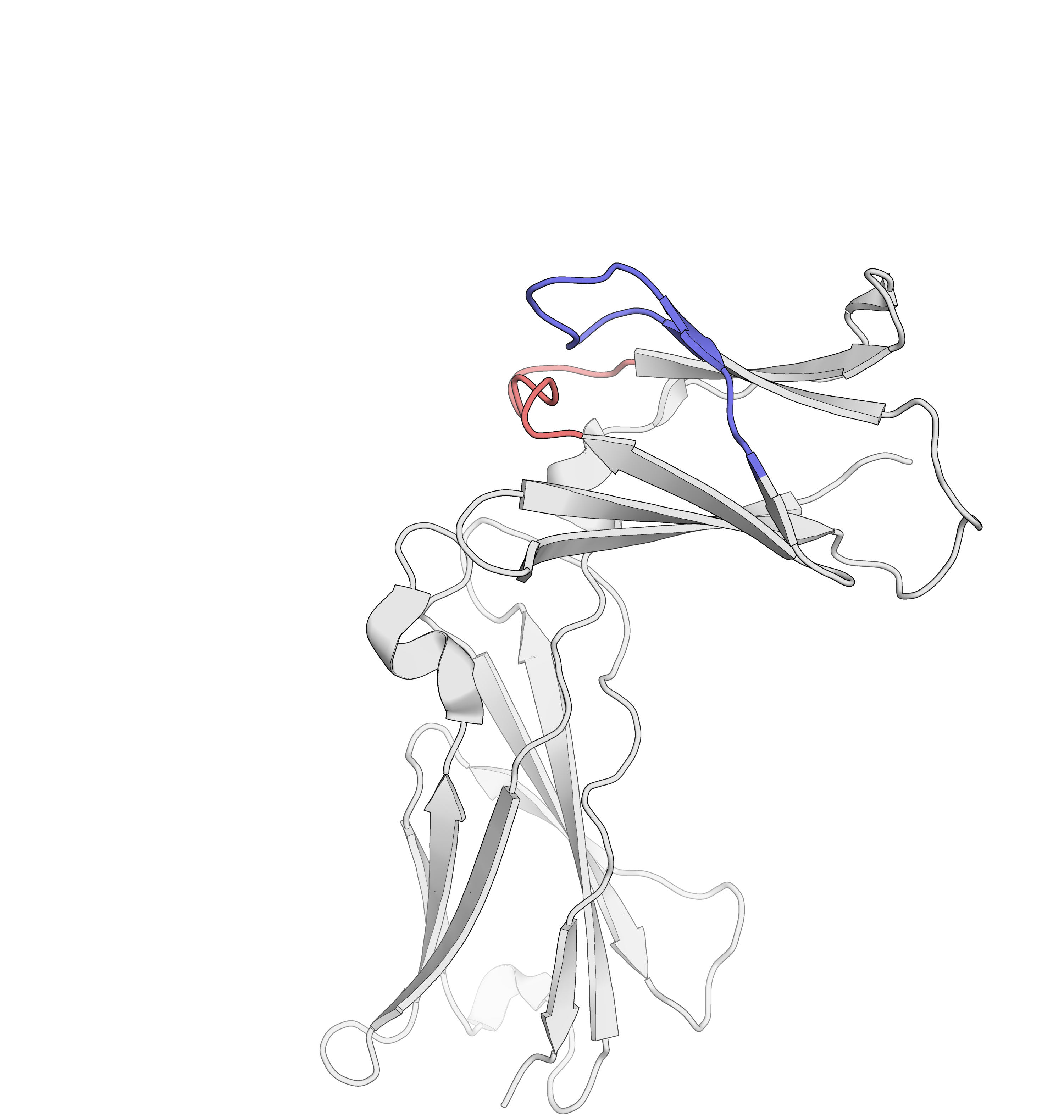
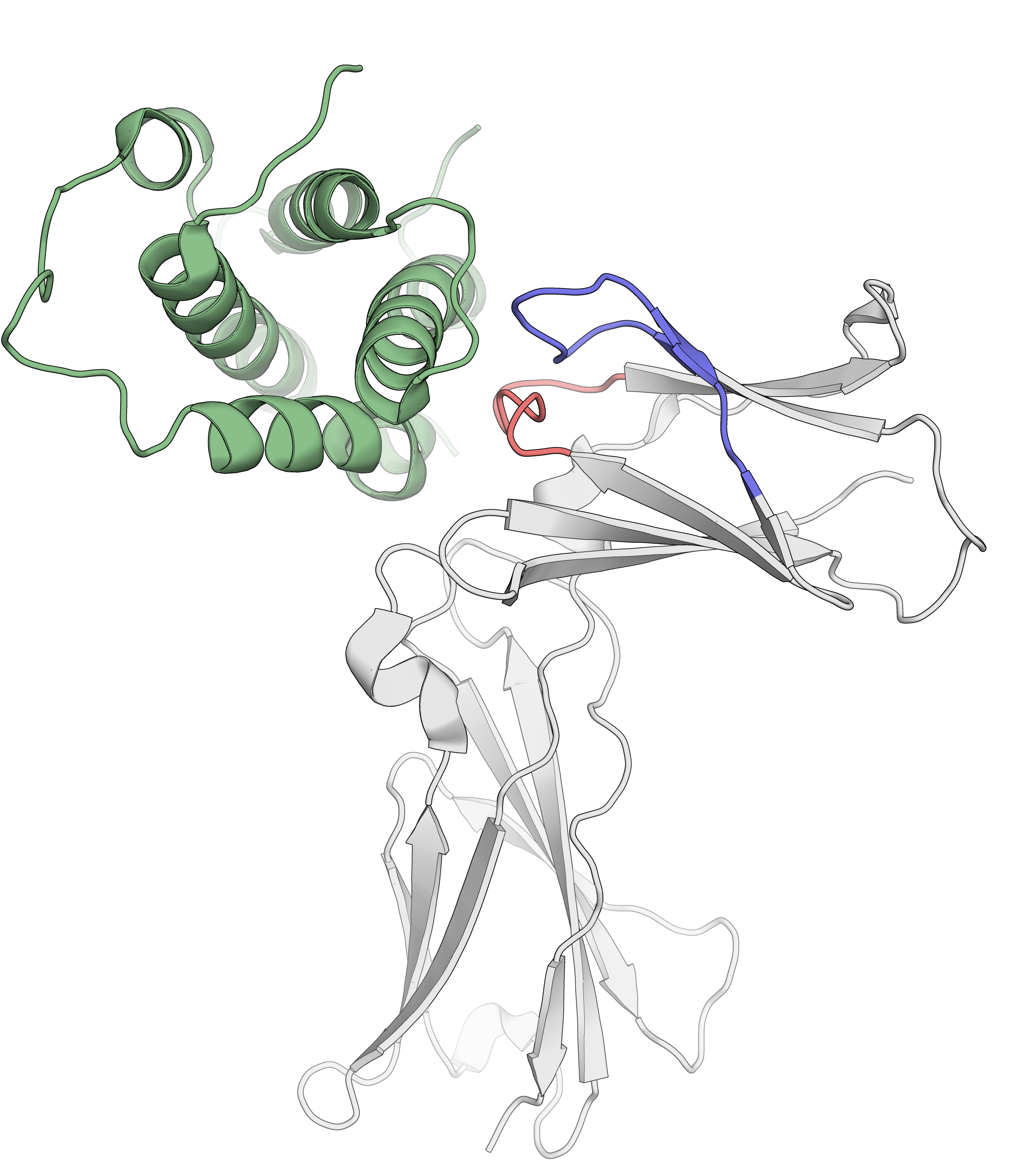
Belarif, L. et al. Nature Communications, 2018, 9, 4483
DNA is amenable to HDX/MS
- H/D exchange ideal for structure probing
- Full sequence coverage
- Directly involved in folding
- Minimal primary/secondary structure change
- Rate of HDX should be structure dependent
- Native MS measurements
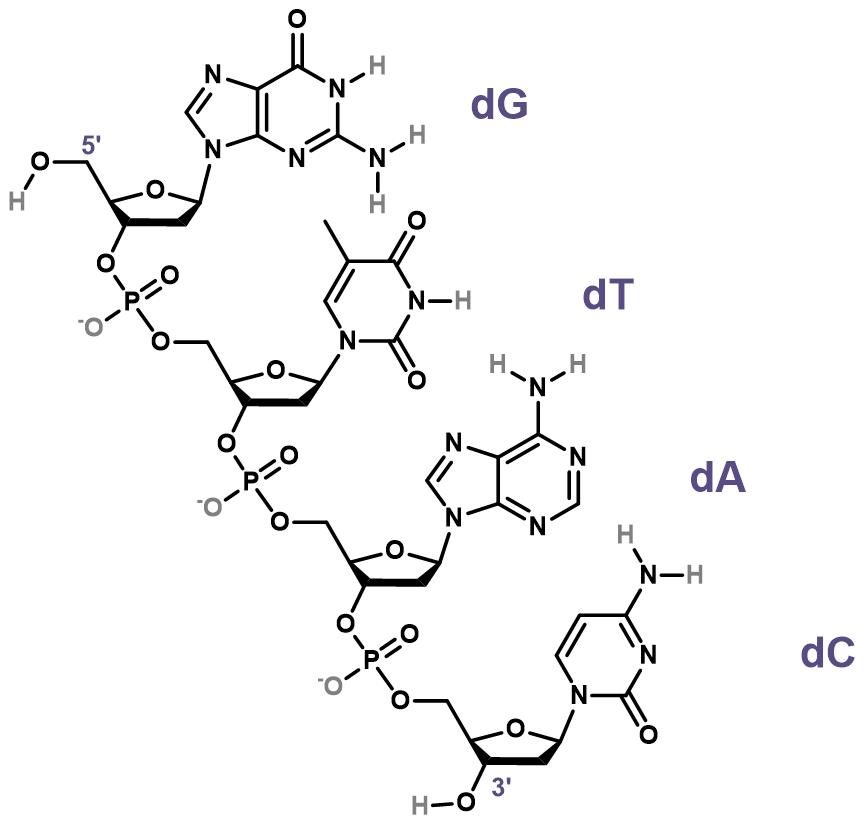

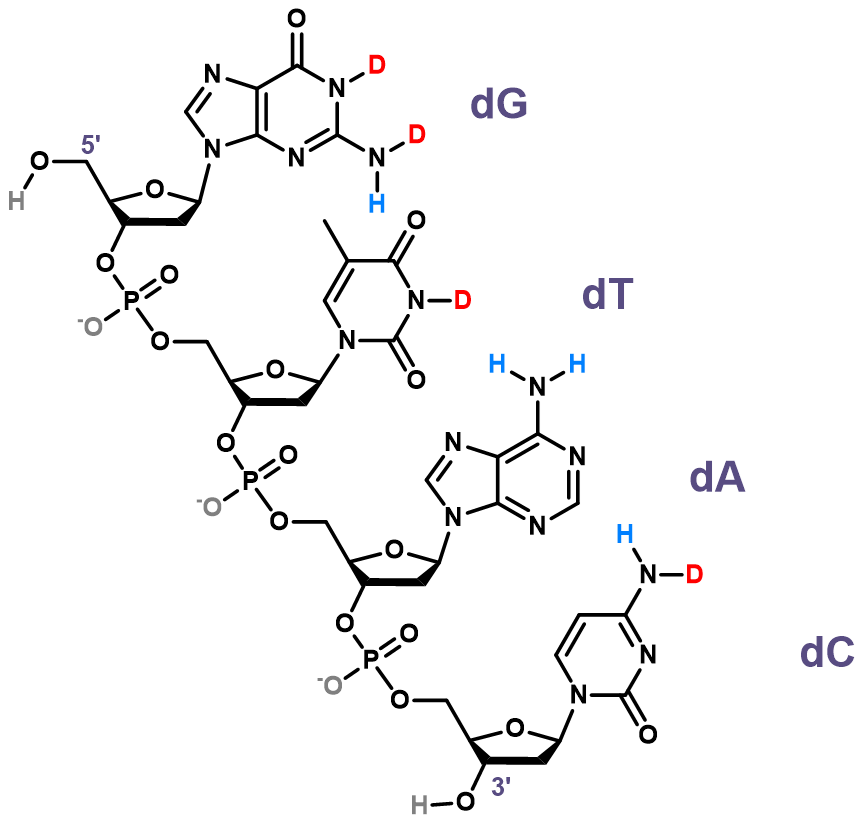

Largy and Gabelica, Anal. Chem., 2020, 92 (6), 4402–10
Coupling in-solution HDX to native MS
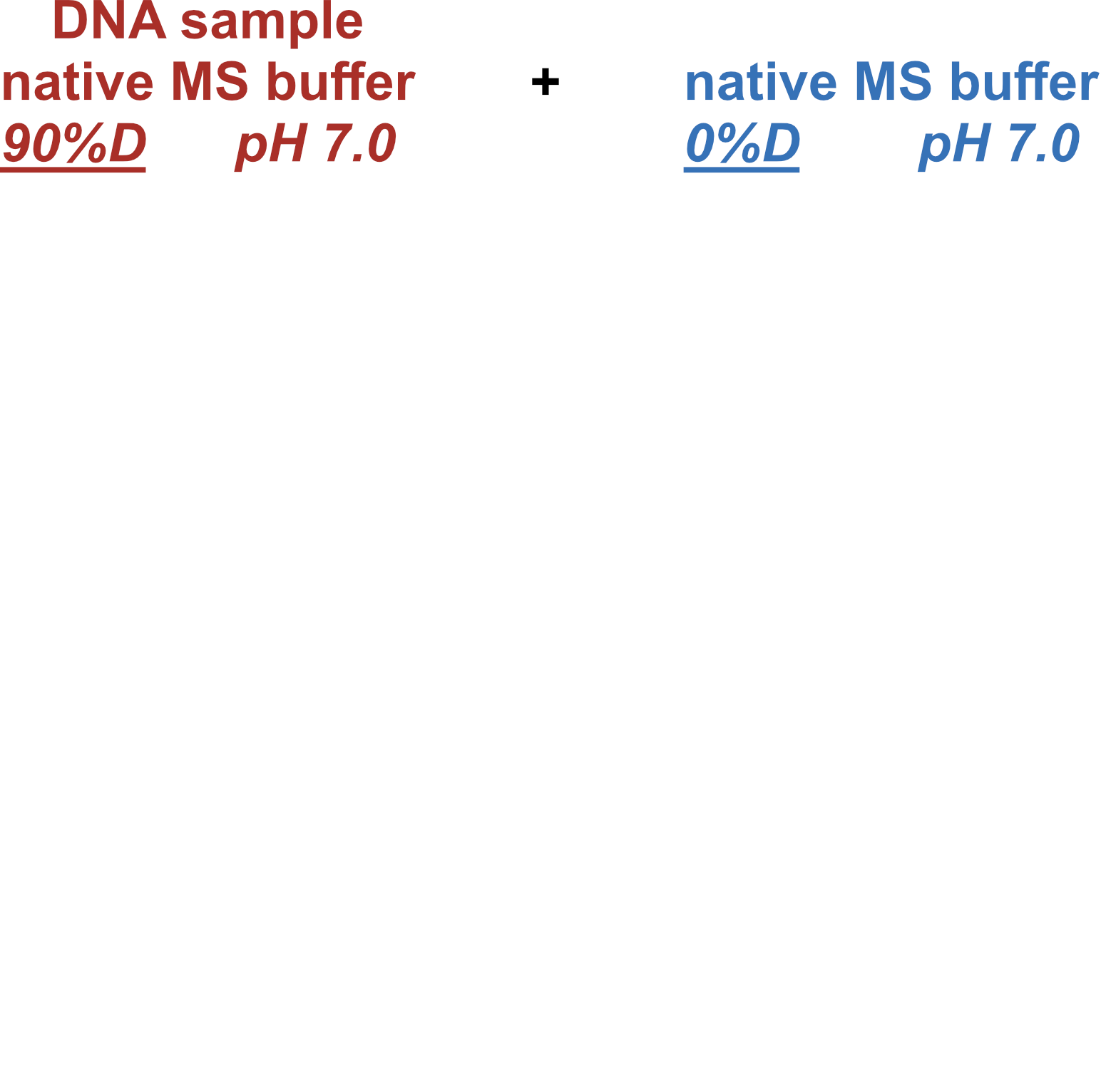

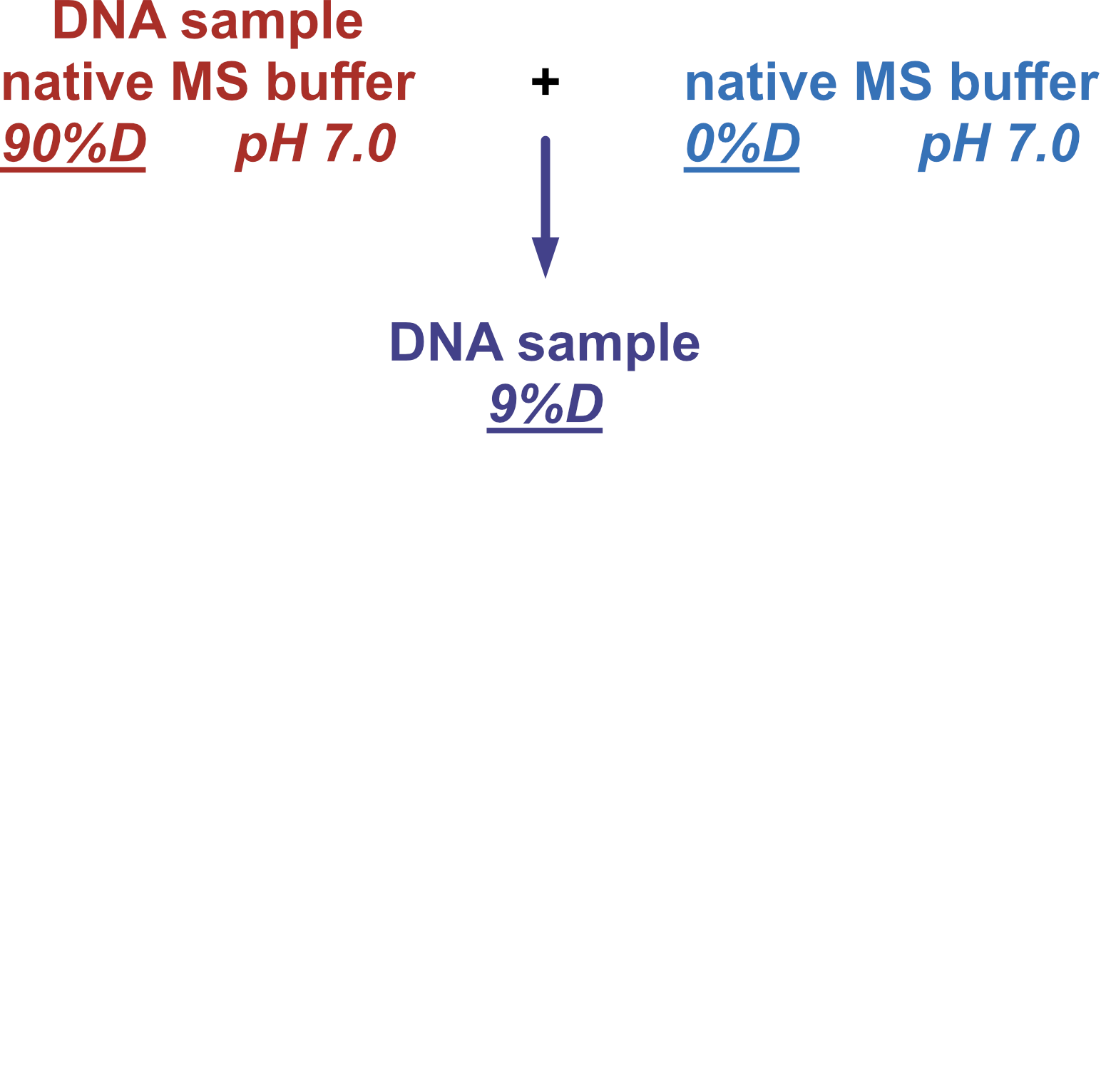
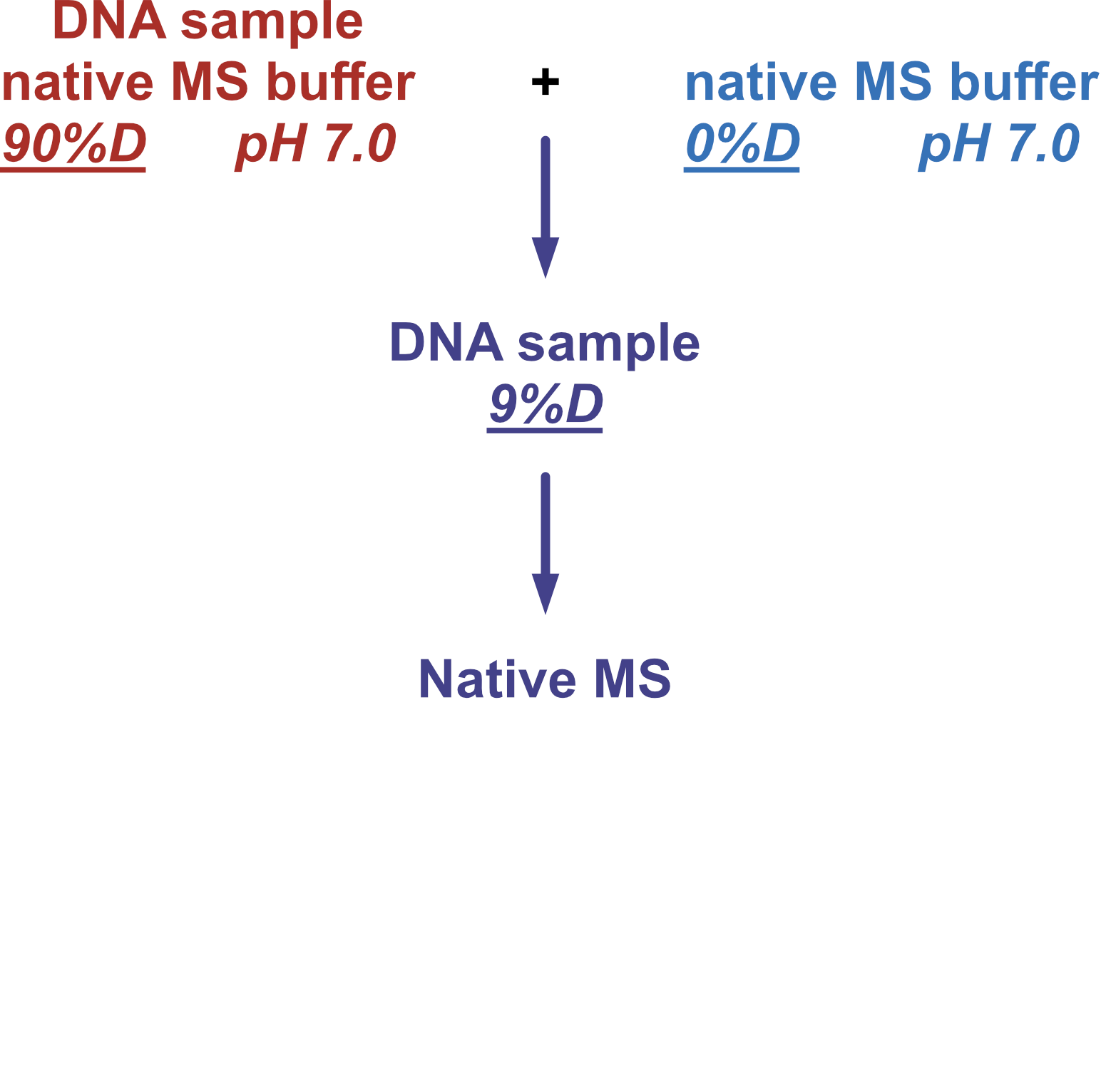
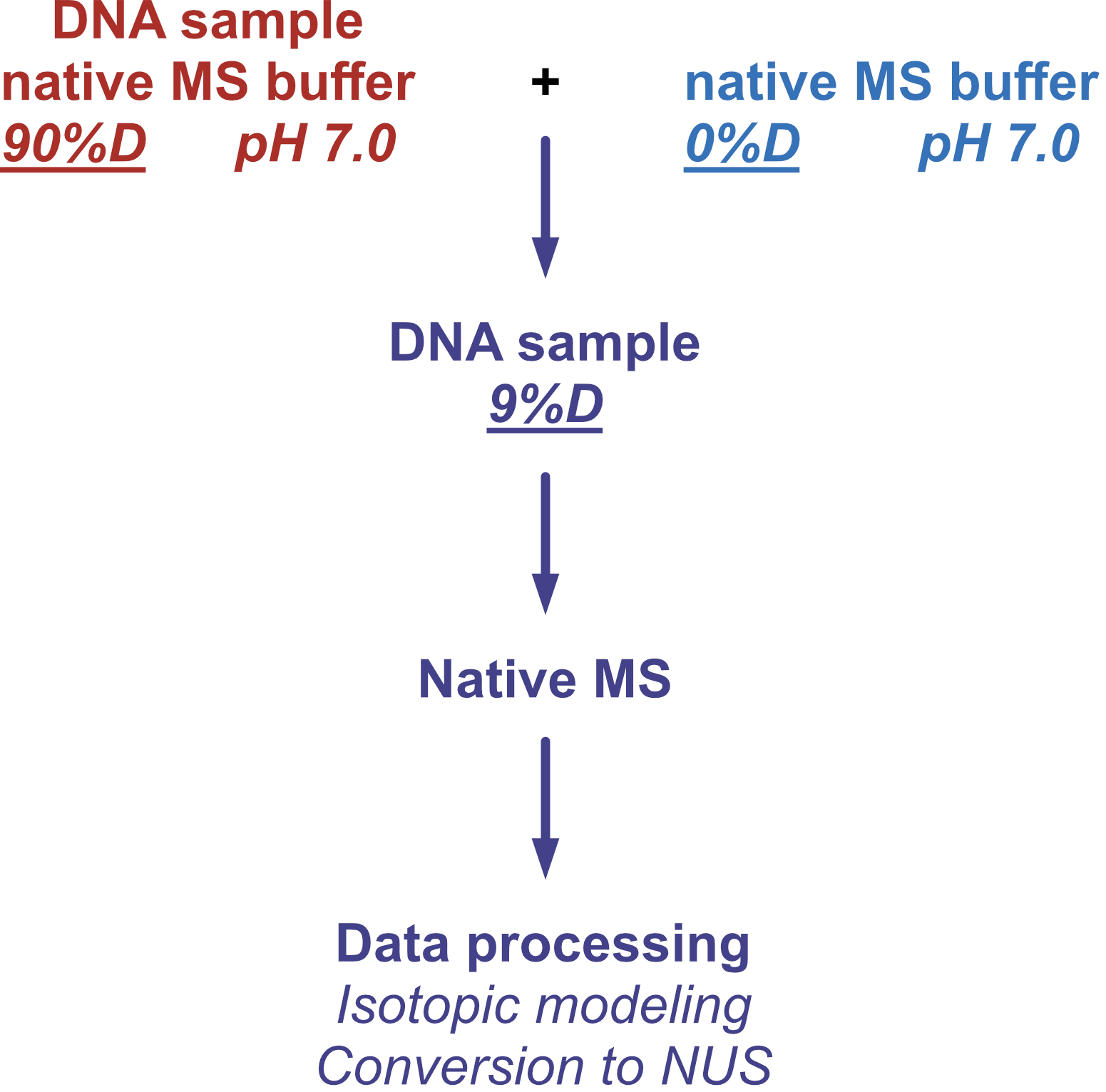
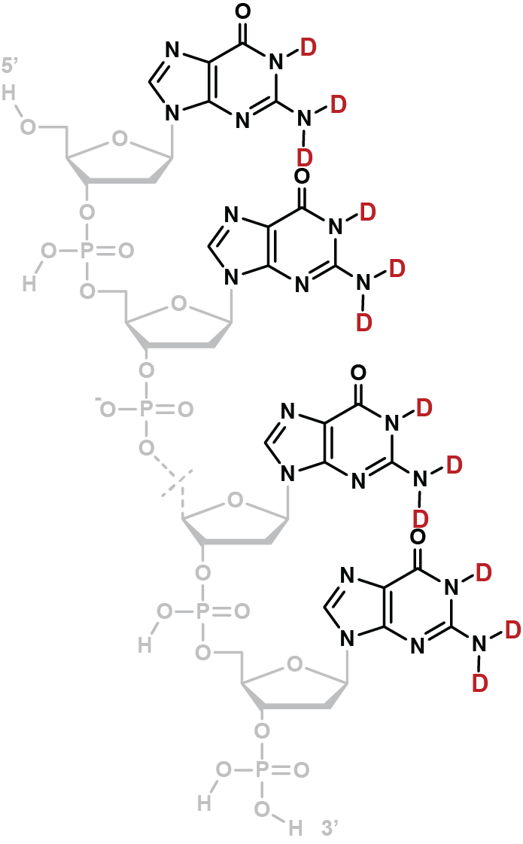
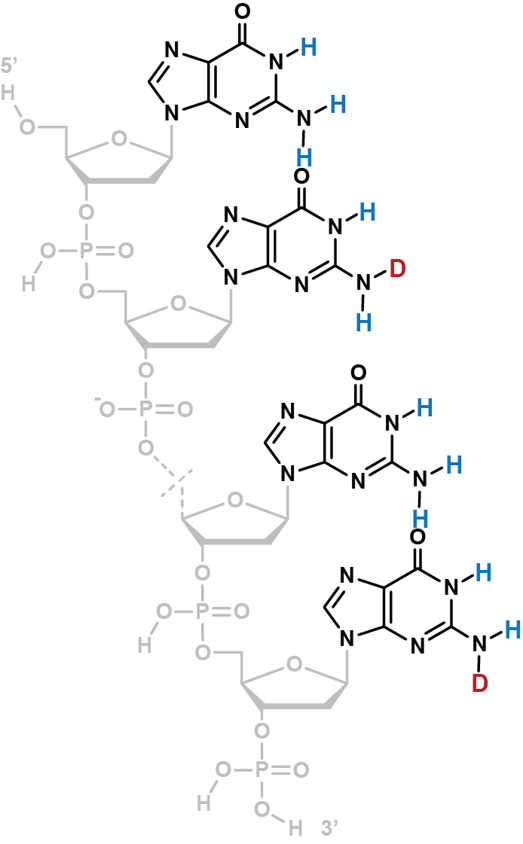
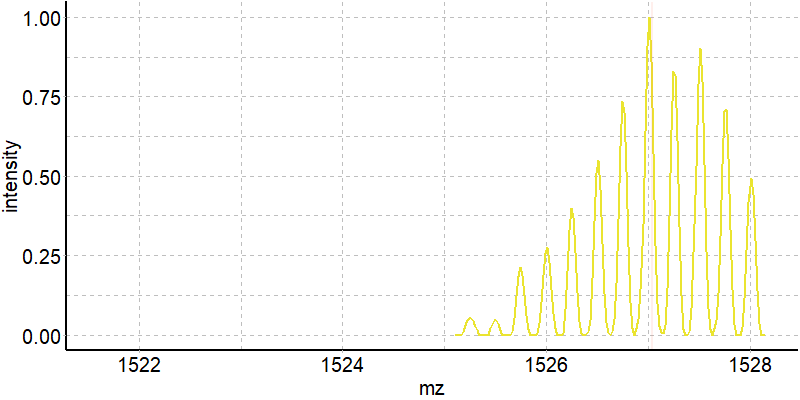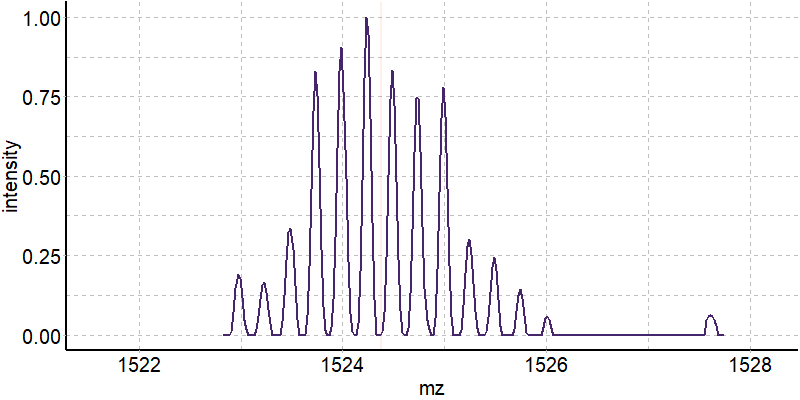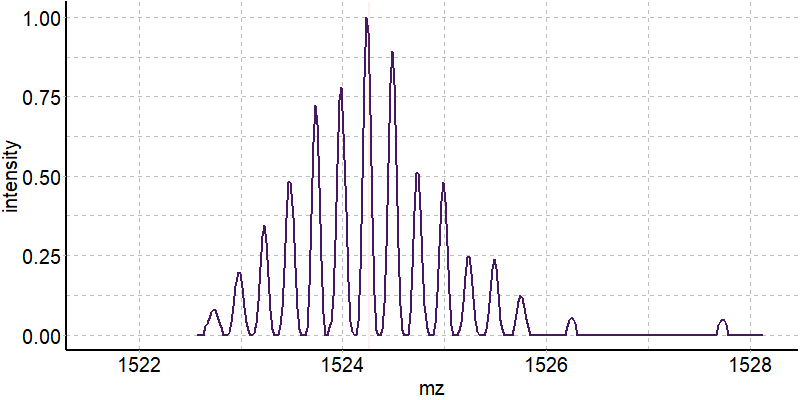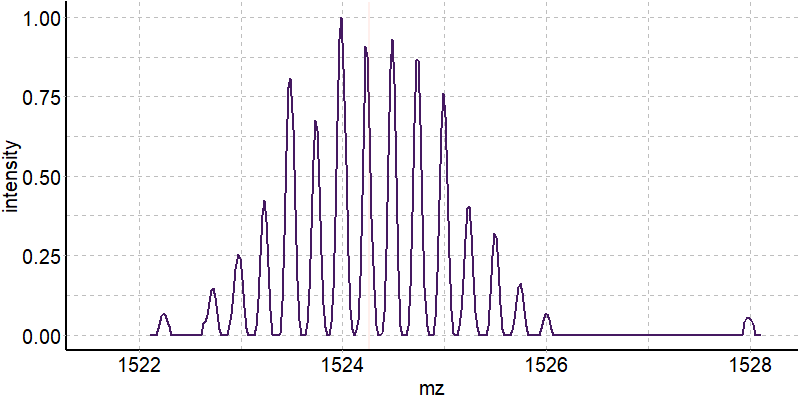
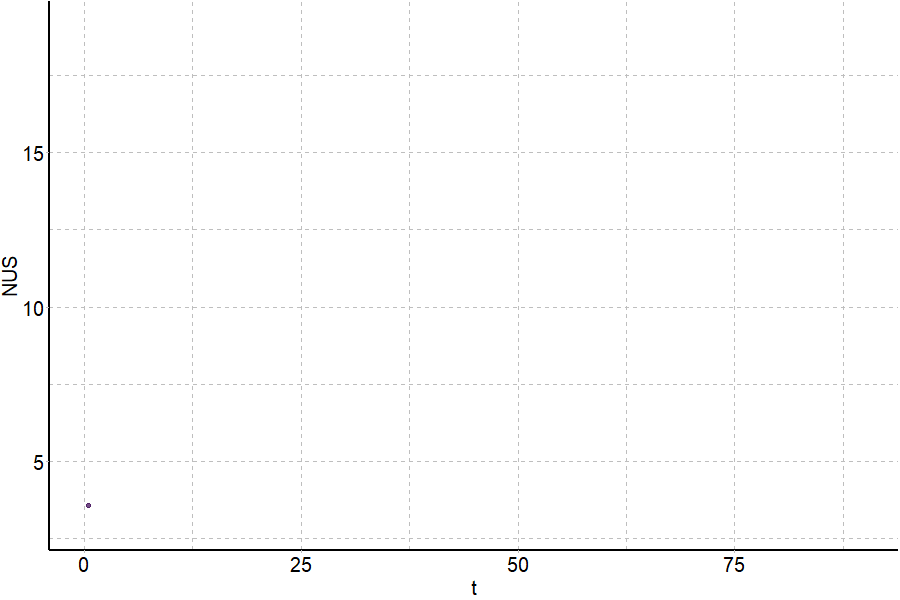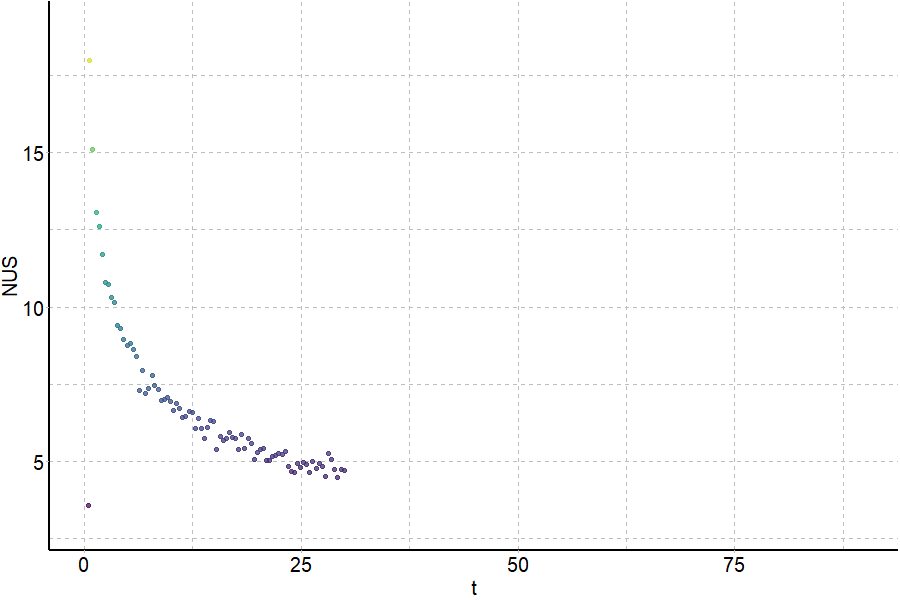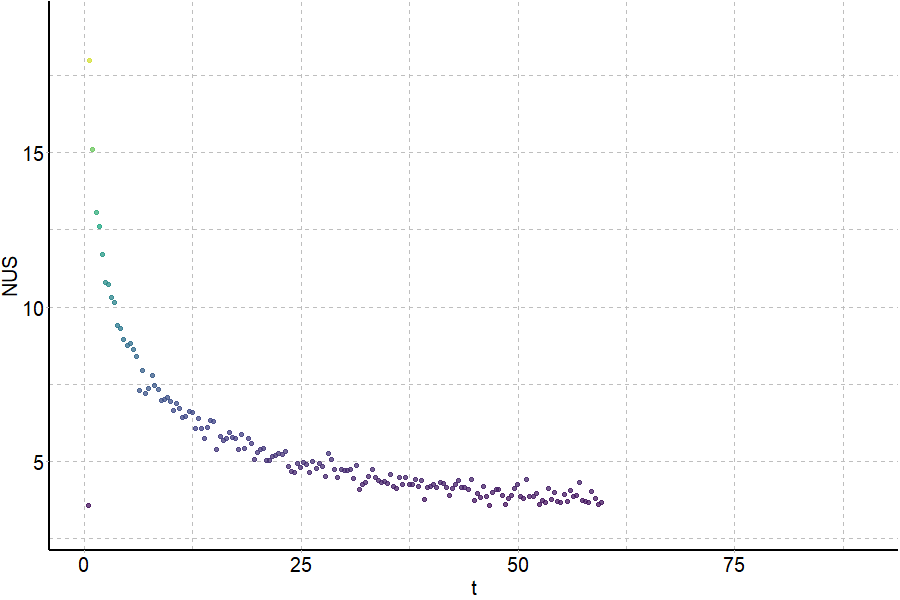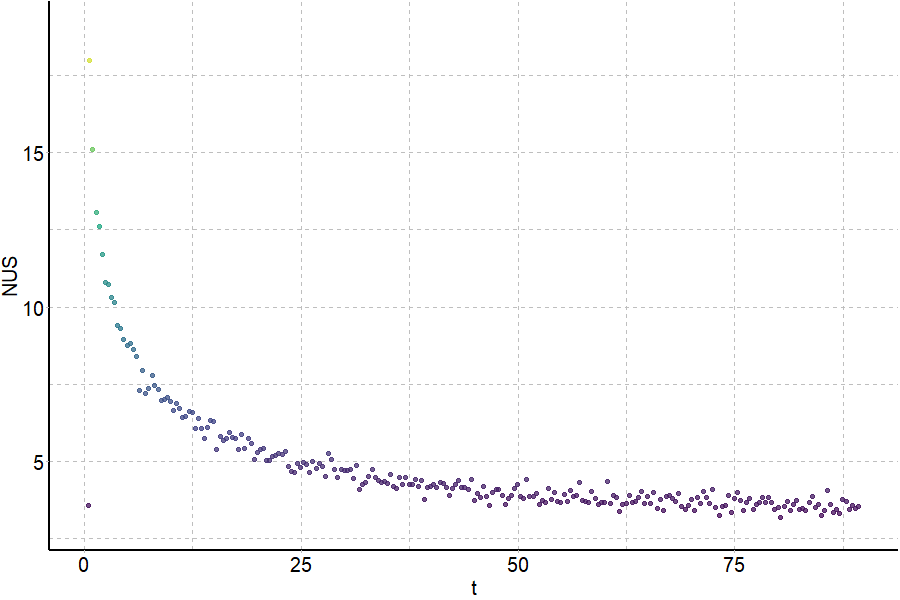
Largy and Gabelica, Anal. Chem., 2020, 92 (6), 4402–10
Largy and Ranz, Anal. Chem., 2023, 95, 9615–22
Exchange rates are sensitive to folding
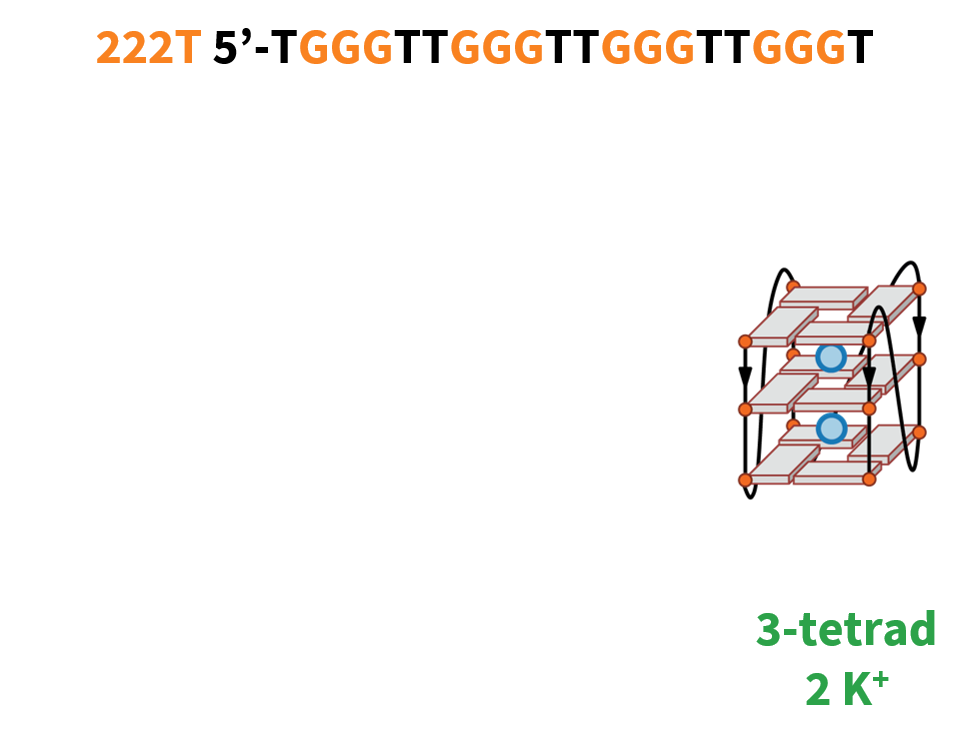
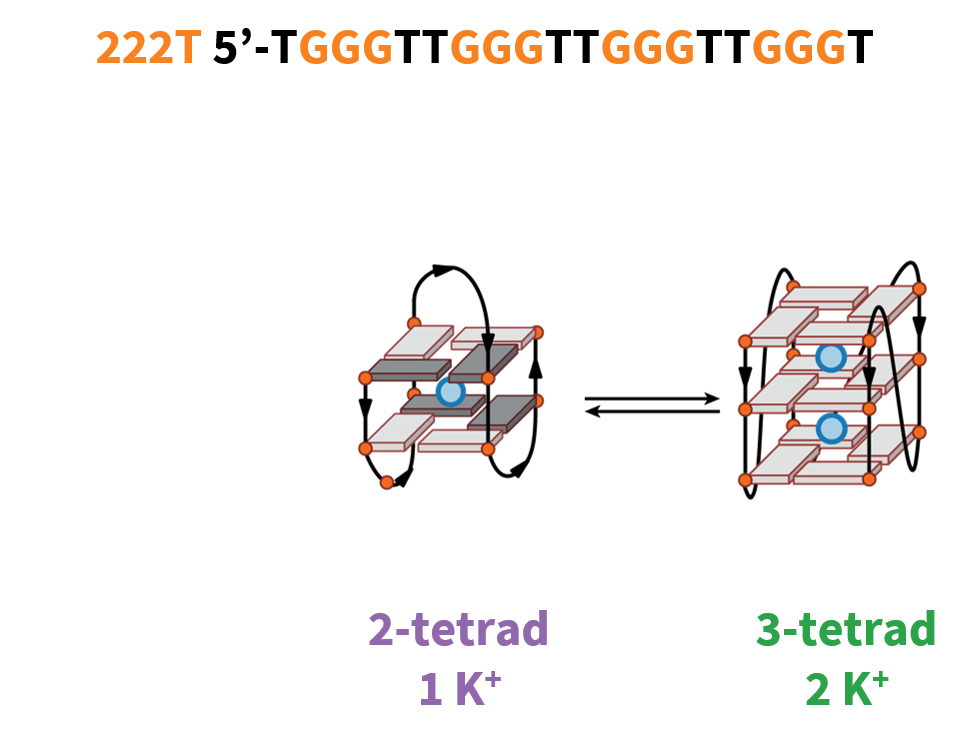

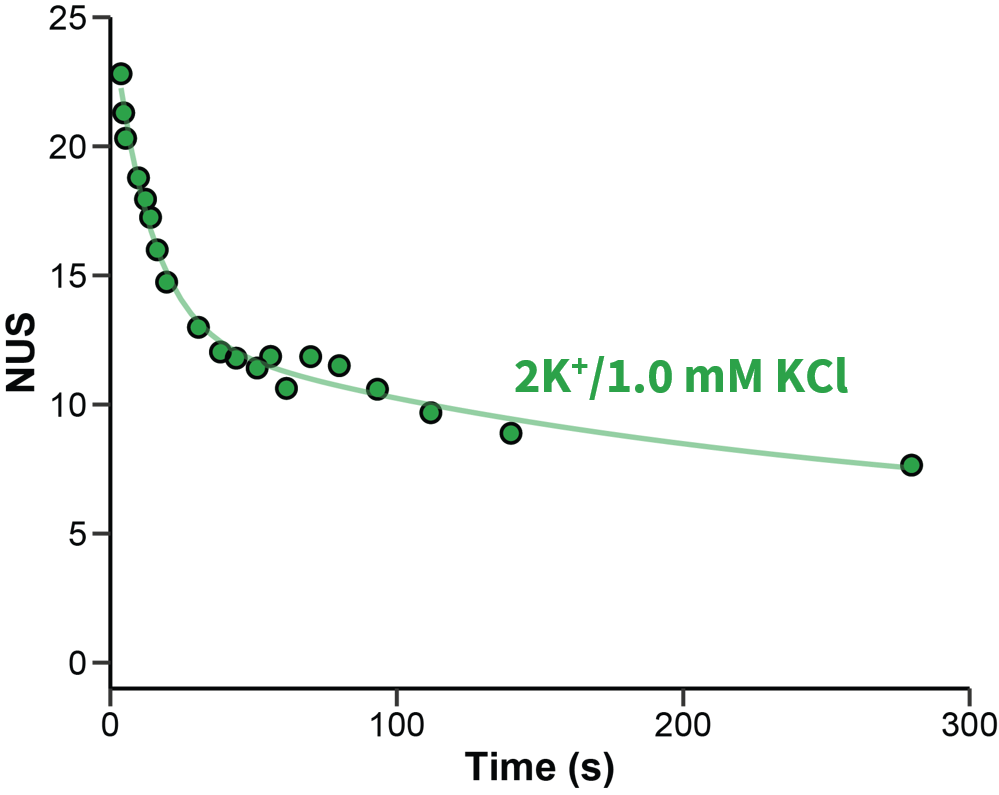
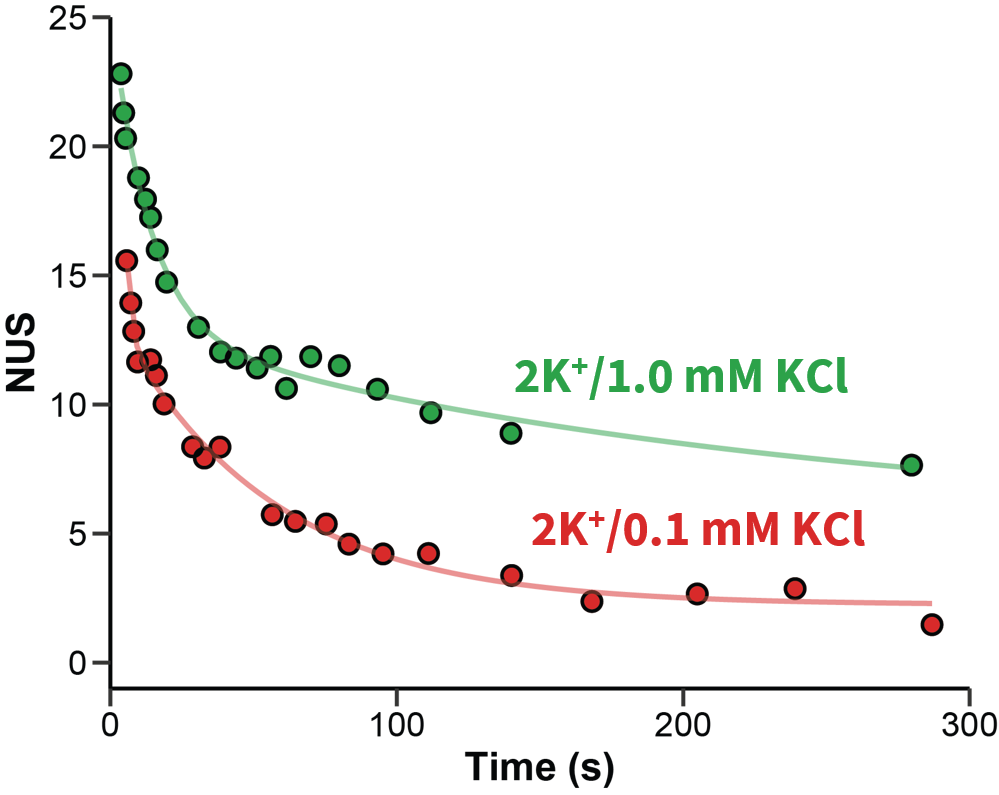
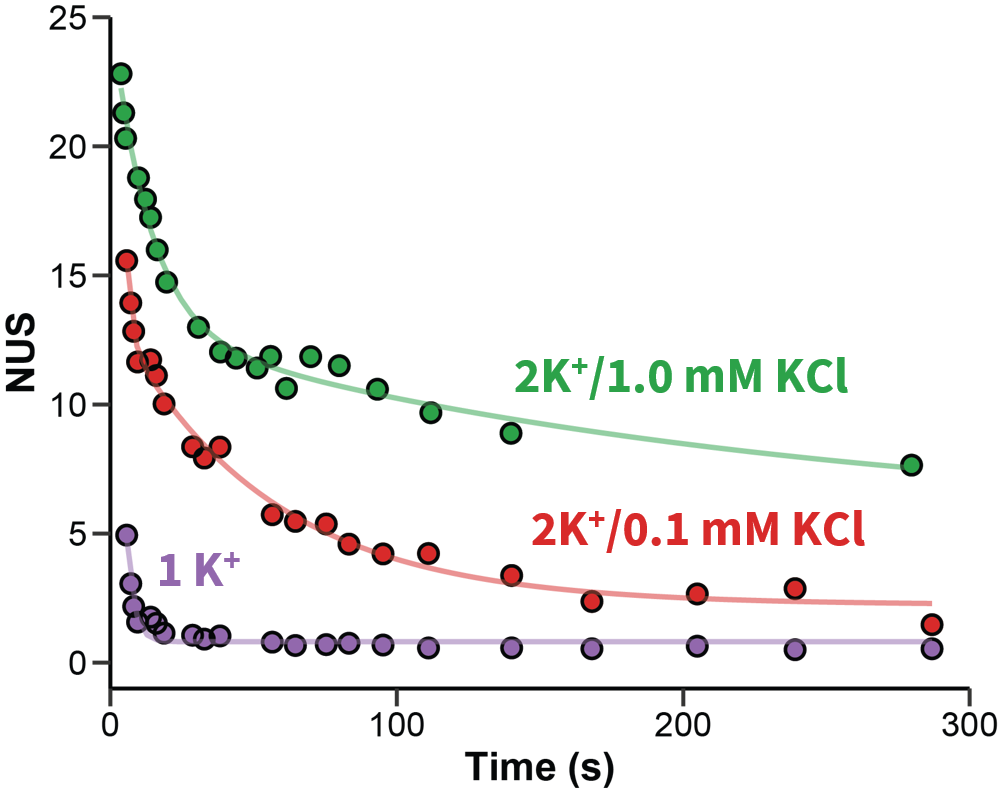
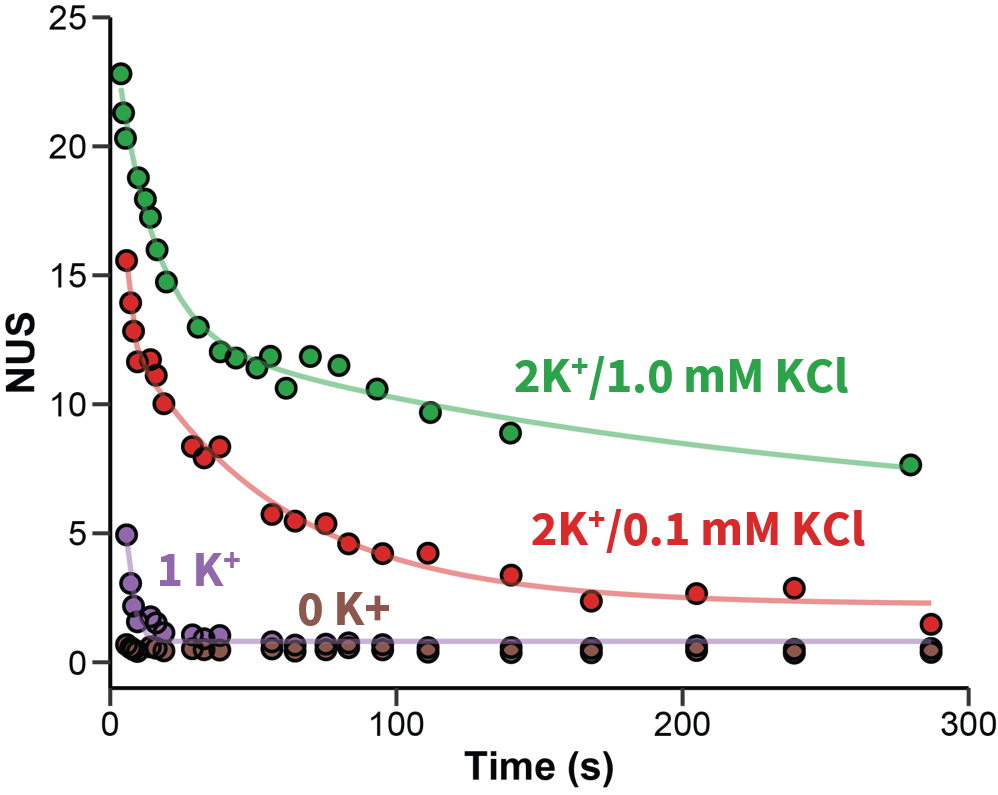
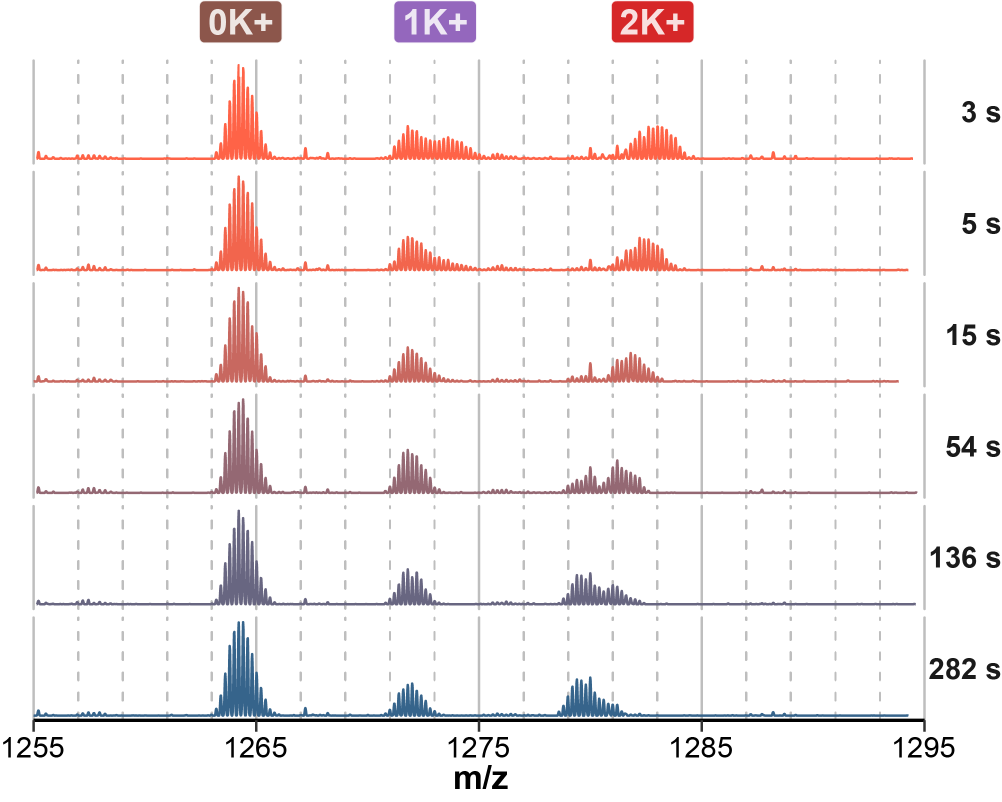
Largy and Gabelica, Anal. Chem., 2020, 92 (6), 4402–10
The number of protected sites scales
with the number of tetrads
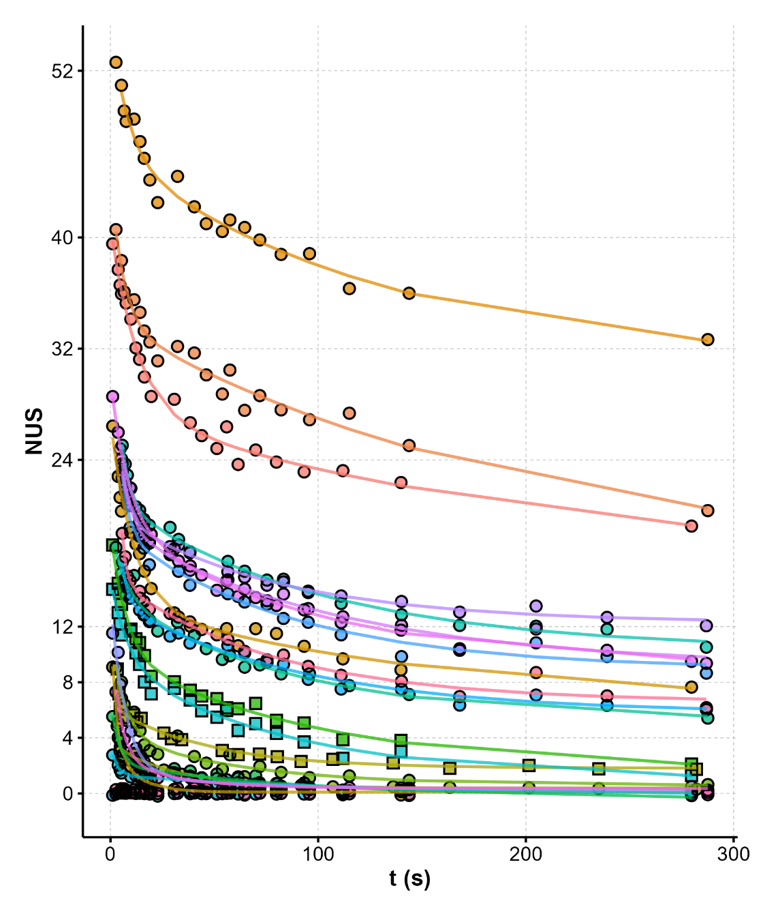
Largy, Ranz, Gabelica, J. Am. Chem. Soc. 2023, 145, 49, 26843
The number of protected sites scales
with the number of tetrads
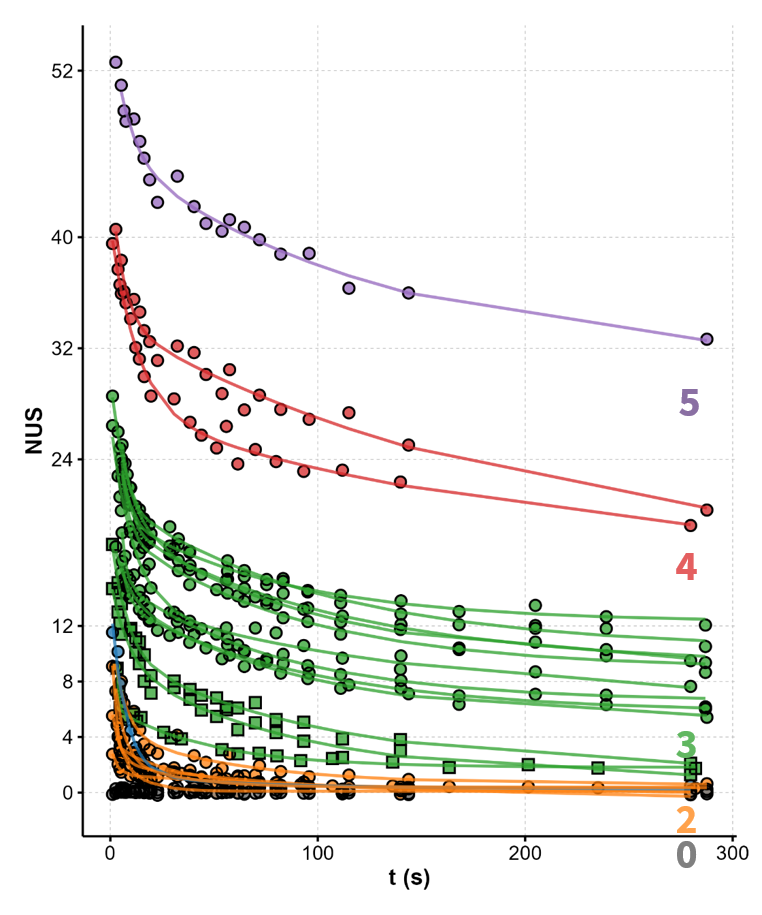
Largy, Ranz, Gabelica, J. Am. Chem. Soc. 2023, 145, 49, 26843
The number of protected sites scales
with the number of tetrads

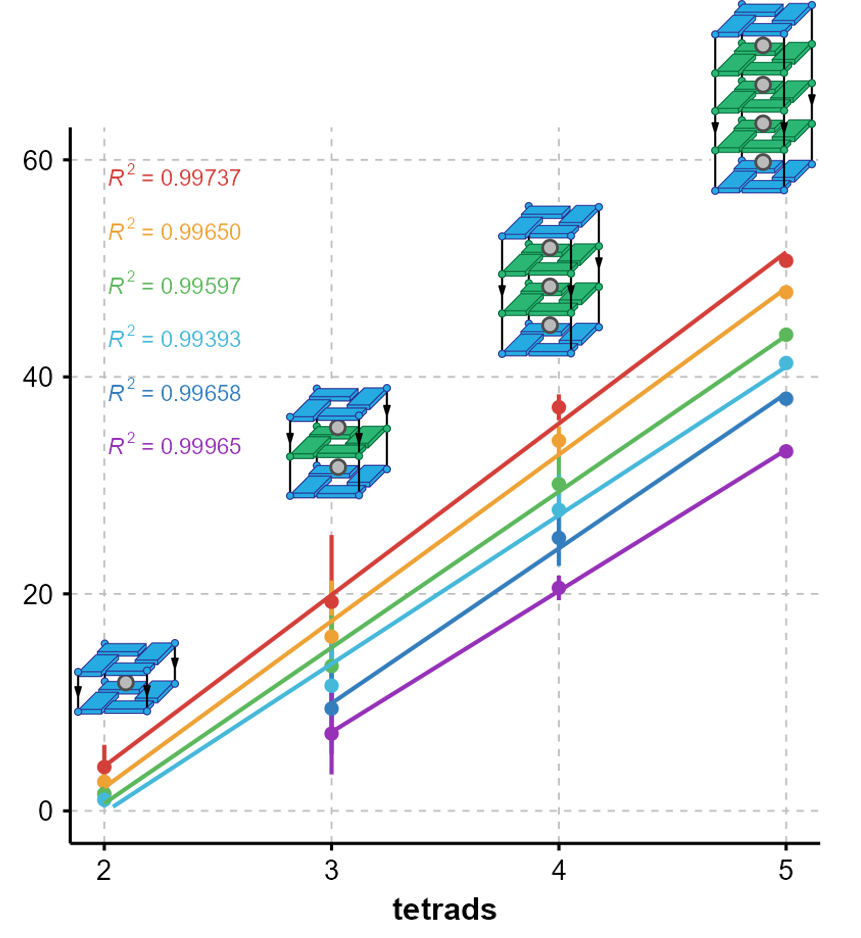
Largy, Ranz, Gabelica, J. Am. Chem. Soc. 2023, 145, 49, 26843
Exchange protection scales with stability
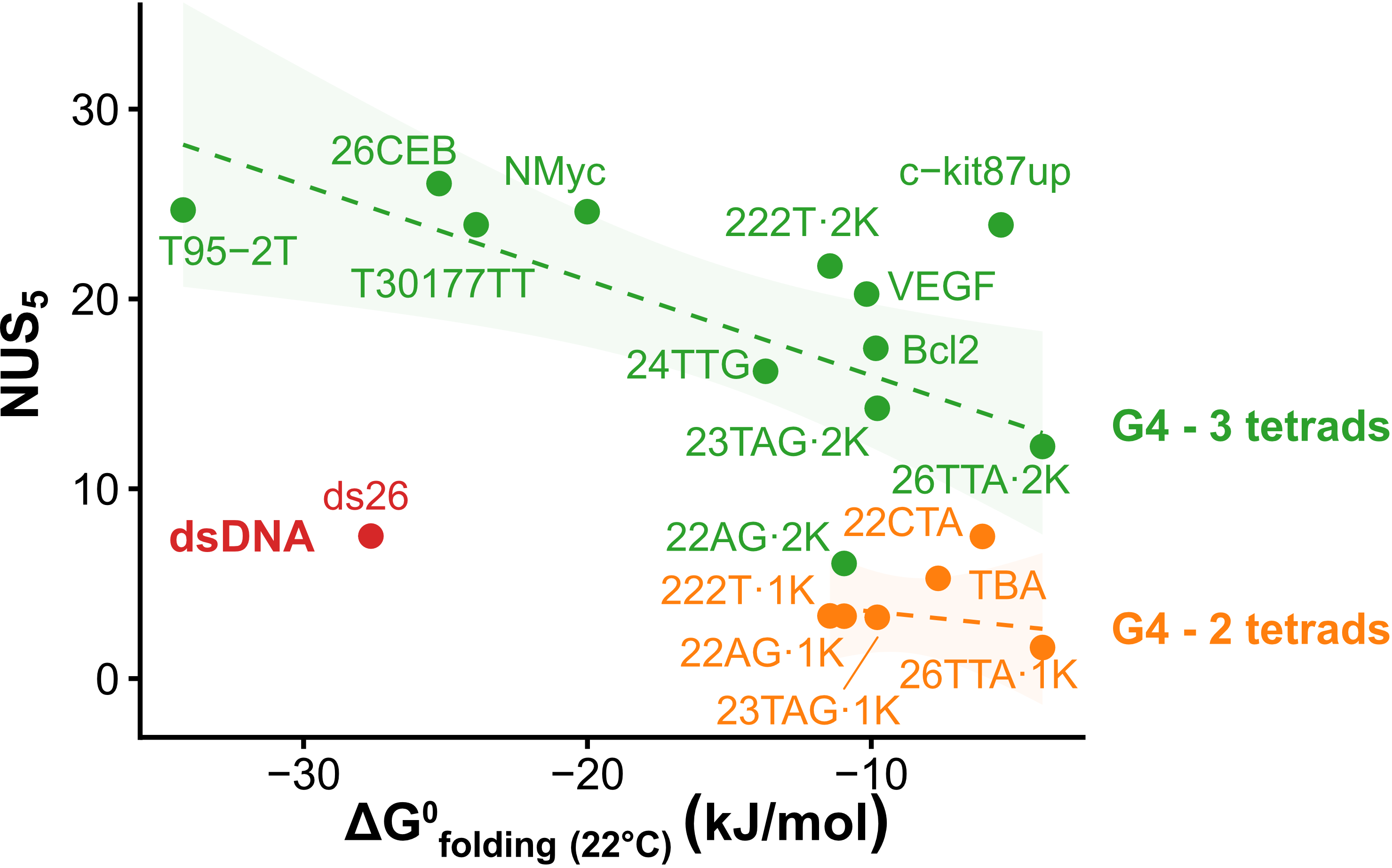
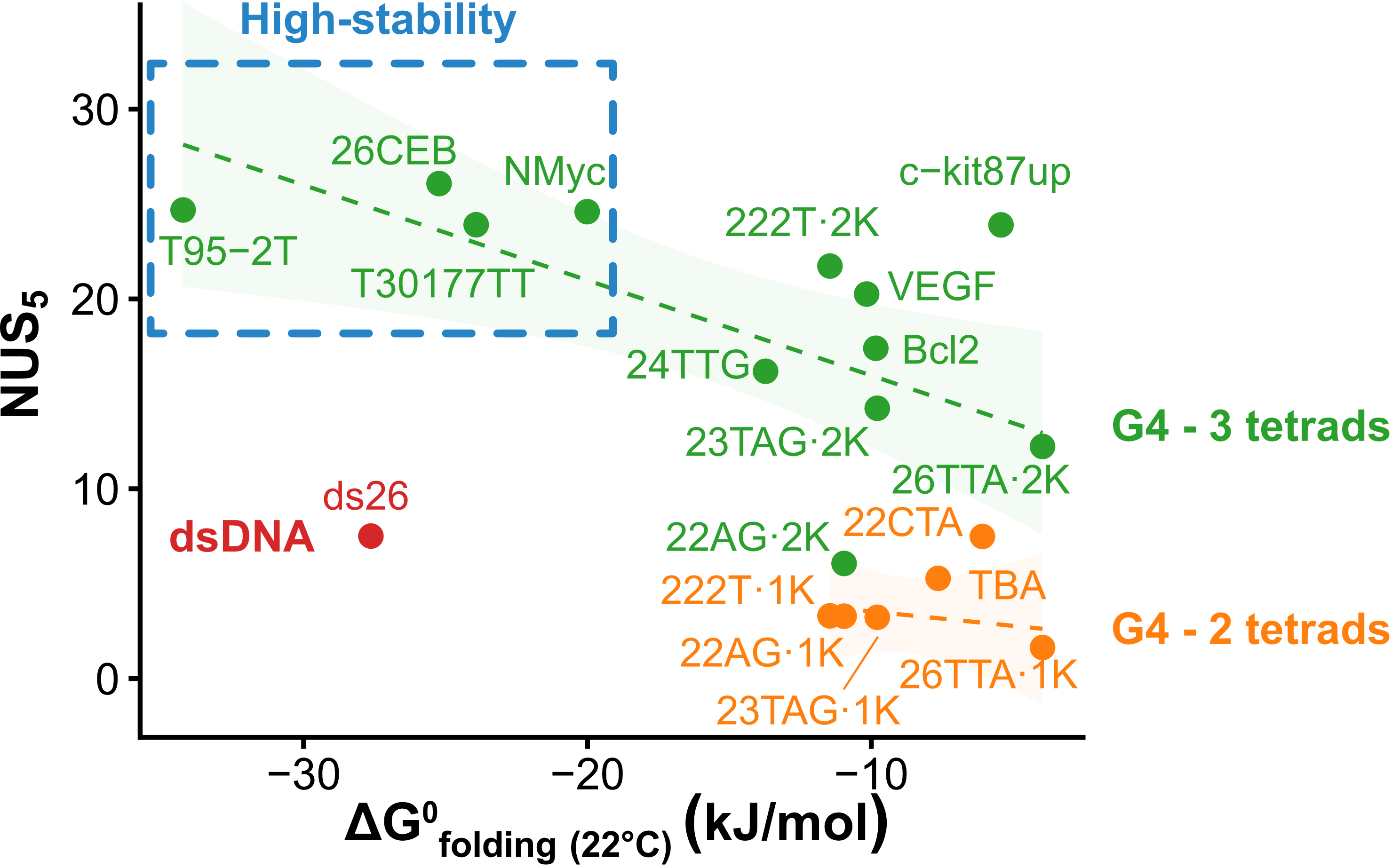
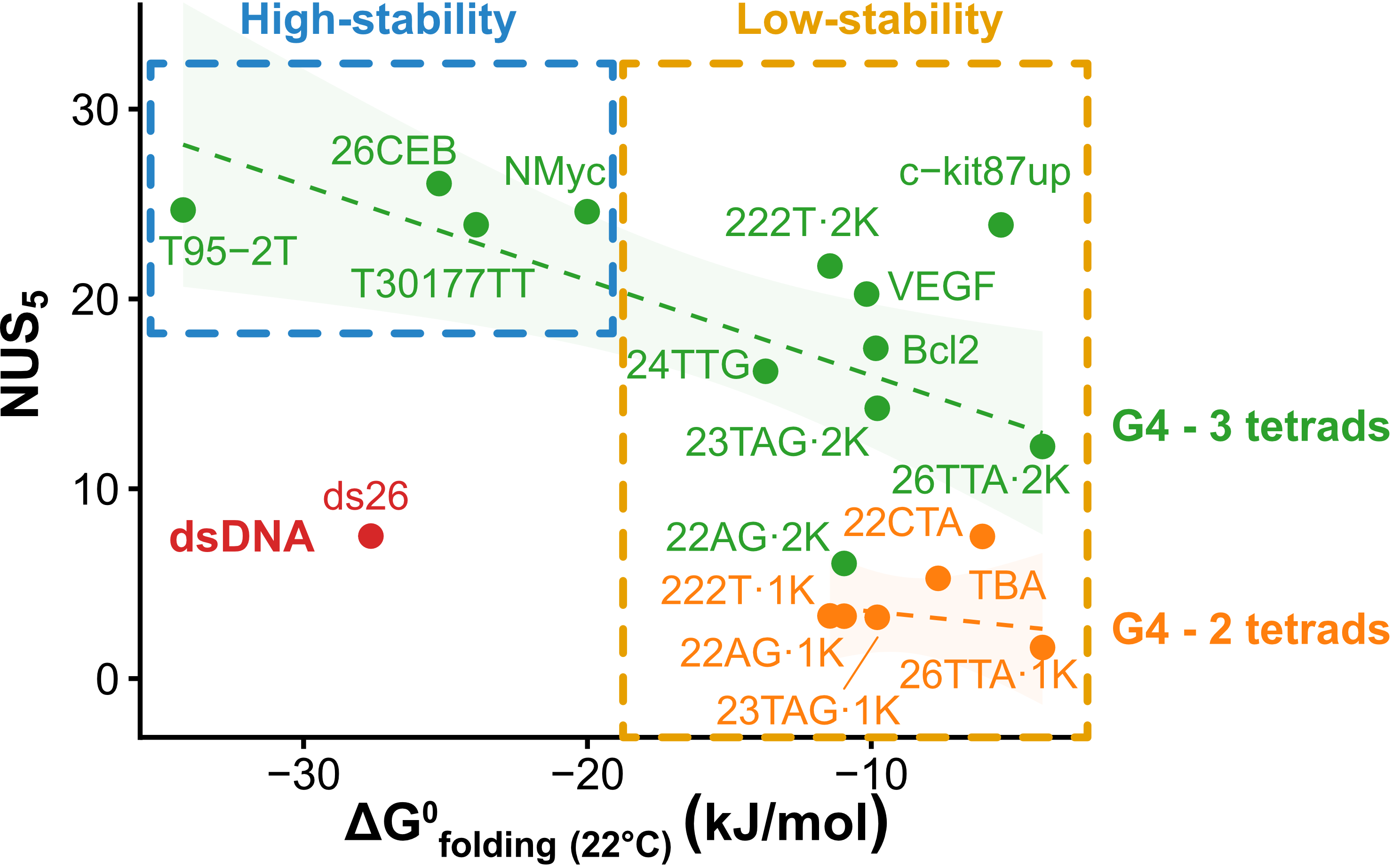
Largy, Ranz, Gabelica, J. Am. Chem. Soc. 2023, 145, 49, 26843
Higher stability G4s exchange via local fluctuations
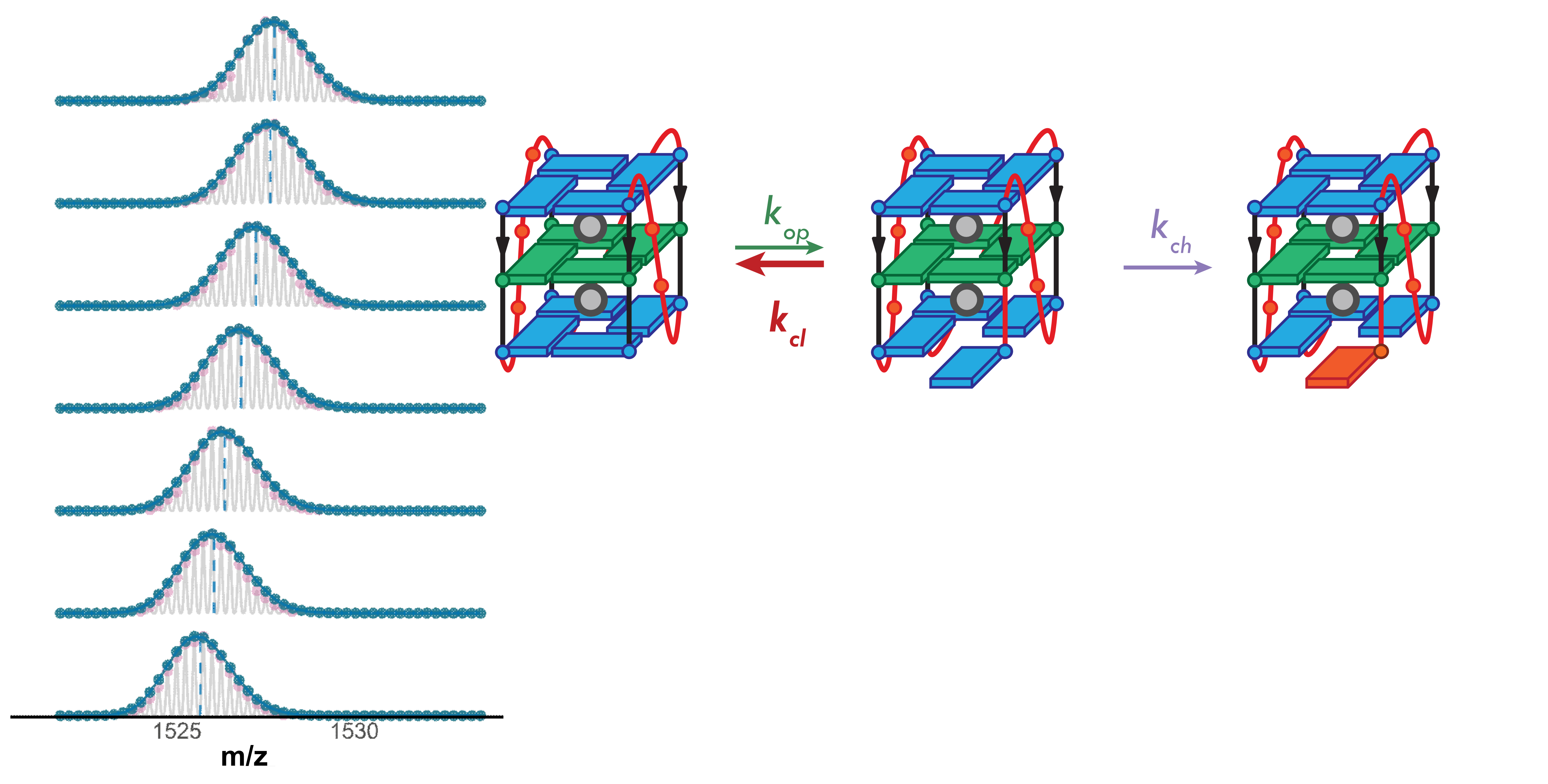
Largy, Ranz, Gabelica, J. Am. Chem. Soc. 2023, 145, 49, 26843
Lower stability G4s exchange via unfolded conformers
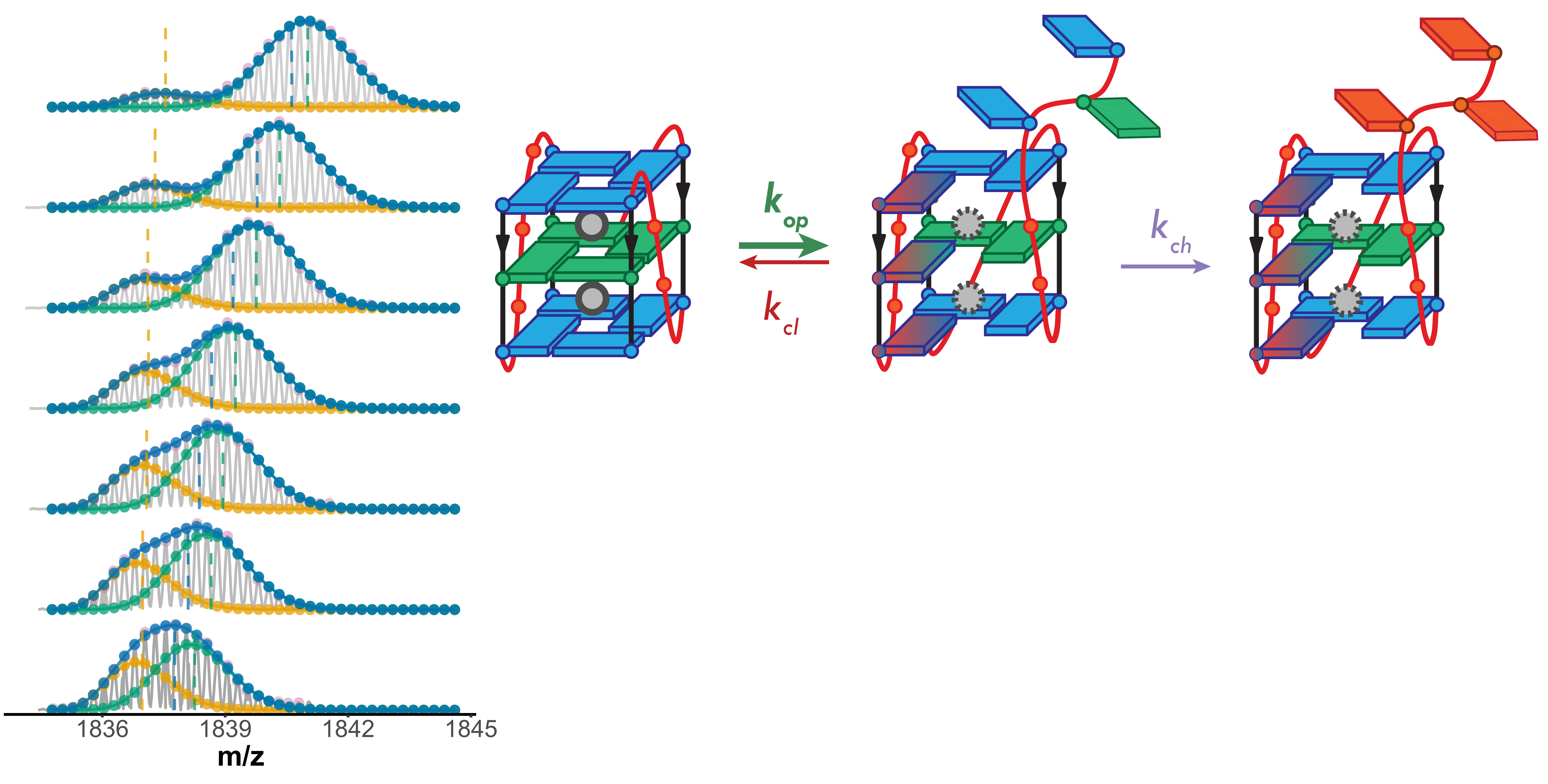
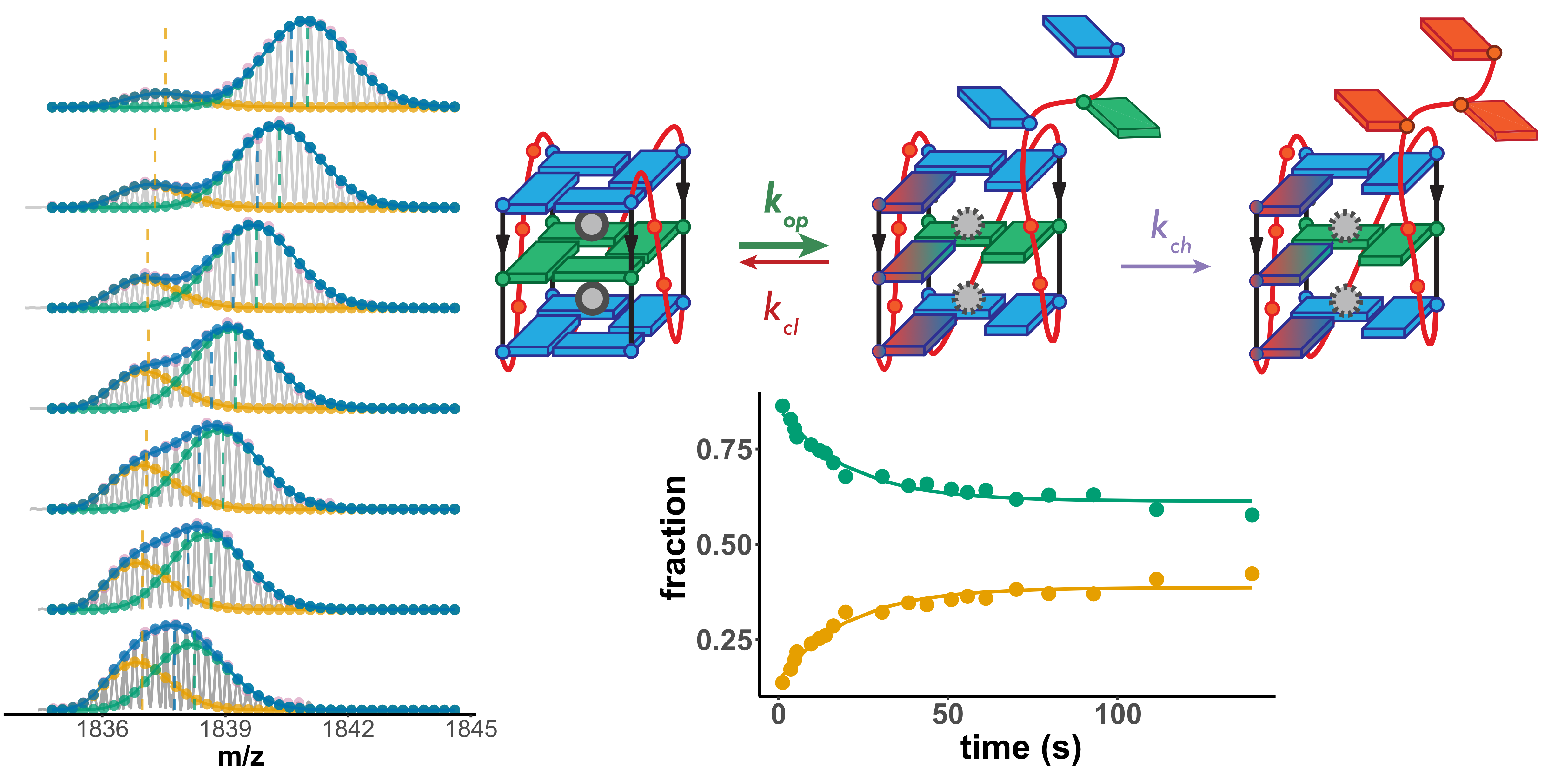
Largy, Ranz, Gabelica, J. Am. Chem. Soc. 2023, 145, 49, 26843
Exchange-competent species
resemble folding intermediates
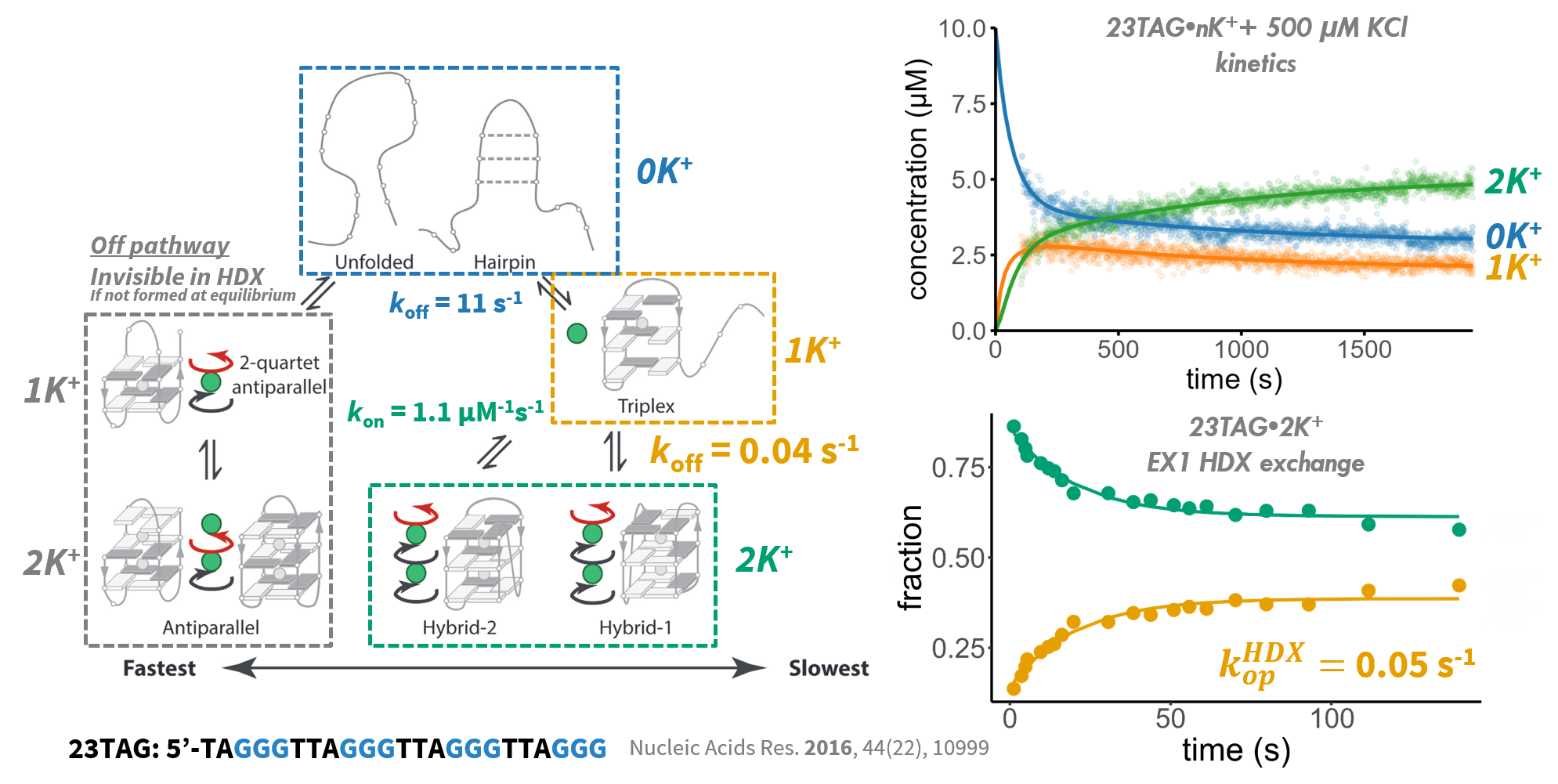

Largy, Ranz, Gabelica, J. Am. Chem. Soc. 2023, 145, 49, 26843
Drift-tube IMS can separate analytes by shape

Drift-tube IMS can separate analytes by shape


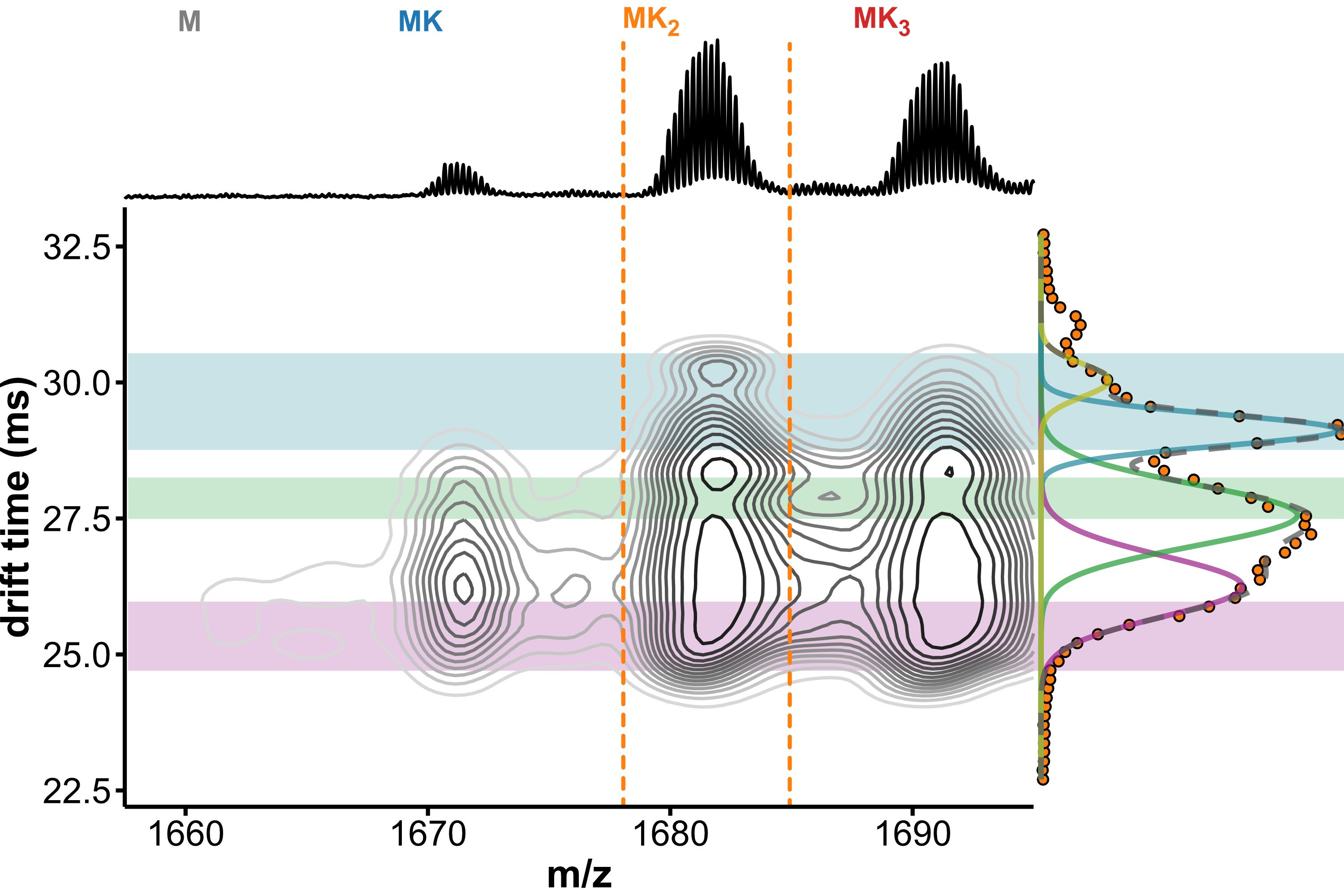
Minor conformers revealed by HDX/IMS
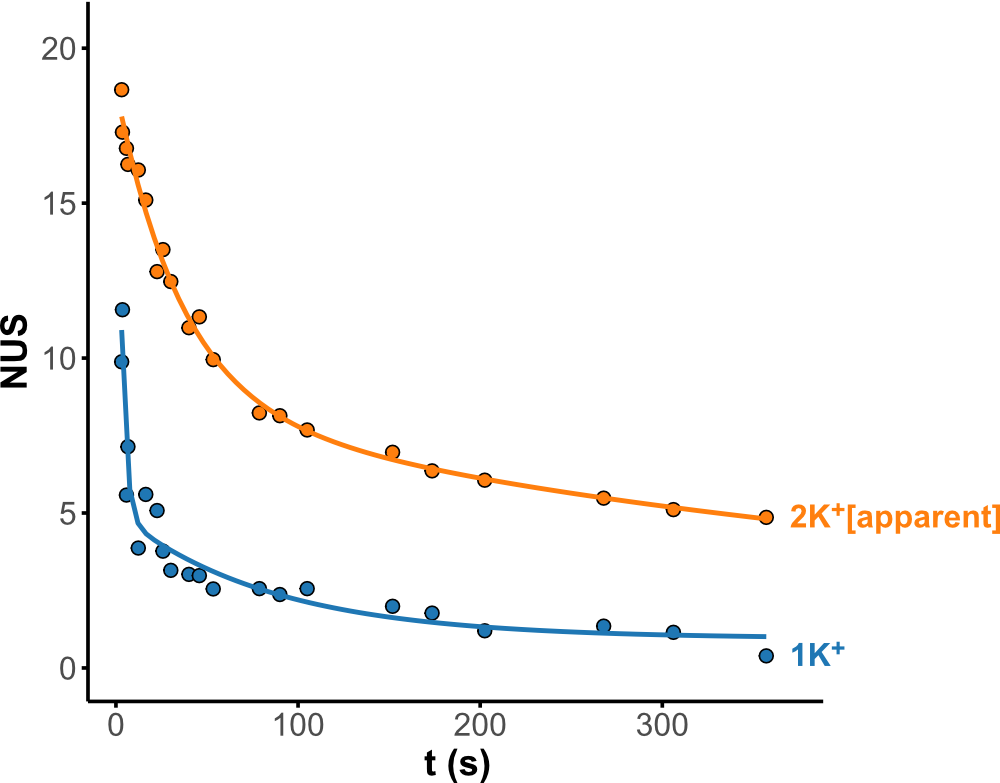

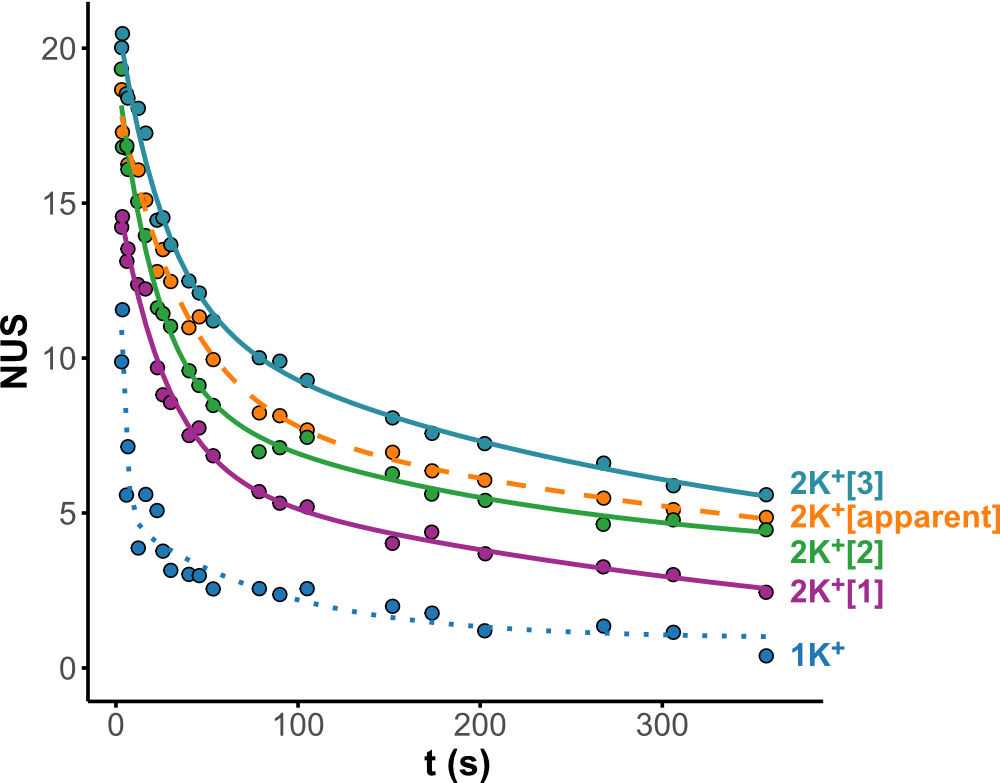
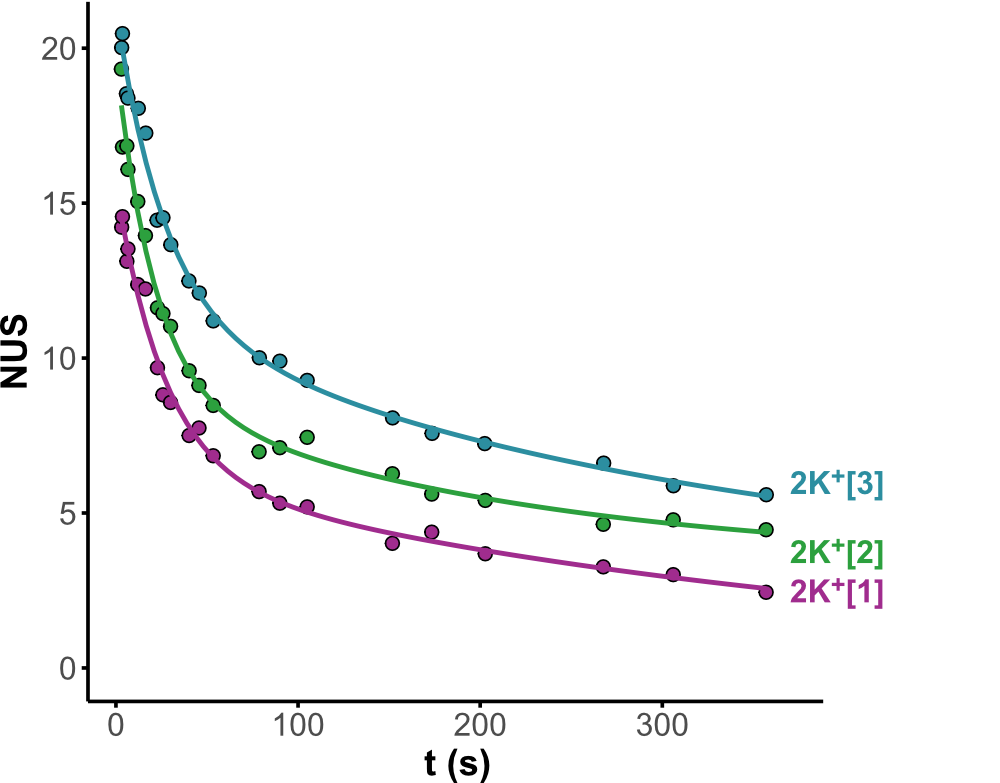
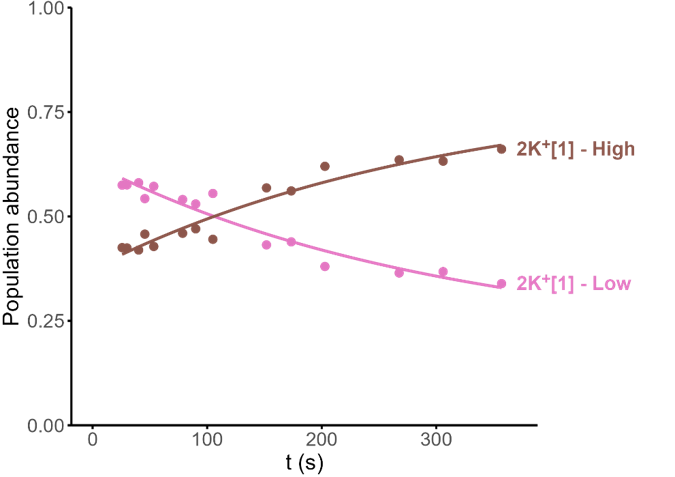
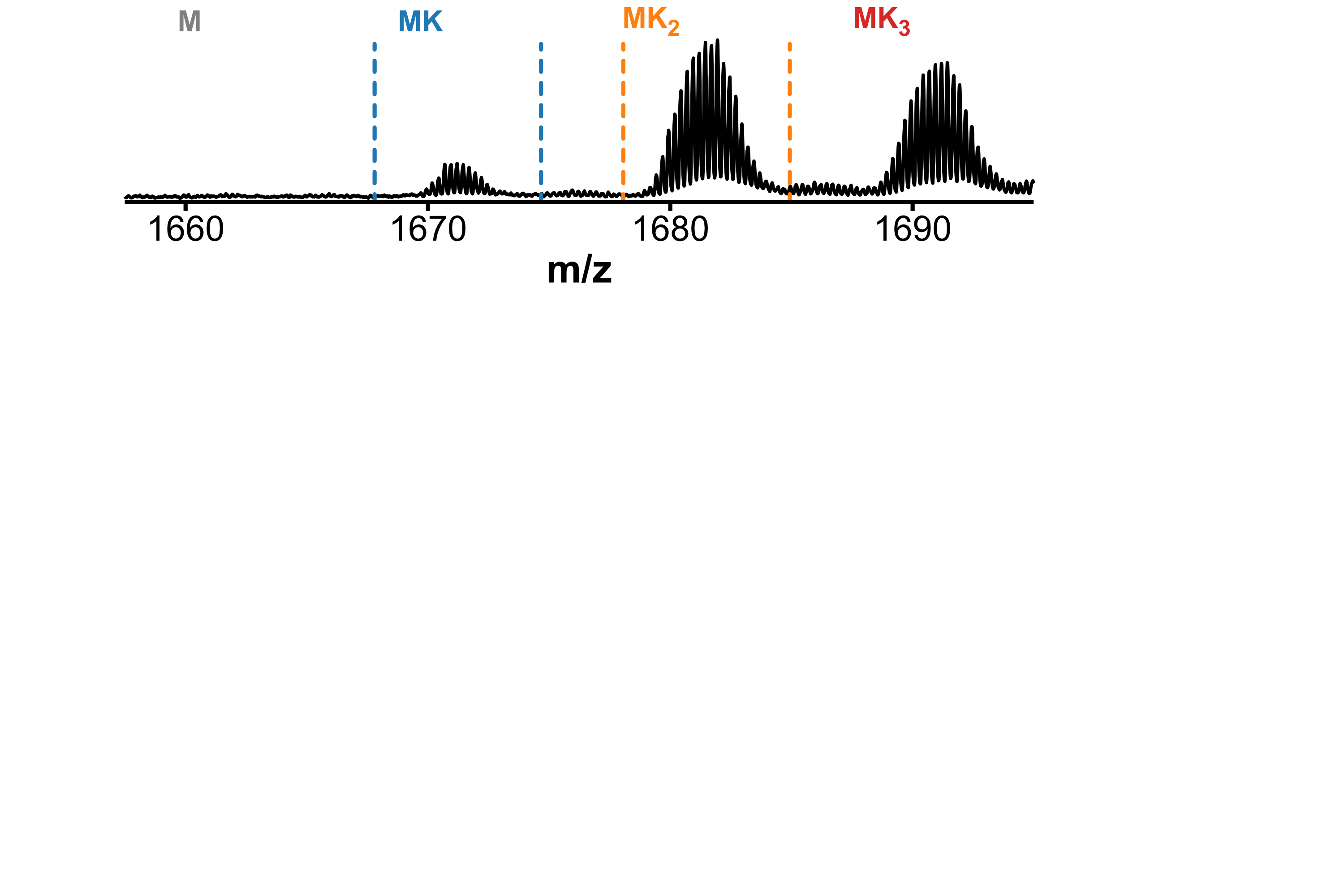
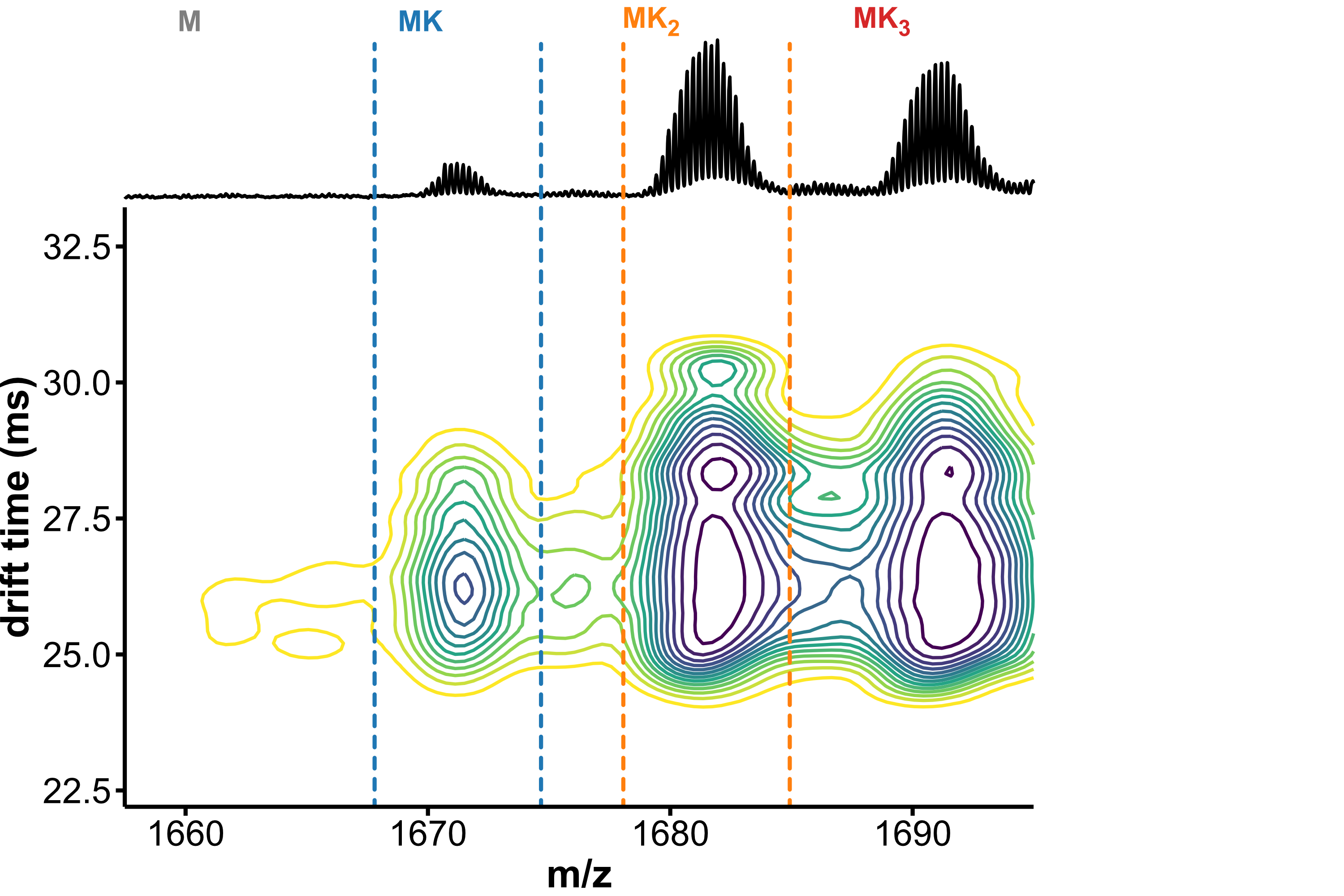

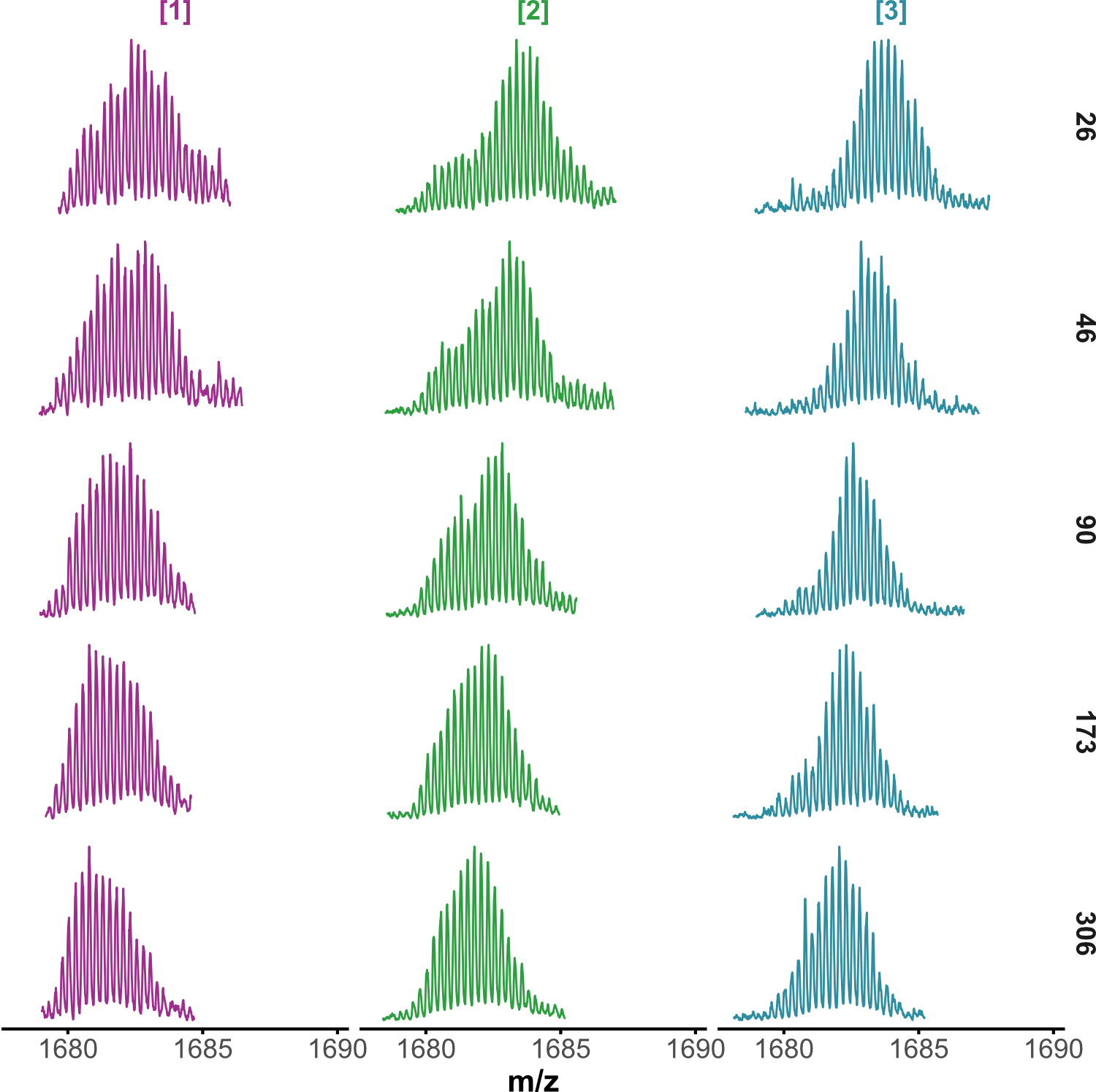
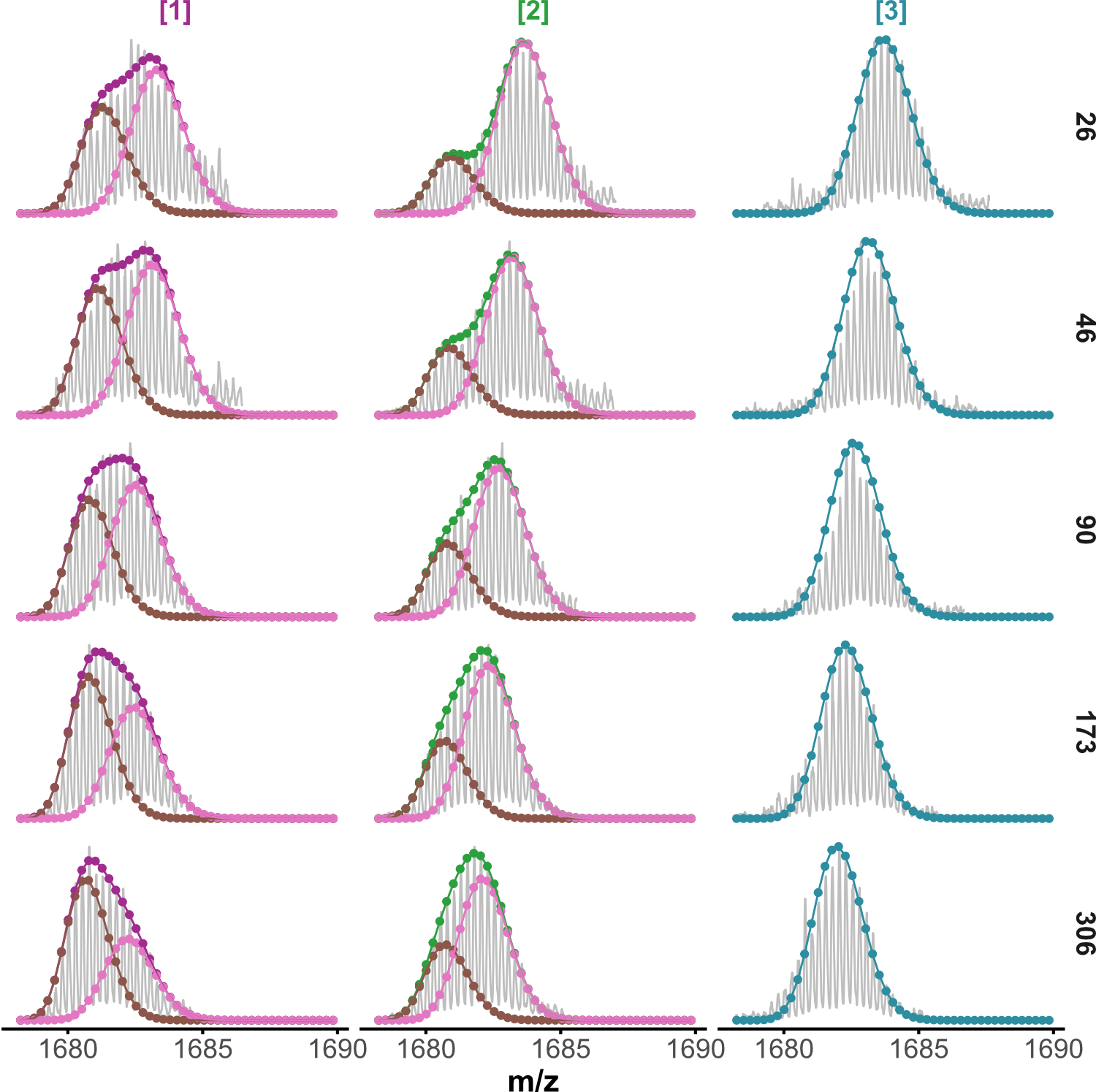



HDX kinetics inform on binding modes
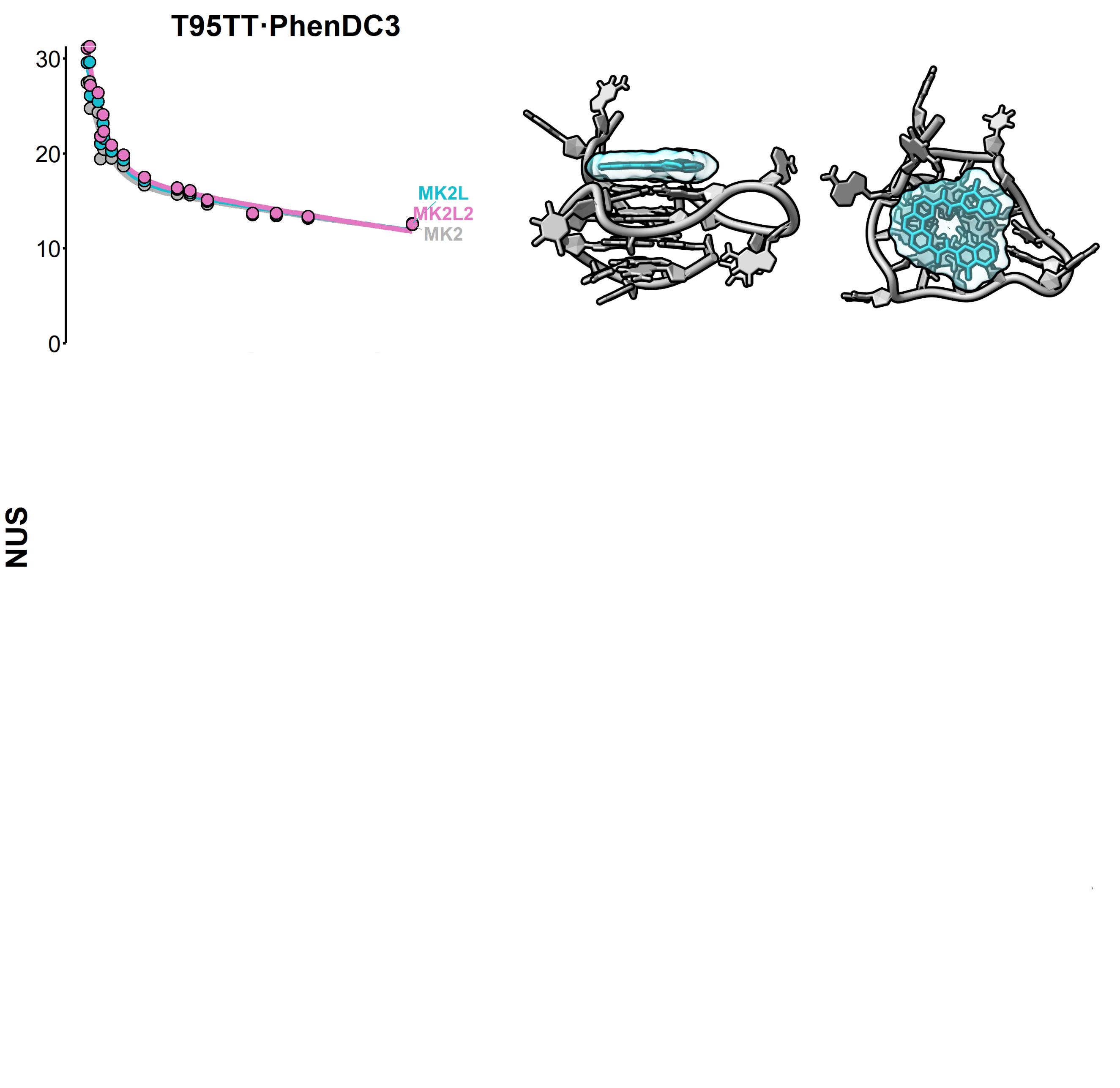
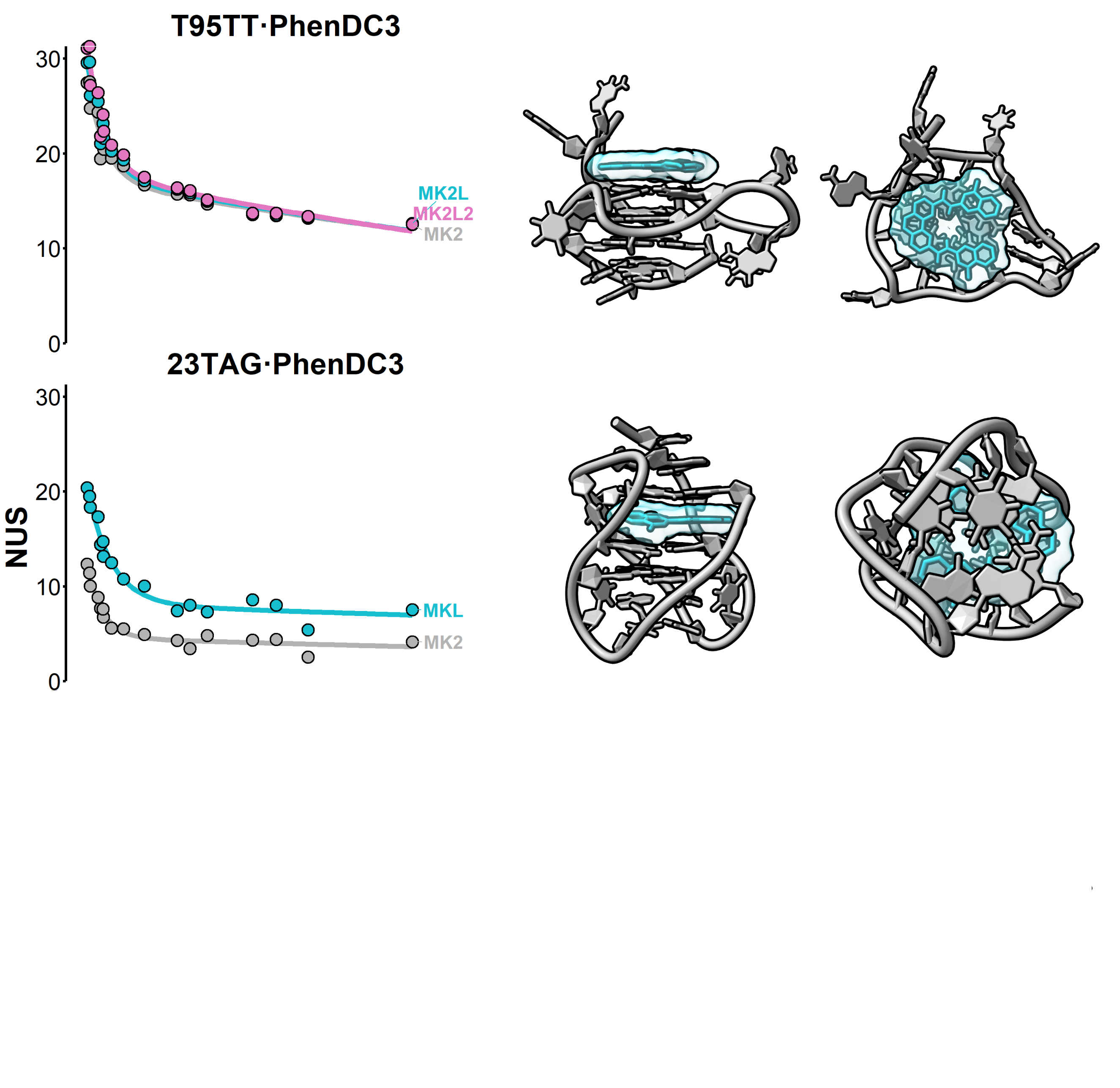
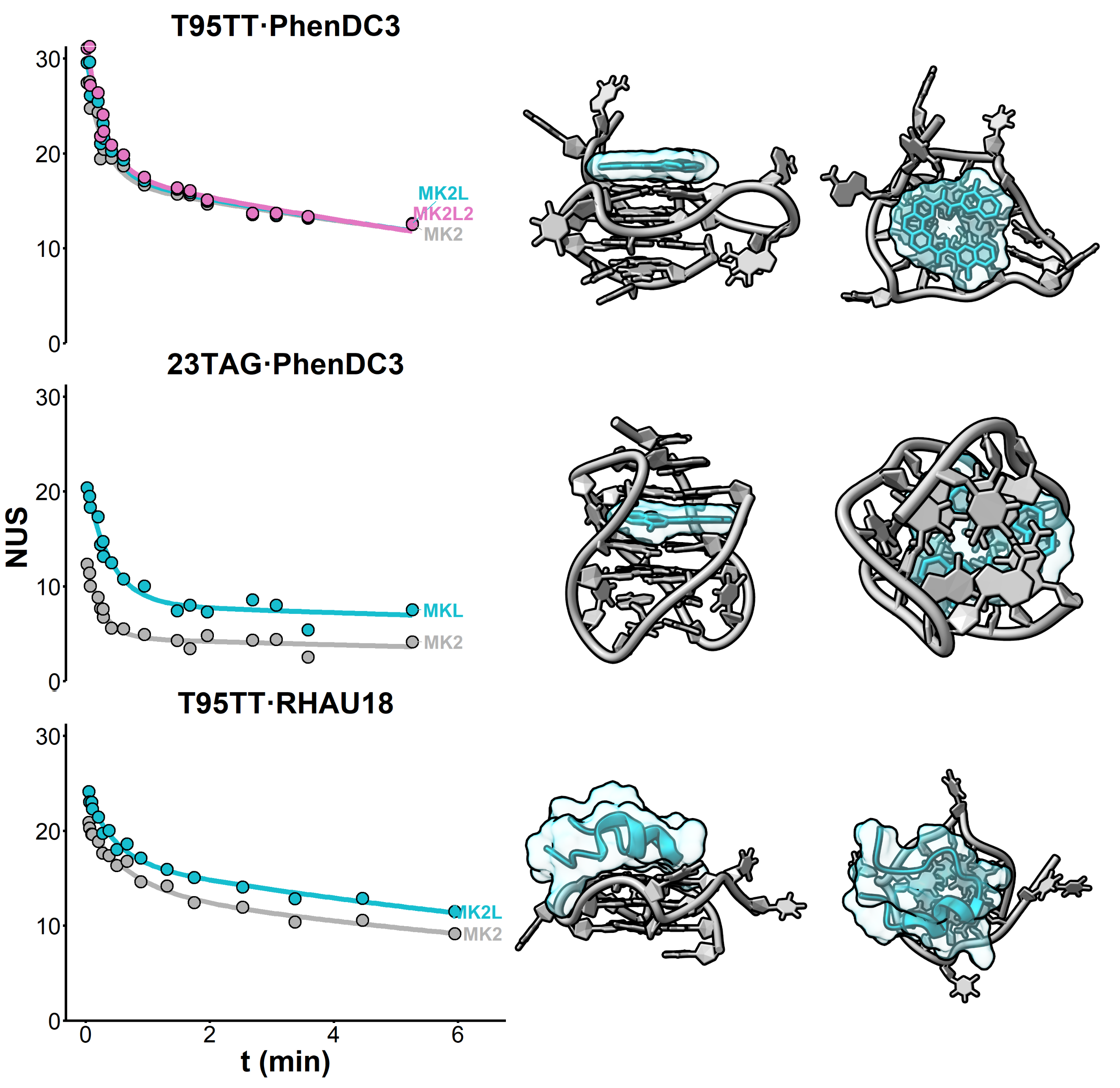


Beating structure prediction models?
Feeding structure prediction models
Aptamer structures are seldom characterized
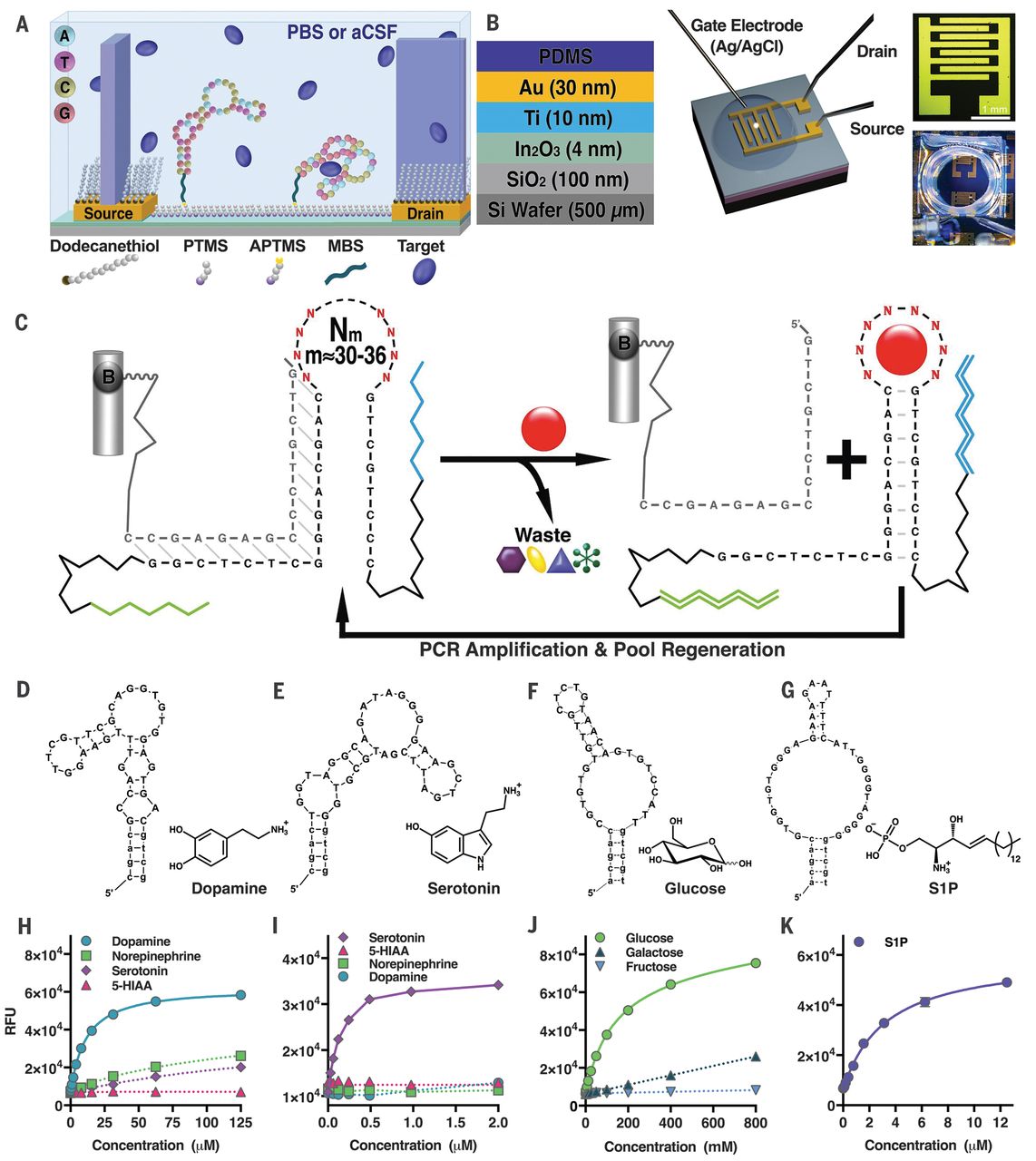
Nakatsuka N., et al., Science, 2018, 362, 319-324
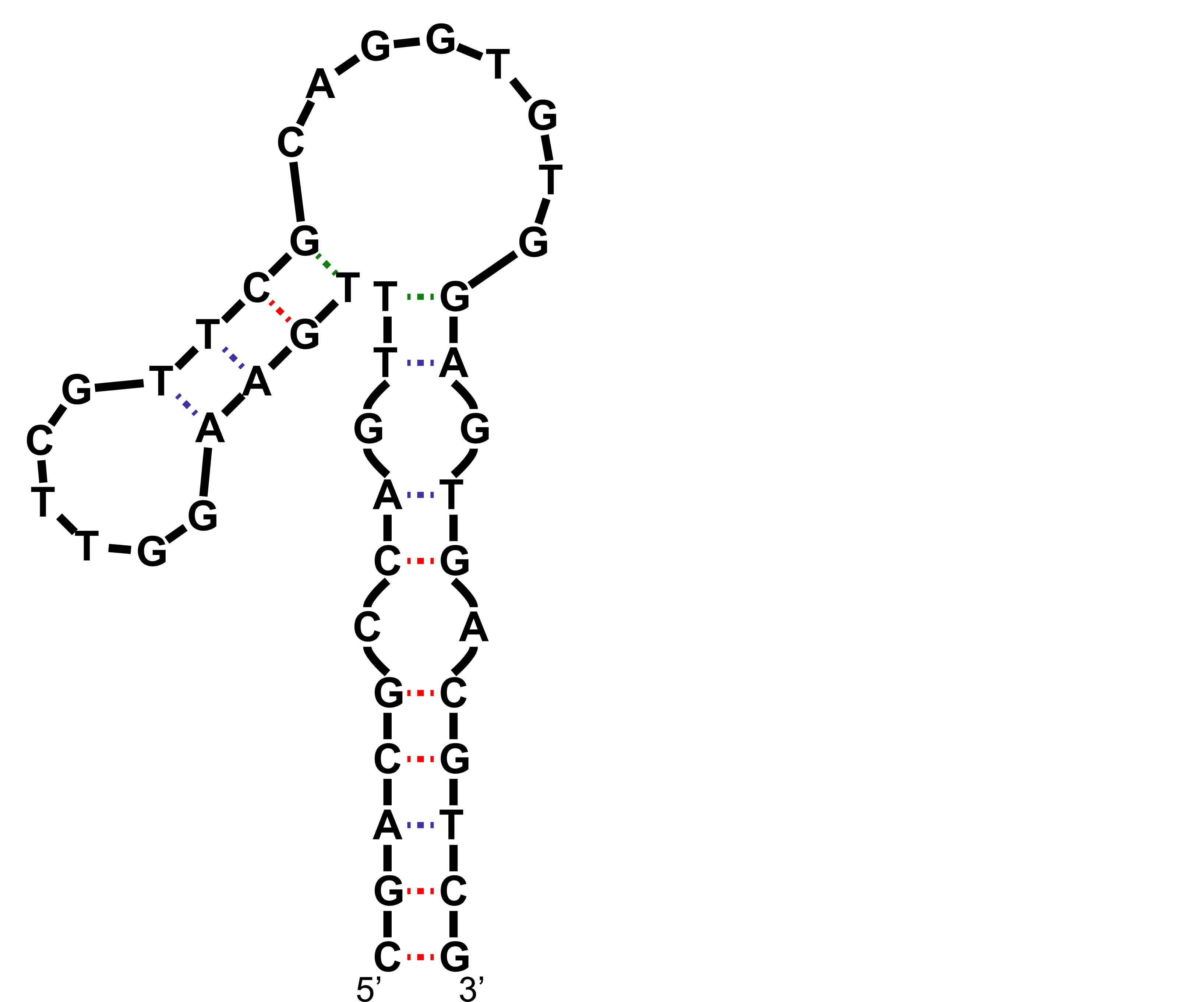
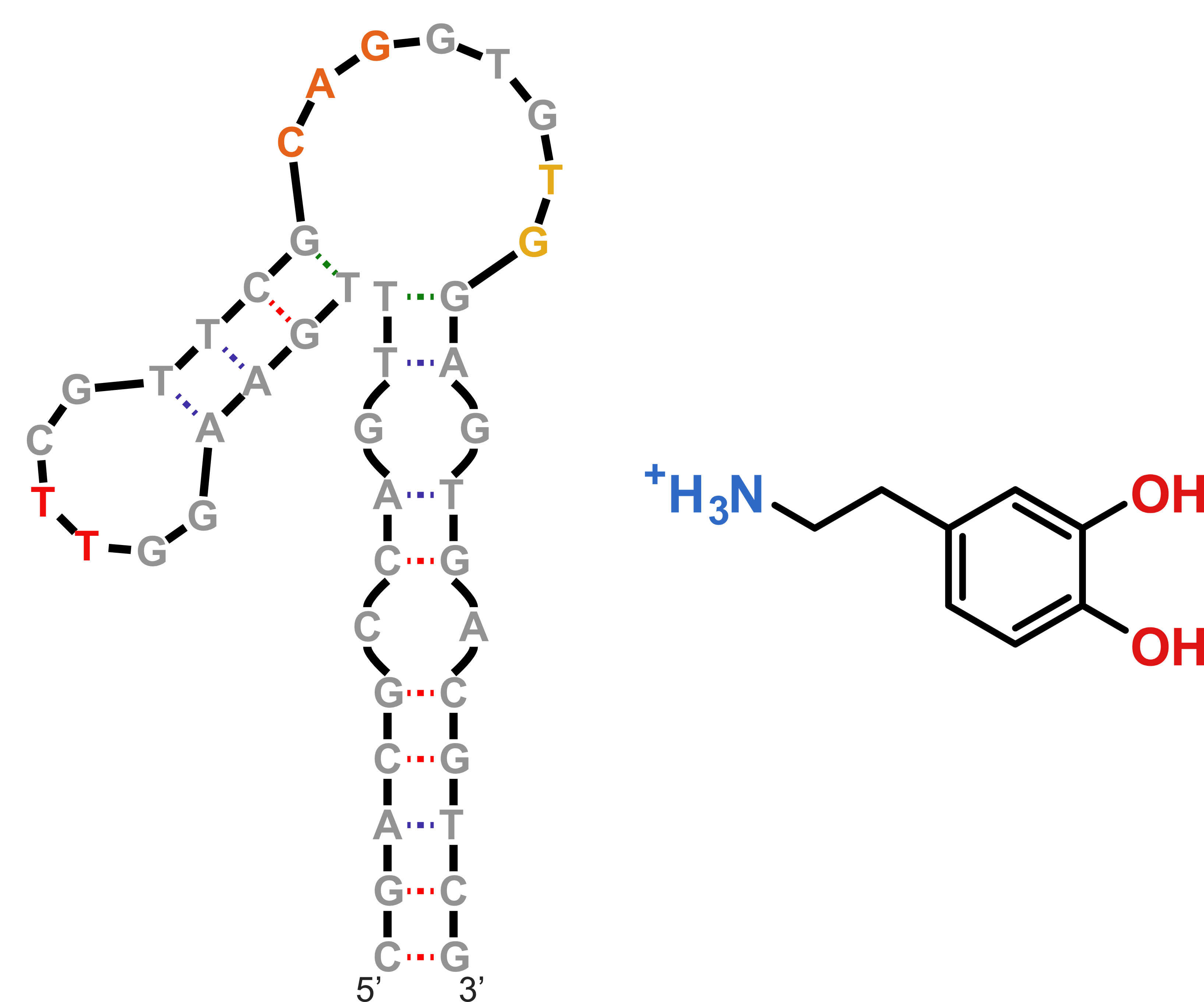
NMR provides (non-canonical !)
starting models
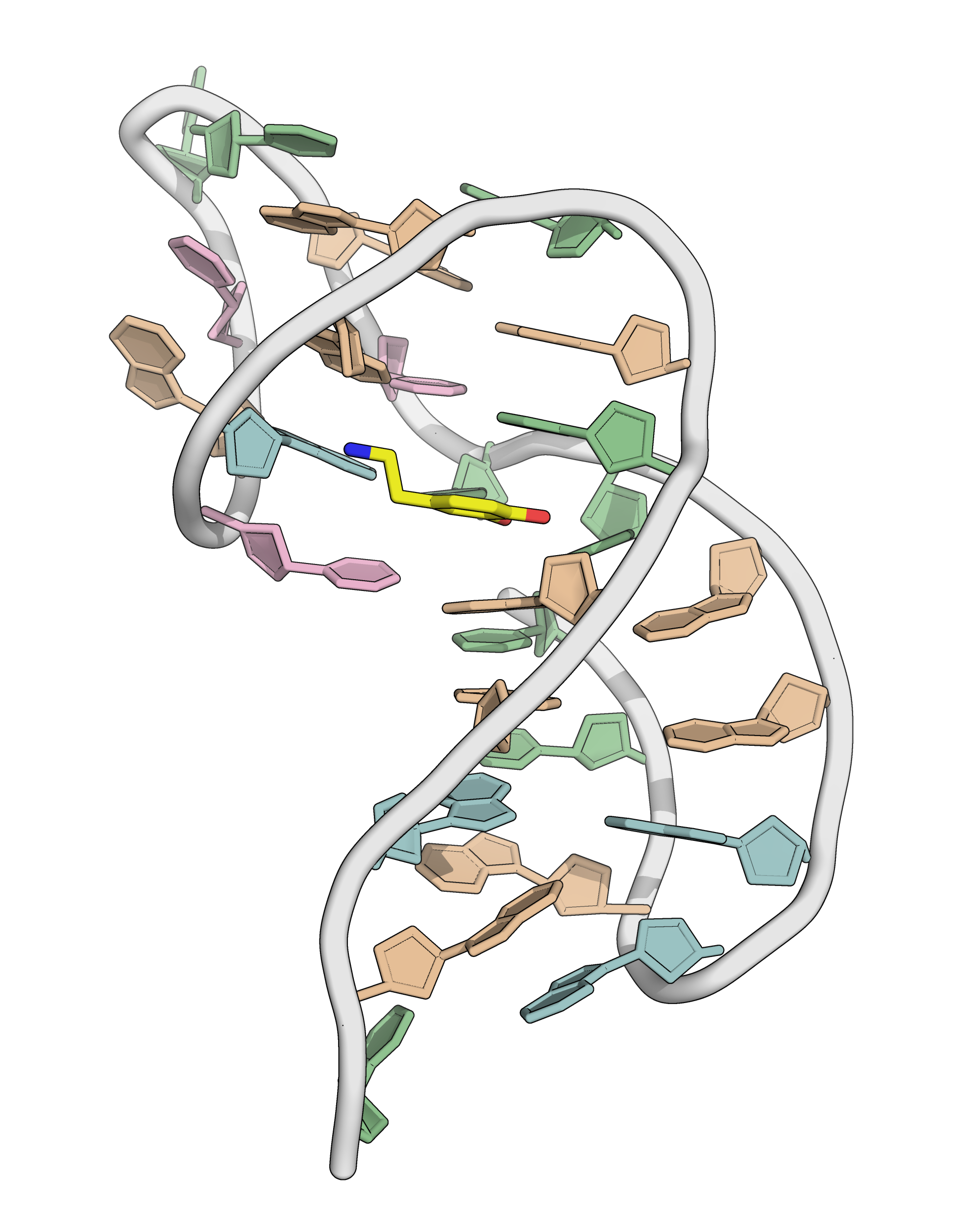

Hoi Pui Chao, E., Largy, E., Kaiyum, Y. A., Nguyen, M.-D., Vialet, B., Dauphin-Ducharme, P., Johnson, P. E., Mackereth, C.D., Nat. Commun., in revision
NMR provides many constraints
- DNA (806)
- intra-residue
- inter-residue
- Dopamine (67)
- intramolecular
- intermolecular
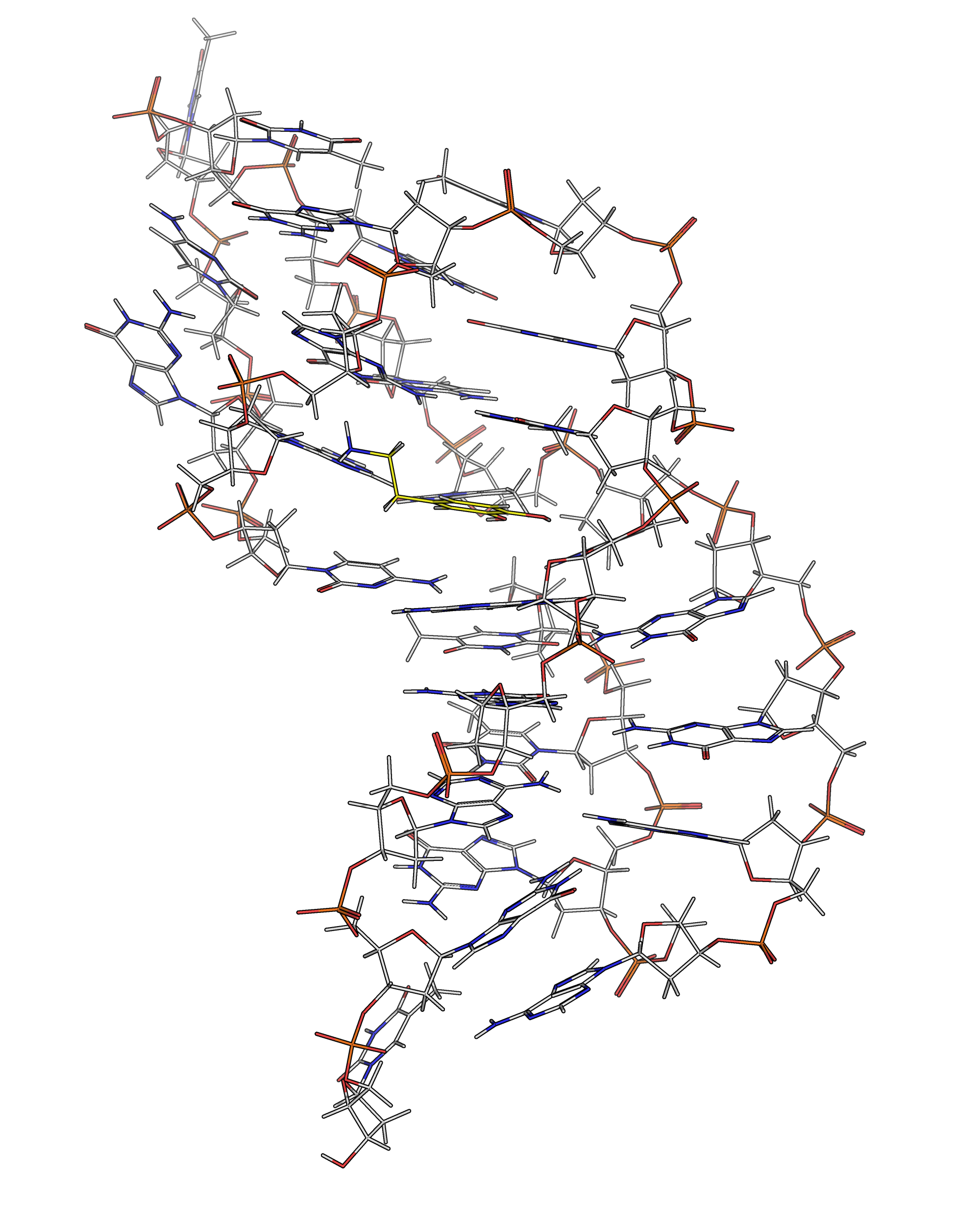
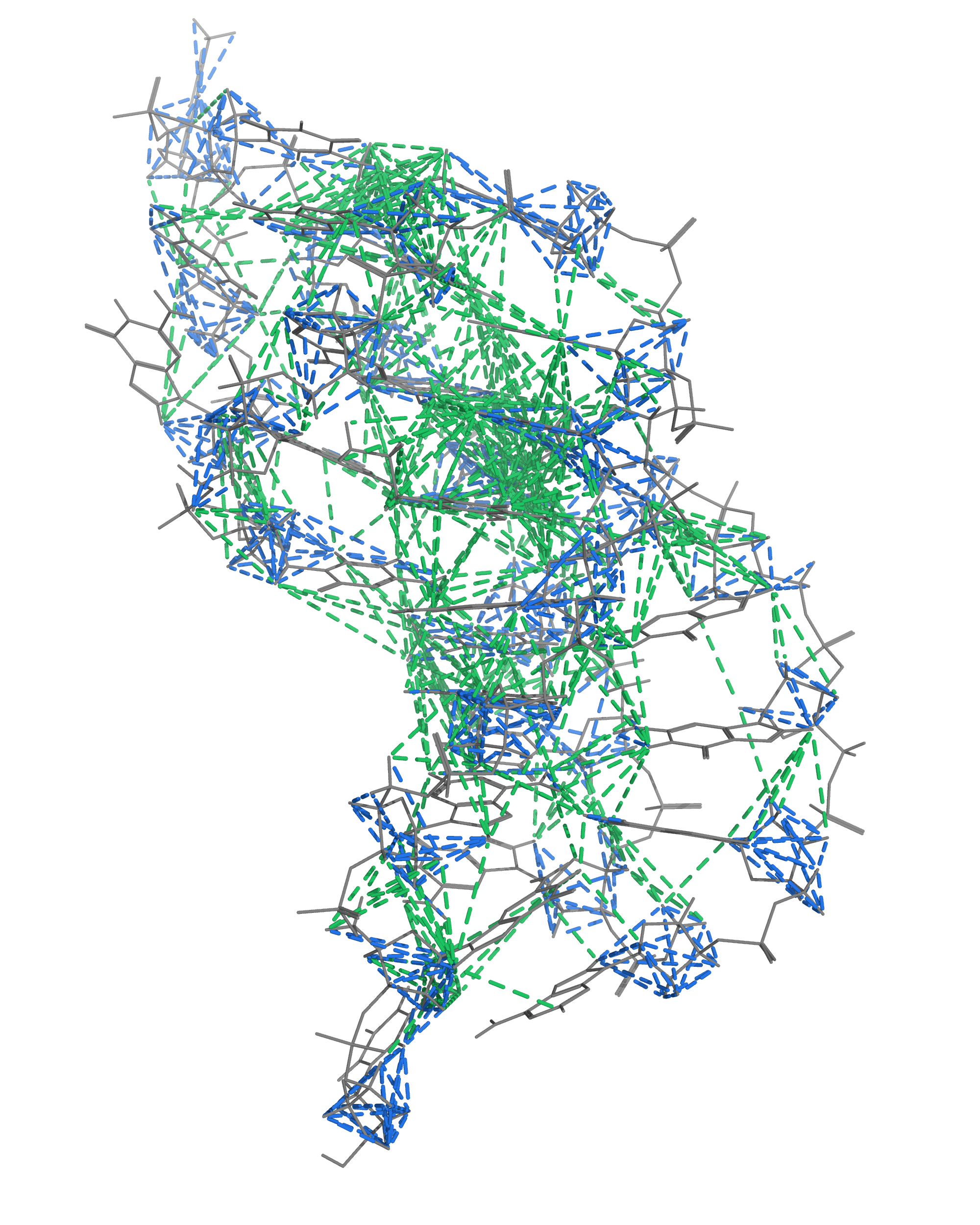
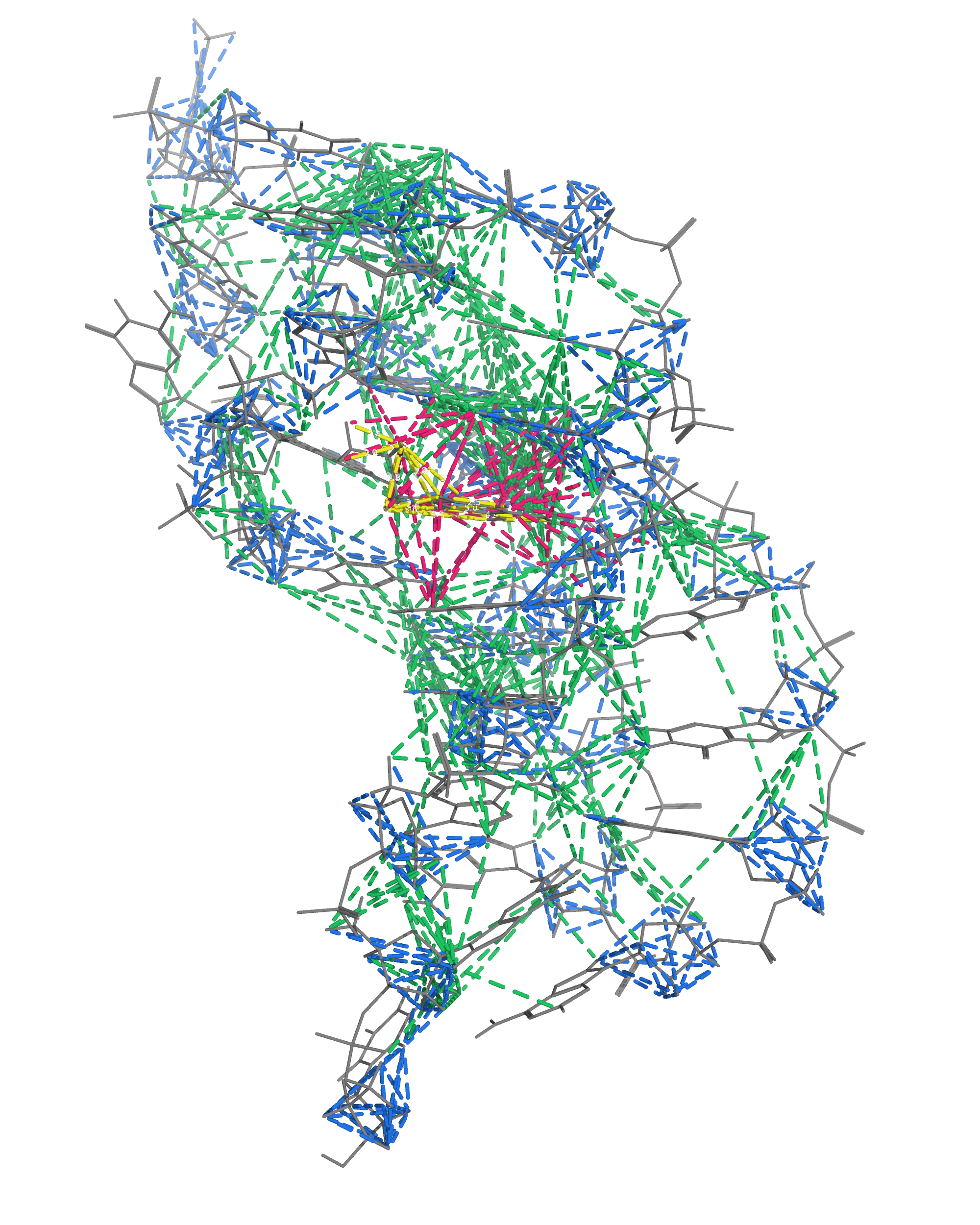

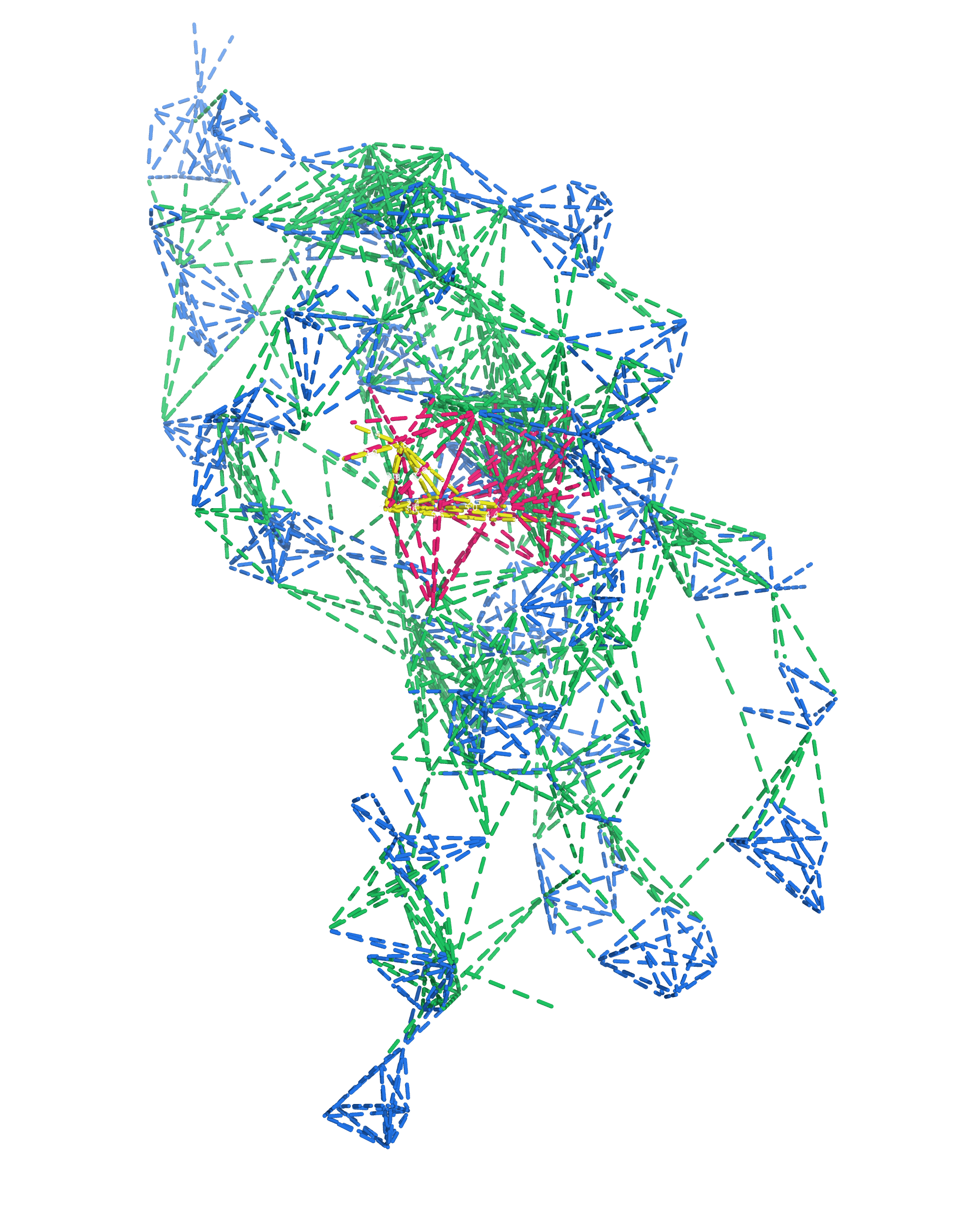

Hoi Pui Chao, E., Largy, E., Kaiyum, Y. A., Nguyen, M.-D., Vialet, B., Dauphin-Ducharme, P., Johnson, P. E., Mackereth, C.D., Nat. Commun., in revision
MD simulations informs on dynamics
Hoi Pui Chao, E., Largy, E., Kaiyum, Y. A., Nguyen, M.-D., Vialet, B., Dauphin-Ducharme, P., Johnson, P. E., Mackereth, C.D., Nat. Commun., in revision
A minimized ensemble that cannot be predicted
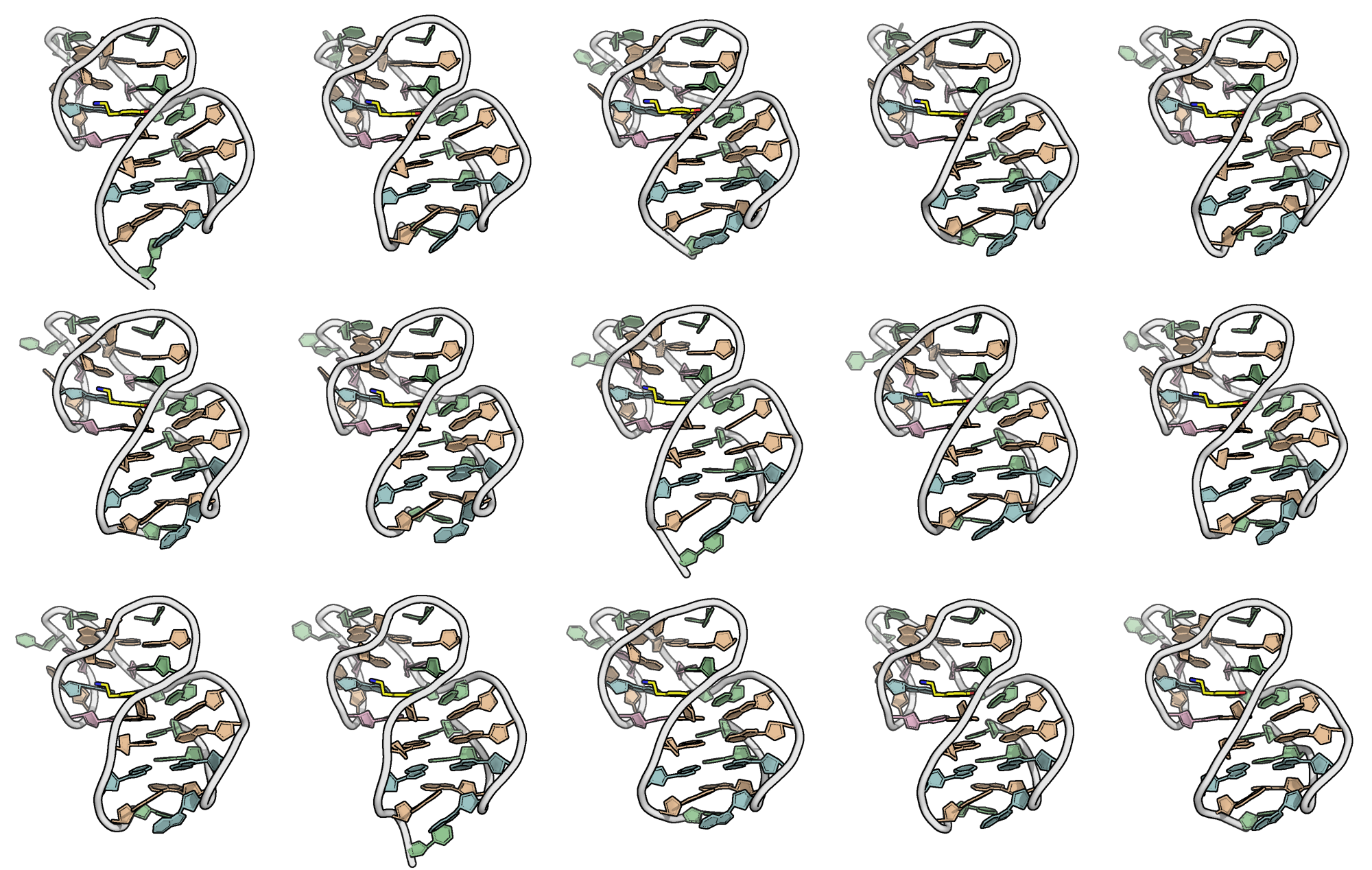
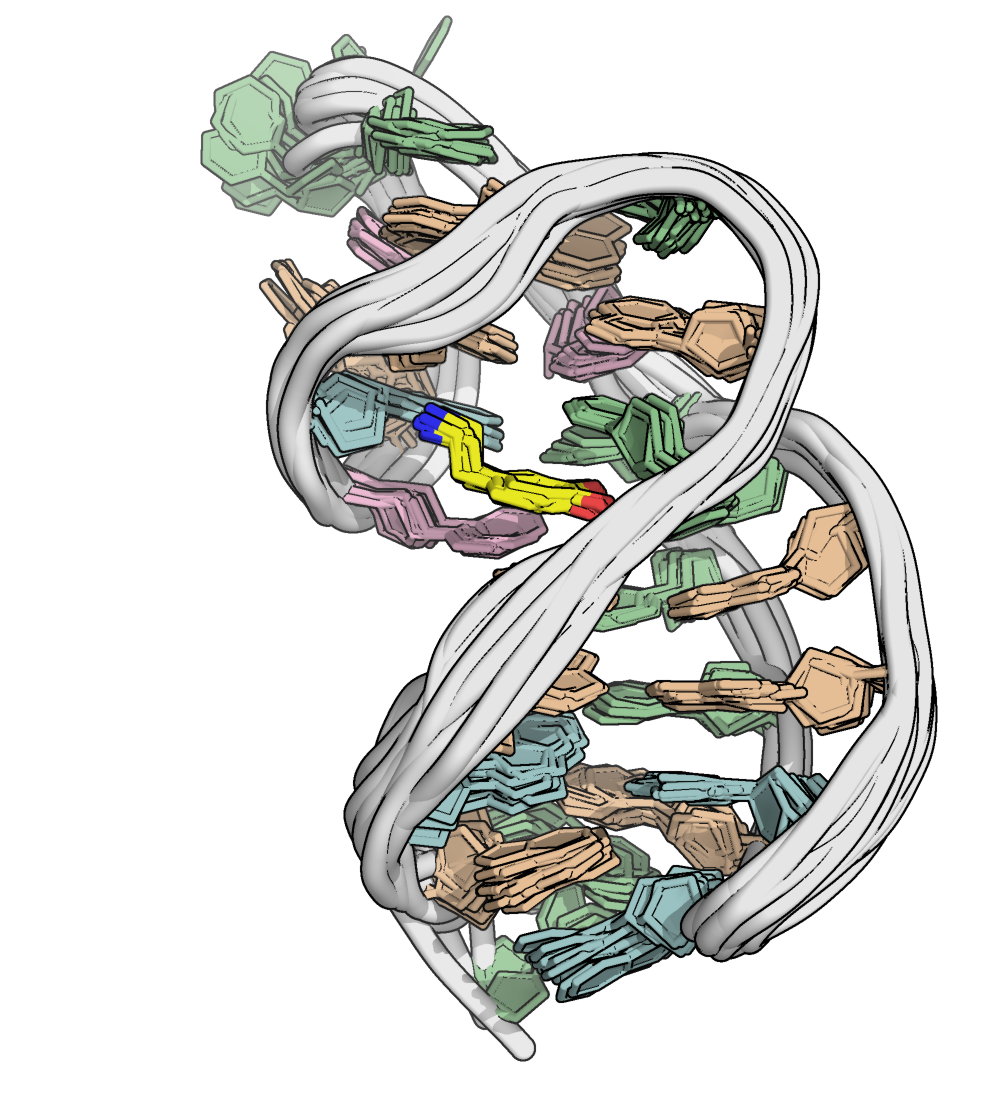
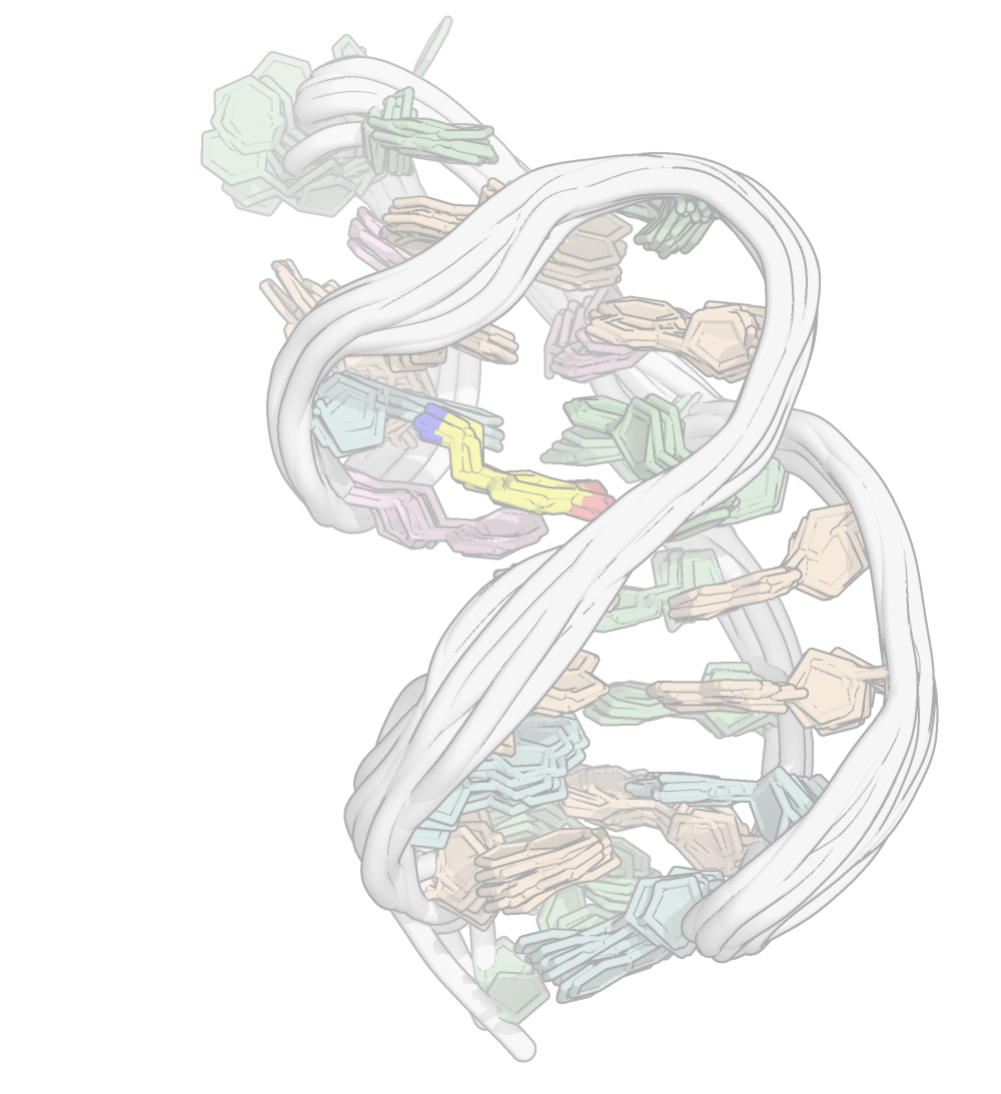

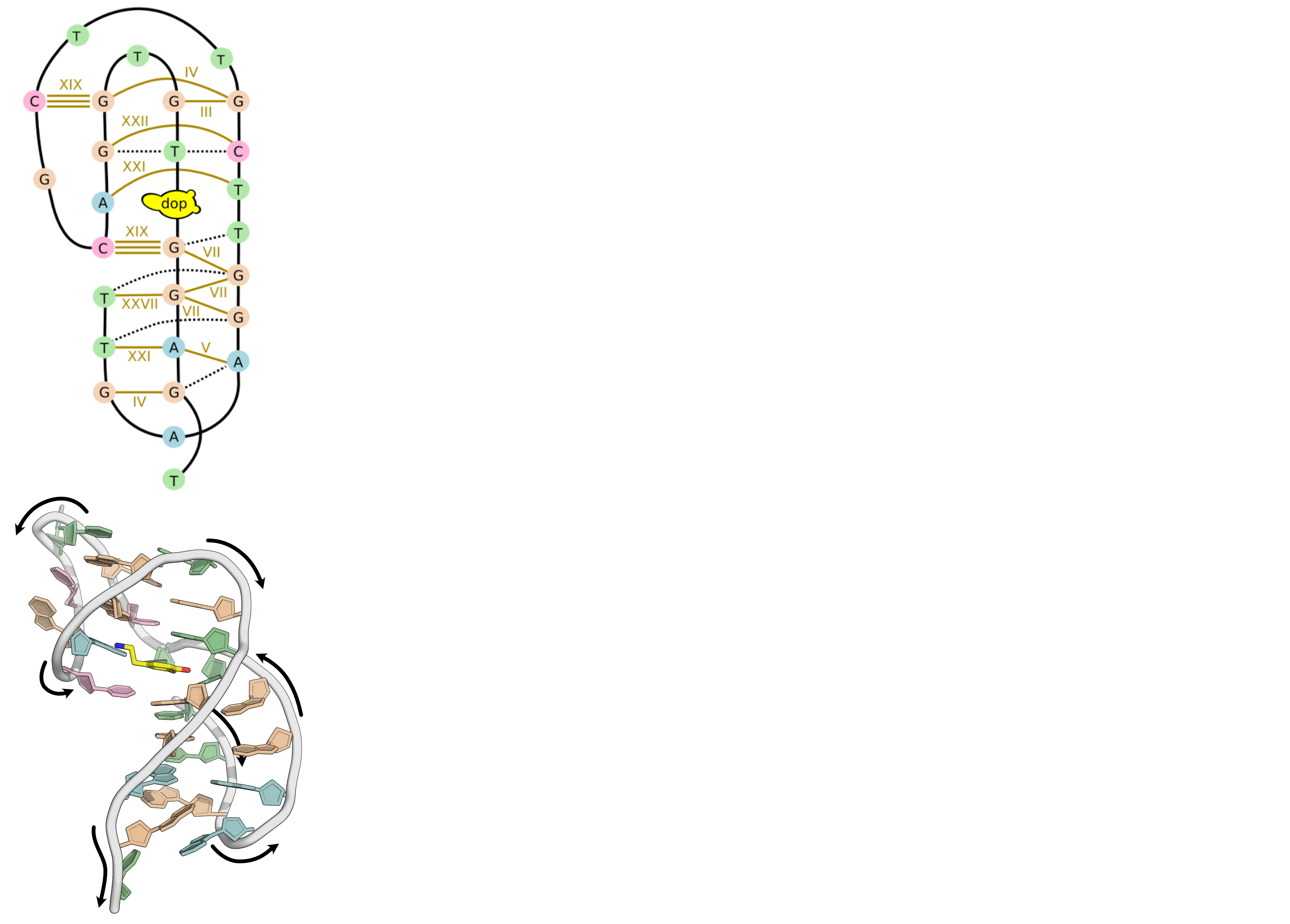
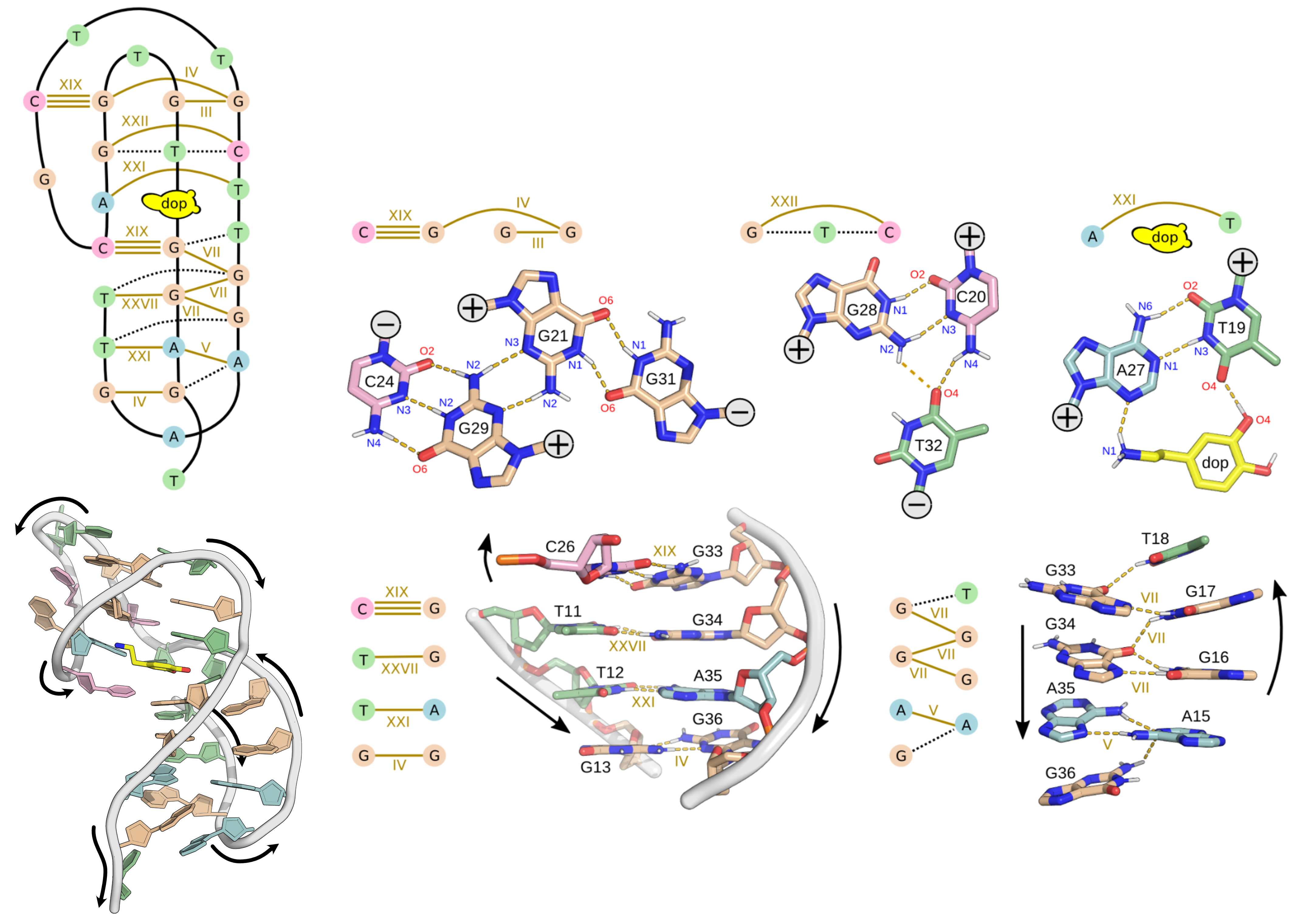
- CASP16
Critical Assessment of Techniques for Protein Structure Prediction- 107 models submitted
- 0 with RMSD < 10 Å

Kretsch R.C., et al., Proteins, 2025, prot.70043
Hoi Pui Chao, E., Largy, E., Kaiyum, Y. A., Nguyen, M.-D., Vialet, B., Dauphin-Ducharme, P., Johnson, P. E., Mackereth, C.D., Nat. Commun., in revision
The structural landscape
of DNA is large
A robust method toolbox is required to explore it
Acknowledgements
Université de Bordeaux
- Cameron Mackereth
- Samir Amrane
- Stéphane Thore
- Yann Ferrand
- Vincent Laffilé
Université de Bordeaux → Institut Polytechnique de Paris
- Jean-Louis Mergny
- Aurore Guedin-Beaurepaire
- Marie Toulisse
Université de Bordeaux → Université de Genève
- Valérie Gabelica
- Frédéric Rosu
- Clarisse Fourel
- Anirban Ghosh
- Romane Guisiano
- Alexander König
- Adrien Marchand
- Matthieu Ranz
- Liliya Yatsunyk
York University
- Philip Johnson
- Emily Chao
Université de Sherbrooke
- Philippe Dauphin Ducharme
LMU München
- Yvan Huc
- Jonas Sigl
Quality Assistance &
Ose Immunotherapeutics
Funding sources


Thank you for your attention!